Details of the Target
General Information of Target
| Target ID | LDTP14136 | |||||
|---|---|---|---|---|---|---|
| Target Name | Mitochondrial import inner membrane translocase subunit Tim13 (TIMM13) | |||||
| Gene Name | TIMM13 | |||||
| Gene ID | 26517 | |||||
| Synonyms |
TIM13B; TIMM13A; TIMM13B; Mitochondrial import inner membrane translocase subunit Tim13 |
|||||
| 3D Structure | ||||||
| Sequence |
MAGKVRSLLPPLLLAAAGLAGLLLLCVPTRDVREPPALKYGIVLDAGSSHTSMFIYKWPA
DKENDTGIVGQHSSCDVPGGGISSYADNPSGASQSLVGCLEQALQDVPKERHAGTPLYLG ATAGMRLLNLTNPEASTSVLMAVTHTLTQYPFDFRGARILSGQEEGVFGWVTANYLLENF IKYGWVGRWFRPRKGTLGAMDLGGASTQITFETTSPAEDRASEVQLHLYGQHYRVYTHSF LCYGRDQVLQRLLASALQTHGFHPCWPRGFSTQVLLGDVYQSPCTMAQRPQNFNSSARVS LSGSSDPHLCRDLVSGLFSFSSCPFSRCSFNGVFQPPVAGNFVAFSAFFYTVDFLRTSMG LPVATLQQLEAAAVNVCNQTWAQLQARVPGQRARLADYCAGAMFVQQLLSRGYGFDERAF GGVIFQKKAADTAVGWALGYMLNLTNLIPADPPGLRKGTDFSSWVVLLLLFASALLAALV LLLRQVHSAKLPSTI |
|||||
| Target Bioclass |
Other
|
|||||
| Family |
Small Tim family
|
|||||
| Subcellular location |
Mitochondrion inner membrane
|
|||||
| Function |
Mitochondrial intermembrane chaperone that participates in the import and insertion of some multi-pass transmembrane proteins into the mitochondrial inner membrane. Also required for the transfer of beta-barrel precursors from the TOM complex to the sorting and assembly machinery (SAM complex) of the outer membrane. Acts as a chaperone-like protein that protects the hydrophobic precursors from aggregation and guide them through the mitochondrial intermembrane space. The TIMM8-TIMM13 complex mediates the import of proteins such as TIMM23, SLC25A12/ARALAR1 and SLC25A13/ARALAR2, while the predominant TIMM9-TIMM10 70 kDa complex mediates the import of much more proteins.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C-Sul Probe Info |
 |
4.15 | LDD0066 | [1] | |
|
STPyne Probe Info |
 |
K27(20.00) | LDD2217 | [2] | |
|
AZ-9 Probe Info |
 |
D8(10.00) | LDD2209 | [3] | |
|
BTD Probe Info |
 |
C50(0.77); C46(0.64) | LDD2097 | [4] | |
|
DBIA Probe Info |
 |
C65(1.37); C69(1.37) | LDD2171 | [5] | |
|
ATP probe Probe Info |
 |
K49(0.00); K53(0.00) | LDD0199 | [6] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
N.A. | LDD0038 | [7] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0162 | [8] | |
|
Lodoacetamide azide Probe Info |
 |
N.A. | LDD0037 | [7] | |
|
JW-RF-010 Probe Info |
 |
N.A. | LDD0026 | [9] | |
|
TFBX Probe Info |
 |
N.A. | LDD0027 | [9] | |
|
WYneC Probe Info |
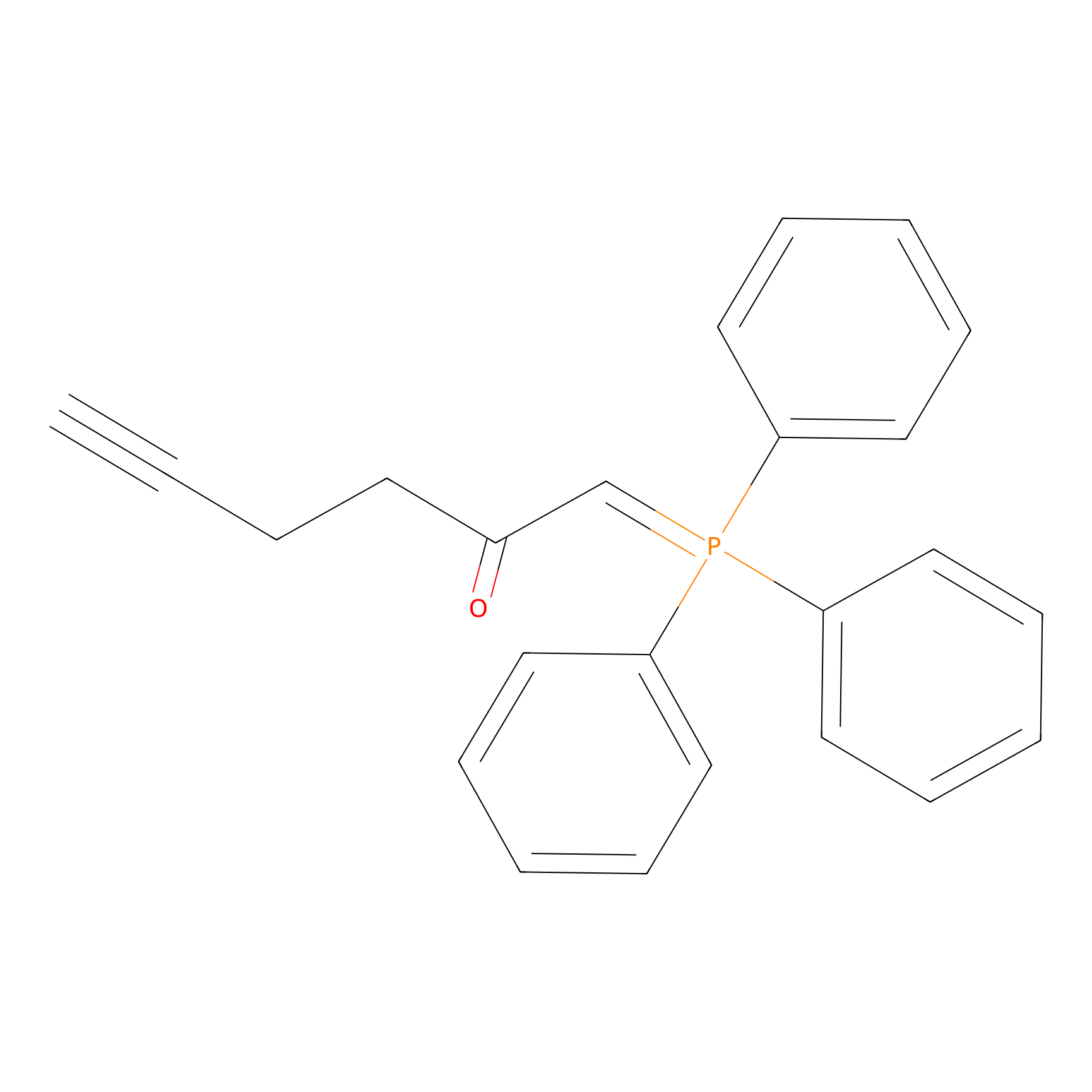 |
N.A. | LDD0014 | [10] | |
|
ENE Probe Info |
 |
N.A. | LDD0006 | [10] | |
|
IPM Probe Info |
 |
N.A. | LDD2156 | [11] | |
|
NPM Probe Info |
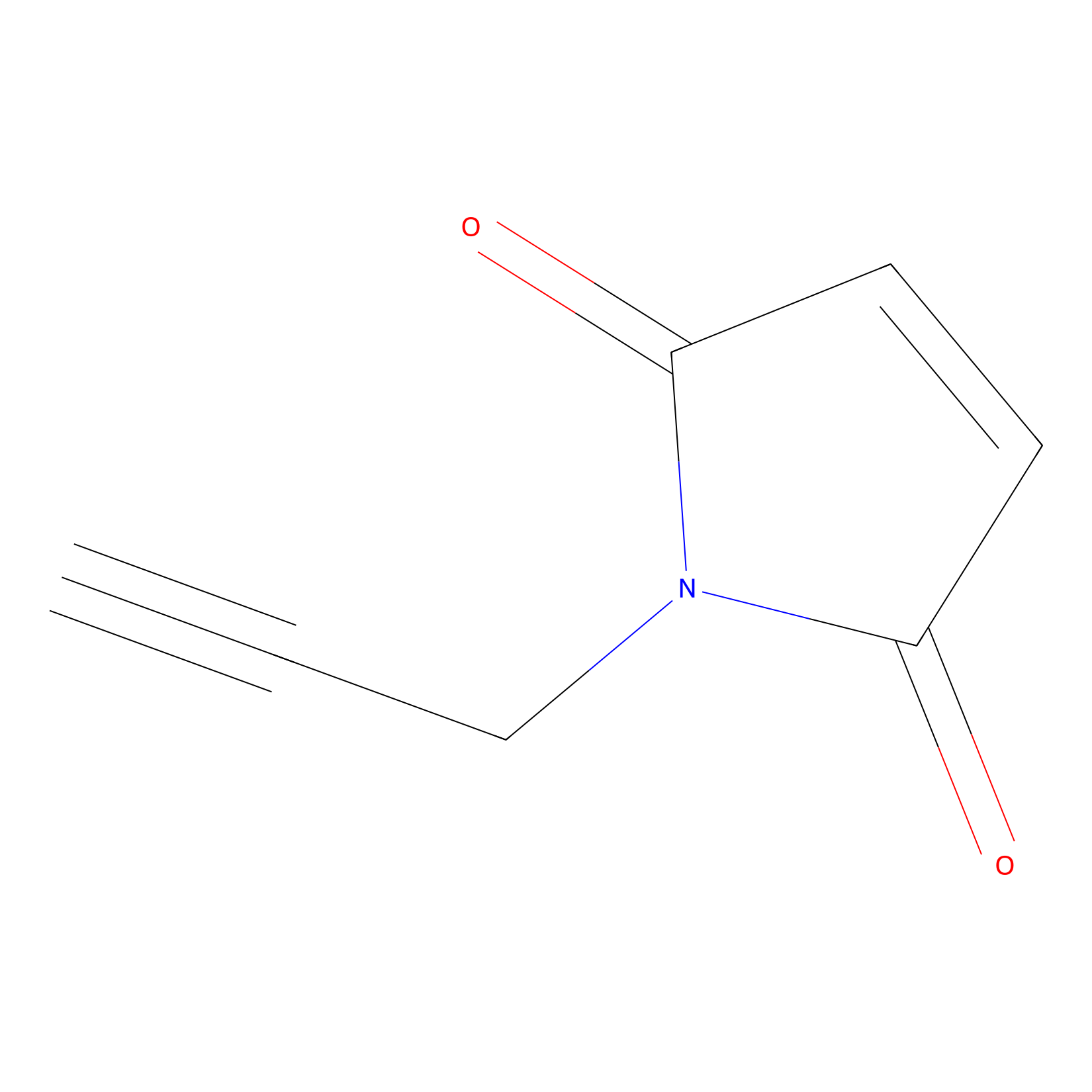 |
N.A. | LDD0016 | [10] | |
|
1c-yne Probe Info |
 |
N.A. | LDD0228 | [12] | |
|
NAIA_5 Probe Info |
 |
C50(0.00); C46(0.00) | LDD2223 | [13] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C004 Probe Info |
 |
8.06 | LDD1714 | [14] | |
|
C106 Probe Info |
 |
16.80 | LDD1793 | [14] | |
|
C218 Probe Info |
 |
11.39 | LDD1892 | [14] | |
|
C349 Probe Info |
 |
11.96 | LDD2010 | [14] | |
|
C350 Probe Info |
 |
24.25 | LDD2011 | [14] | |
|
FFF probe11 Probe Info |
 |
17.14 | LDD0471 | [15] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0520 | AKOS000195272 | MDA-MB-231 | C50(0.58) | LDD2113 | [4] |
| LDCM0020 | ARS-1620 | HCC44 | C65(1.37); C69(1.37) | LDD2171 | [5] |
| LDCM0504 | Nucleophilic fragment 15a | MDA-MB-231 | C50(0.77); C46(0.64) | LDD2097 | [4] |
| LDCM0505 | Nucleophilic fragment 15b | MDA-MB-231 | C46(1.59) | LDD2098 | [4] |
| LDCM0507 | Nucleophilic fragment 16b | MDA-MB-231 | C50(0.26) | LDD2100 | [4] |
| LDCM0508 | Nucleophilic fragment 17a | MDA-MB-231 | C50(1.10); C46(0.65) | LDD2101 | [4] |
| LDCM0511 | Nucleophilic fragment 18b | MDA-MB-231 | C50(0.34) | LDD2104 | [4] |
| LDCM0522 | Nucleophilic fragment 24a | MDA-MB-231 | C50(0.39); C46(0.87) | LDD2115 | [4] |
| LDCM0541 | Nucleophilic fragment 36 | MDA-MB-231 | C50(0.72); C46(4.37) | LDD2134 | [4] |
| LDCM0553 | Nucleophilic fragment 6b | MDA-MB-231 | C50(0.47) | LDD2147 | [4] |
| LDCM0554 | Nucleophilic fragment 7a | MDA-MB-231 | C50(0.75) | LDD2148 | [4] |
| LDCM0021 | THZ1 | HCT 116 | C65(1.37); C69(1.37) | LDD2173 | [5] |
The Interaction Atlas With This Target
References
