Details of the Target
General Information of Target
Target Site Mutations in Different Cell Lines
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
YN-1 Probe Info |
 |
100.00 | LDD0444 | [1] | |
|
STPyne Probe Info |
 |
K277(7.08) | LDD0277 | [2] | |
|
OPA-S-S-alkyne Probe Info |
 |
K63(2.75); K78(3.33); K47(5.07); K178(5.39) | LDD3494 | [3] | |
|
Probe 1 Probe Info |
 |
Y75(42.19); Y280(18.89) | LDD3495 | [4] | |
|
DBIA Probe Info |
 |
C50(1.96) | LDD3310 | [5] | |
|
BTD Probe Info |
 |
C74(1.90) | LDD1700 | [6] | |
|
IA-alkyne Probe Info |
 |
C50(0.00); C153(0.00) | LDD0162 | [7] | |
|
NAIA_5 Probe Info |
 |
C153(0.00); C50(0.00); C74(0.00) | LDD2223 | [8] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C170 Probe Info |
 |
11.88 | LDD1850 | [9] | |
|
C186 Probe Info |
 |
7.62 | LDD1864 | [9] | |
|
C208 Probe Info |
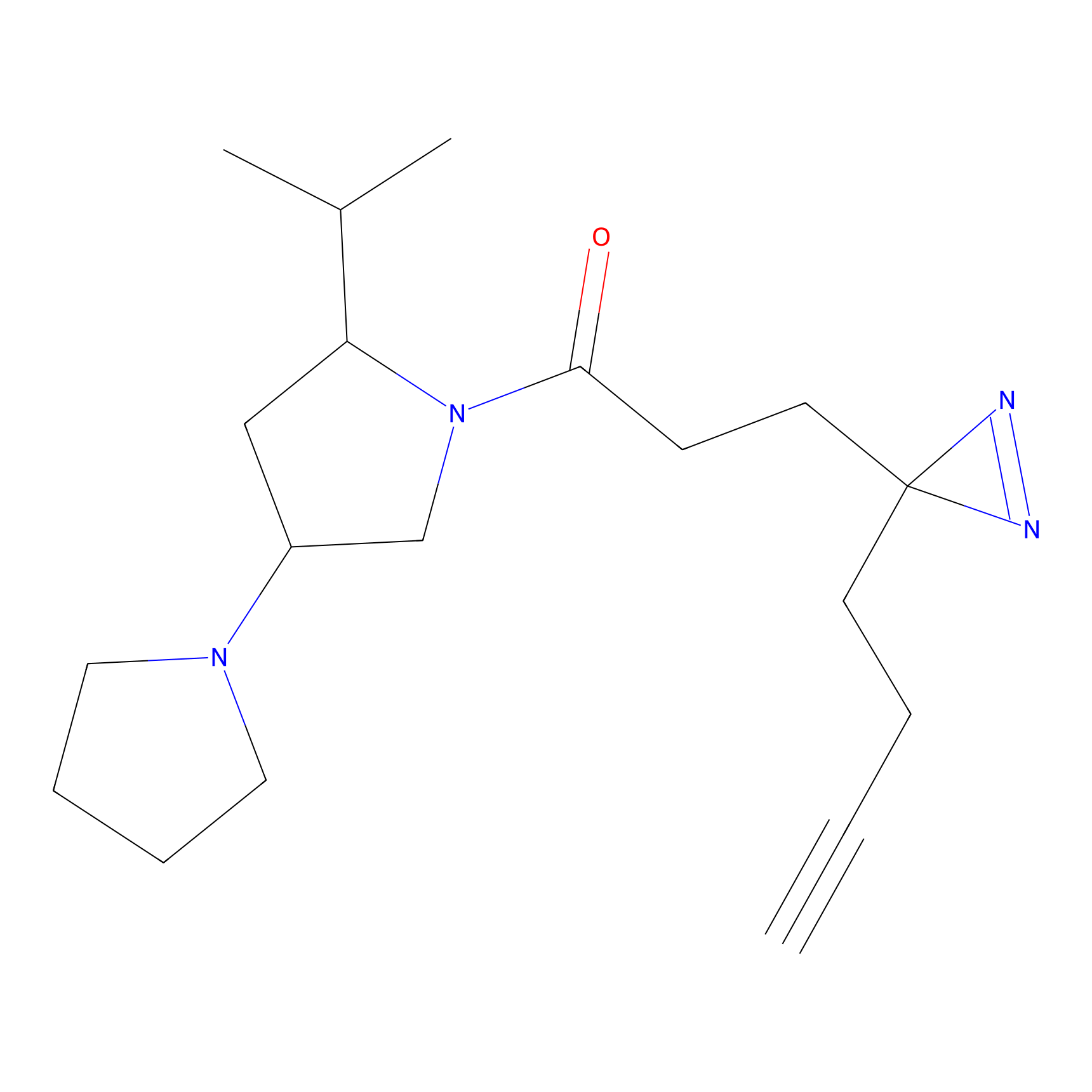 |
4.92 | LDD1883 | [9] | |
|
C302 Probe Info |
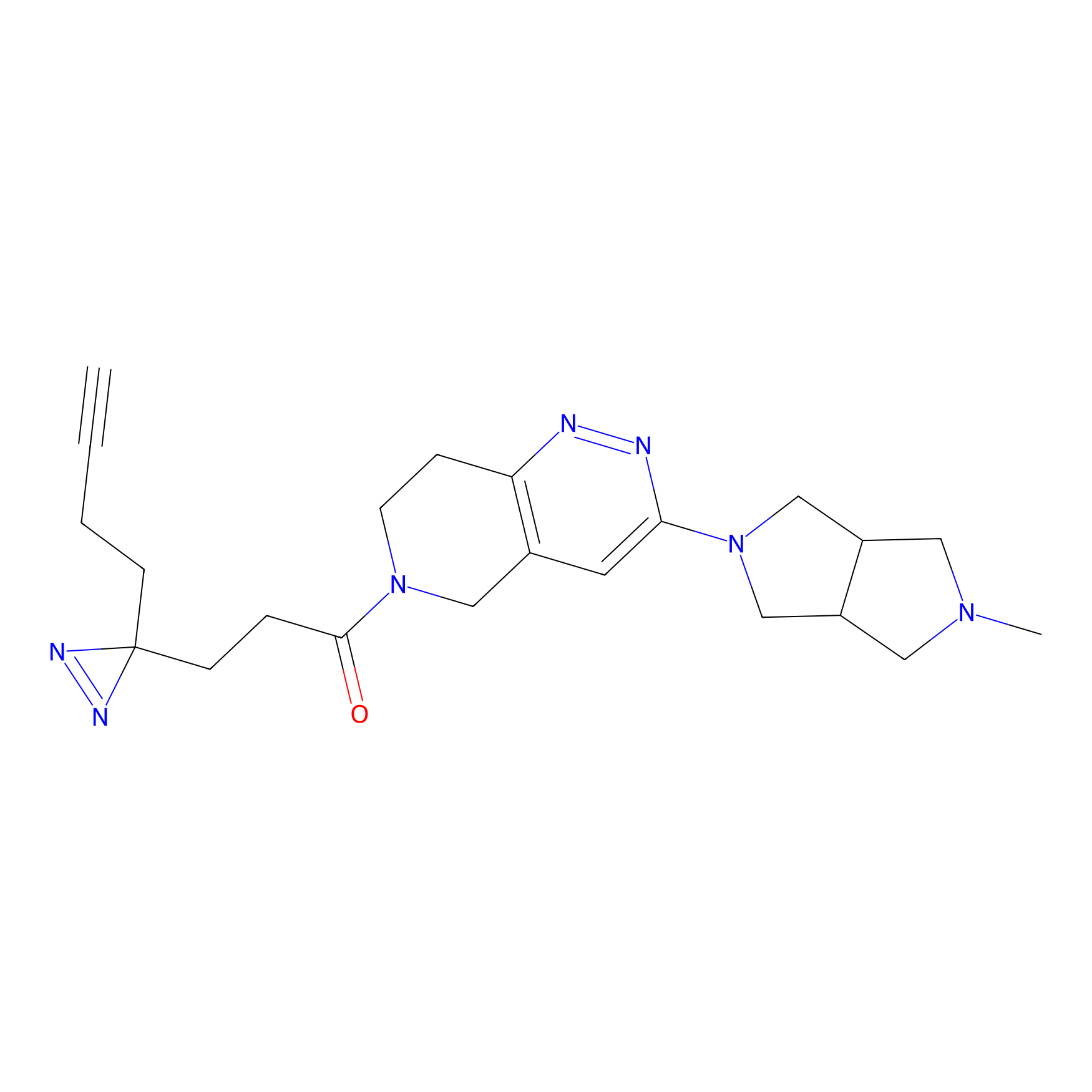 |
6.82 | LDD1971 | [9] | |
|
C310 Probe Info |
 |
8.46 | LDD1977 | [9] | |
|
C425 Probe Info |
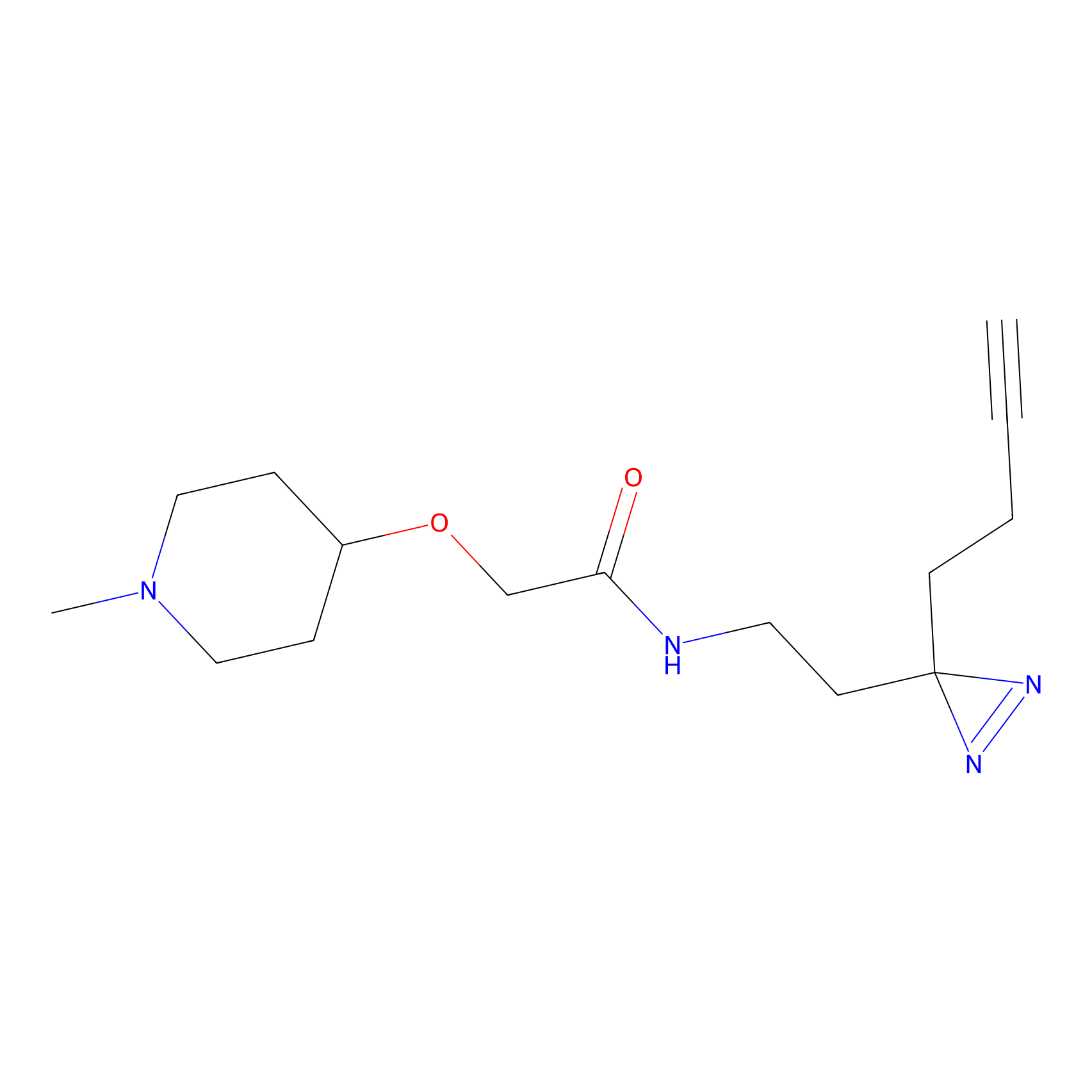 |
5.10 | LDD2080 | [9] | |
|
FFF probe11 Probe Info |
 |
17.41 | LDD0472 | [10] | |
|
FFF probe12 Probe Info |
 |
20.00 | LDD0473 | [10] | |
|
VE-P Probe Info |
 |
N.A. | LDD0396 | [11] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0371 | CL102 | HEK-293T | C74(0.94) | LDD1575 | [12] |
| LDCM0375 | CL106 | HEK-293T | C74(0.91) | LDD1579 | [12] |
| LDCM0380 | CL110 | HEK-293T | C74(0.92) | LDD1584 | [12] |
| LDCM0384 | CL114 | HEK-293T | C74(0.93) | LDD1588 | [12] |
| LDCM0388 | CL118 | HEK-293T | C74(1.02) | LDD1592 | [12] |
| LDCM0393 | CL122 | HEK-293T | C74(1.12) | LDD1597 | [12] |
| LDCM0397 | CL126 | HEK-293T | C74(0.93) | LDD1601 | [12] |
| LDCM0401 | CL14 | HEK-293T | C74(1.05) | LDD1605 | [12] |
| LDCM0407 | CL2 | HEK-293T | C74(1.07) | LDD1611 | [12] |
| LDCM0414 | CL26 | HEK-293T | C74(0.99) | LDD1618 | [12] |
| LDCM0441 | CL50 | HEK-293T | C74(1.10) | LDD1645 | [12] |
| LDCM0454 | CL62 | HEK-293T | C74(1.03) | LDD1657 | [12] |
| LDCM0467 | CL74 | HEK-293T | C74(1.15) | LDD1670 | [12] |
| LDCM0480 | CL86 | HEK-293T | C74(0.93) | LDD1683 | [12] |
| LDCM0493 | CL98 | HEK-293T | C74(0.98) | LDD1696 | [12] |
| LDCM0427 | Fragment51 | HEK-293T | C74(1.00) | LDD1631 | [12] |
| LDCM0022 | KB02 | A-549 | C153(1.50) | LDD2260 | [5] |
| LDCM0023 | KB03 | A2780 | C50(1.65) | LDD2671 | [5] |
| LDCM0024 | KB05 | COLO792 | C50(1.96) | LDD3310 | [5] |
| LDCM0506 | Nucleophilic fragment 16a | MDA-MB-231 | C74(1.09) | LDD2099 | [6] |
| LDCM0514 | Nucleophilic fragment 20a | MDA-MB-231 | C74(0.89) | LDD2107 | [6] |
| LDCM0516 | Nucleophilic fragment 21a | MDA-MB-231 | C74(0.66) | LDD2109 | [6] |
| LDCM0522 | Nucleophilic fragment 24a | MDA-MB-231 | C74(0.61) | LDD2115 | [6] |
| LDCM0530 | Nucleophilic fragment 28a | MDA-MB-231 | C74(0.88) | LDD2123 | [6] |
| LDCM0532 | Nucleophilic fragment 29a | MDA-MB-231 | C74(1.10) | LDD2125 | [6] |
| LDCM0534 | Nucleophilic fragment 30a | MDA-MB-231 | C74(0.68) | LDD2127 | [6] |
| LDCM0541 | Nucleophilic fragment 36 | MDA-MB-231 | C74(0.60) | LDD2134 | [6] |
| LDCM0543 | Nucleophilic fragment 38 | MDA-MB-231 | C74(0.83) | LDD2136 | [6] |
| LDCM0544 | Nucleophilic fragment 39 | MDA-MB-231 | C74(0.78) | LDD2137 | [6] |
| LDCM0211 | Nucleophilic fragment 3b | MDA-MB-231 | C74(1.90) | LDD1700 | [6] |
| LDCM0554 | Nucleophilic fragment 7a | MDA-MB-231 | C74(0.55) | LDD2148 | [6] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transporter and channel
Other
References
