Details of the Target
General Information of Target
Target Site Mutations in Different Cell Lines
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
15.00 | LDD0402 | [1] | |
|
C-Sul Probe Info |
 |
5.77 | LDD0066 | [2] | |
|
STPyne Probe Info |
 |
K154(1.18) | LDD0277 | [3] | |
|
Probe 1 Probe Info |
 |
Y55(7.89) | LDD3495 | [4] | |
|
Curcusone 37 Probe Info |
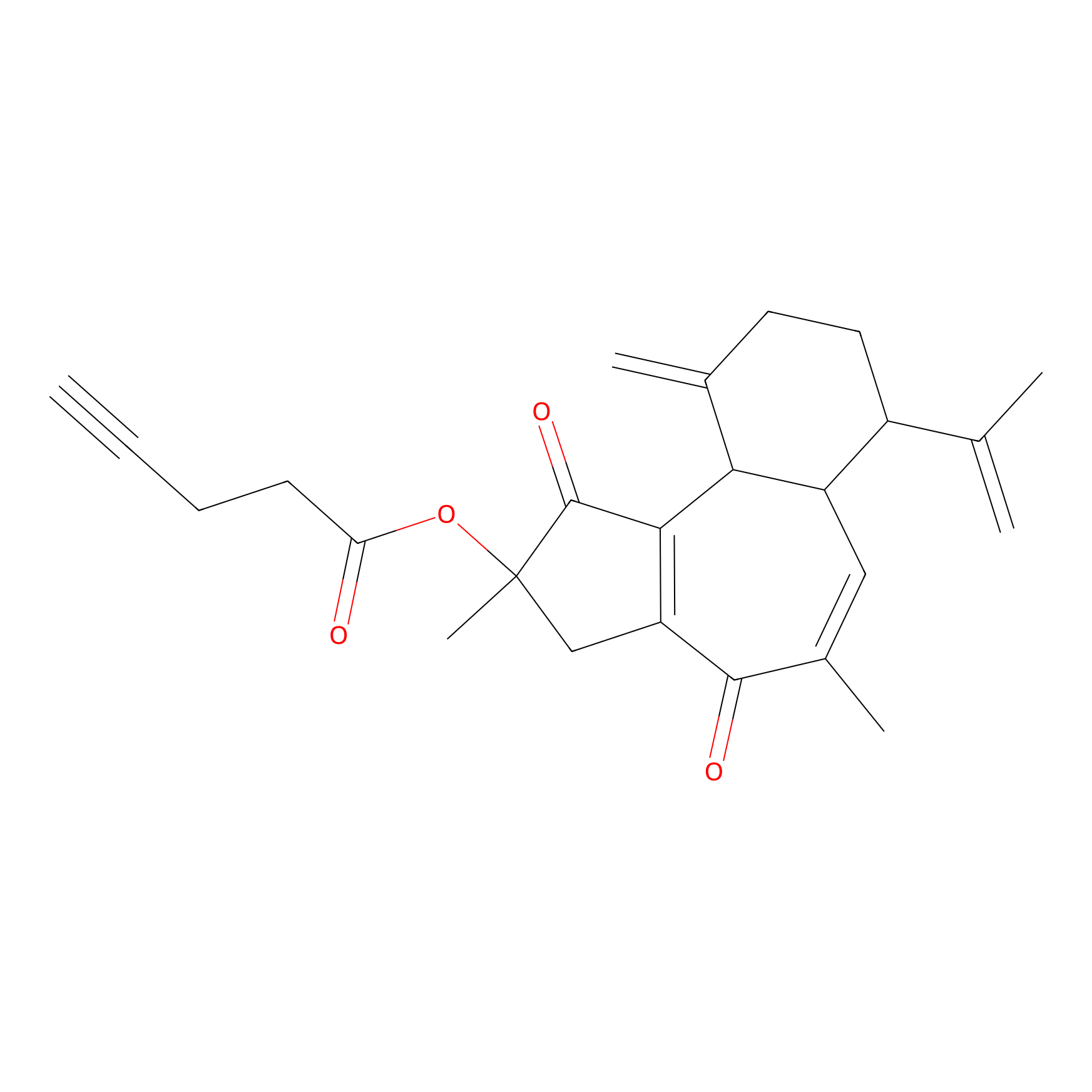 |
3.02 | LDD0188 | [5] | |
|
YY4-yne Probe Info |
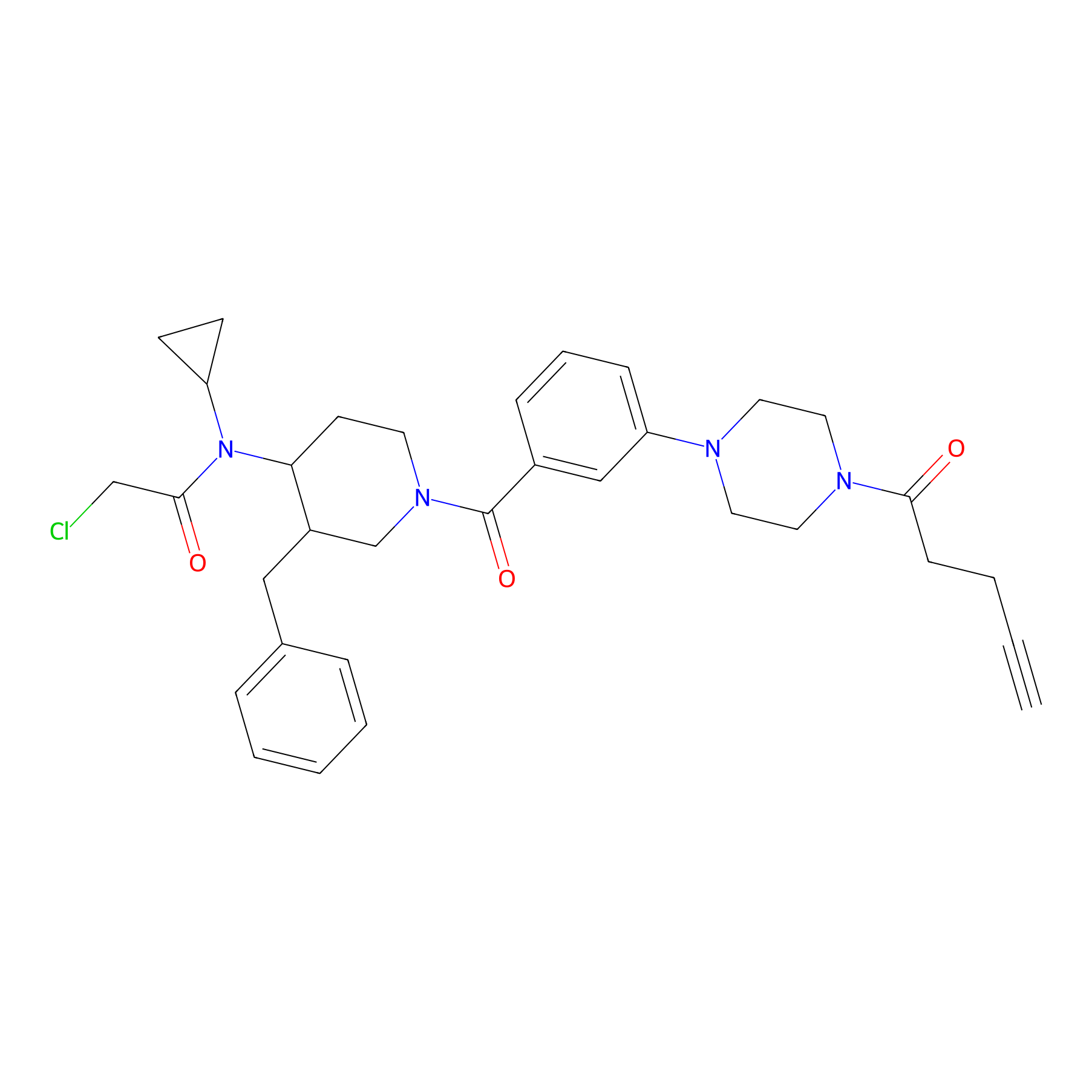 |
3.39 | LDD0400 | [6] | |
|
IA-alkyne Probe Info |
 |
C64(2.29) | LDD0372 | [7] | |
|
DBIA Probe Info |
 |
C61(4.70) | LDD0209 | [8] | |
|
IPM Probe Info |
 |
N.A. | LDD0025 | [9] | |
|
JW-RF-010 Probe Info |
 |
N.A. | LDD0026 | [9] | |
|
TFBX Probe Info |
 |
N.A. | LDD0027 | [9] | |
|
ENE Probe Info |
 |
N.A. | LDD0006 | [10] | |
|
PF-06672131 Probe Info |
 |
N.A. | LDD0017 | [11] | |
|
Phosphinate-6 Probe Info |
 |
N.A. | LDD0018 | [12] | |
|
AOyne Probe Info |
 |
14.40 | LDD0443 | [13] | |
|
NAIA_5 Probe Info |
 |
N.A. | LDD2223 | [14] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C055 Probe Info |
 |
11.63 | LDD1752 | [15] | |
|
C100 Probe Info |
 |
4.99 | LDD1789 | [15] | |
|
C178 Probe Info |
 |
22.78 | LDD1857 | [15] | |
|
C429 Probe Info |
 |
12.64 | LDD2084 | [15] | |
|
C431 Probe Info |
 |
24.25 | LDD2086 | [15] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0026 | 4SU-RNA+native RNA | HEK-293T | C64(2.29) | LDD0372 | [7] |
| LDCM0156 | Aniline | NCI-H1299 | 15.00 | LDD0403 | [1] |
| LDCM0632 | CL-Sc | Hep-G2 | C13(1.30); C13(0.89) | LDD2227 | [14] |
| LDCM0033 | Curcusone1d | MCF-7 | 3.02 | LDD0188 | [5] |
| LDCM0022 | KB02 | T cell-activated | C64(12.95) | LDD1708 | [16] |
| LDCM0023 | KB03 | Jurkat | C61(4.70) | LDD0209 | [8] |
| LDCM0024 | KB05 | NCI-N87 | C64(2.58) | LDD3364 | [17] |
| LDCM0154 | YY4 | T cell | 3.39 | LDD0400 | [6] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Transporter and channel
Transcription factor
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Nuclear factor erythroid 2-related factor 2 (NFE2L2) | BZIP family | Q16236 | |||
| Krueppel-like factor 11 (KLF11) | Sp1 C2H2-type zinc-finger protein family | O14901 | |||
Other
References
