Details of the Target
General Information of Target
| Target ID | LDTP06809 | |||||
|---|---|---|---|---|---|---|
| Target Name | Tubulin beta-8 chain (TUBB8) | |||||
| Gene Name | TUBB8 | |||||
| Gene ID | 347688 | |||||
| Synonyms |
Tubulin beta-8 chain; Tubulin beta 8 class VIII |
|||||
| 3D Structure | ||||||
| Sequence |
MREIVLTQIGQCGNQIGAKFWEVISDEHAIDSAGTYHGDSHLQLERINVYYNEASGGRYV
PRAVLVDLEPGTMDSVRSGPFGQVFRPDNFIFGQCGAGNNWAKGHYTEGAELMESVMDVV RKEAESCDCLQGFQLTHSLGGGTGSGMGTLLLSKIREEYPDRIINTFSILPSPKVSDTVV EPYNATLSVHQLIENADETFCIDNEALYDICSKTLKLPTPTYGDLNHLVSATMSGVTTCL RFPGQLNADLRKLAVNMVPFPRLHFFMPGFAPLTSRGSQQYRALTVAELTQQMFDAKNMM AACDPRHGRYLTAAAIFRGRMPMREVDEQMFNIQDKNSSYFADWLPNNVKTAVCDIPPRG LKMSATFIGNNTAIQELFKRVSEQFTAMFRRKAFLHWYTGEGMDEMEFTEAESNMNDLVS EYQQYQDATAEEEEDEEYAEEEVA |
|||||
| Target Bioclass |
Other
|
|||||
| Family |
Tubulin family
|
|||||
| Subcellular location |
Cytoplasm, cytoskeleton
|
|||||
| Function |
Tubulin is the major constituent of microtubules, a cylinder consisting of laterally associated linear protofilaments composed of alpha- and beta-tubulin heterodimers. Microtubules grow by the addition of GTP-tubulin dimers to the microtubule end, where a stabilizing cap forms. Below the cap, tubulin dimers are in GDP-bound state, owing to GTPase activity of alpha-tubulin. TUBB8 has a key role in meiotic spindle assembly and oocyte maturation.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
15.00 | LDD0402 | [1] | |
|
AZ-5 Probe Info |
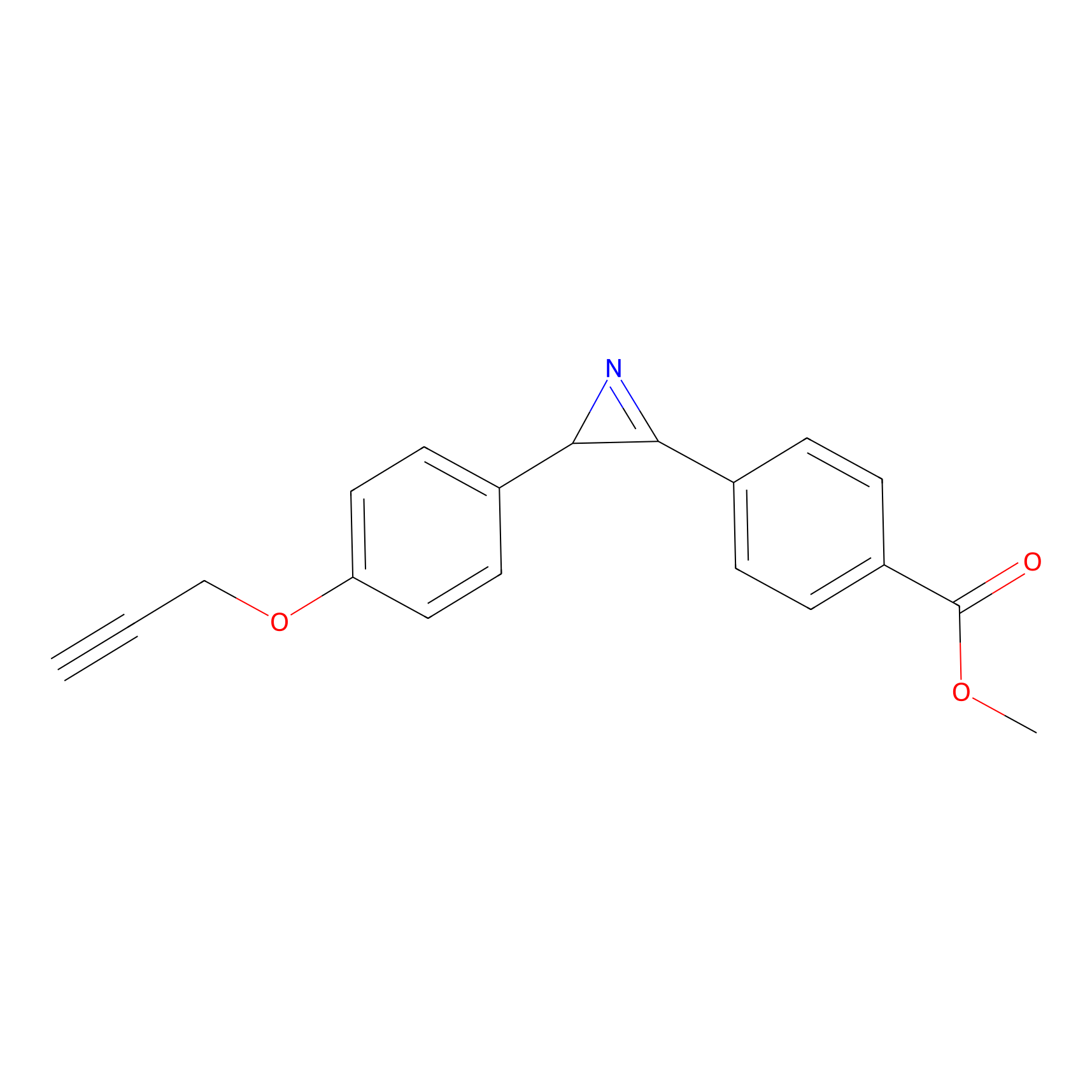 |
2.43 | LDD0394 | [2] | |
|
N1 Probe Info |
 |
100.00 | LDD0242 | [3] | |
|
DBIA Probe Info |
 |
C406(1.32) | LDD3314 | [4] | |
|
P26 Probe Info |
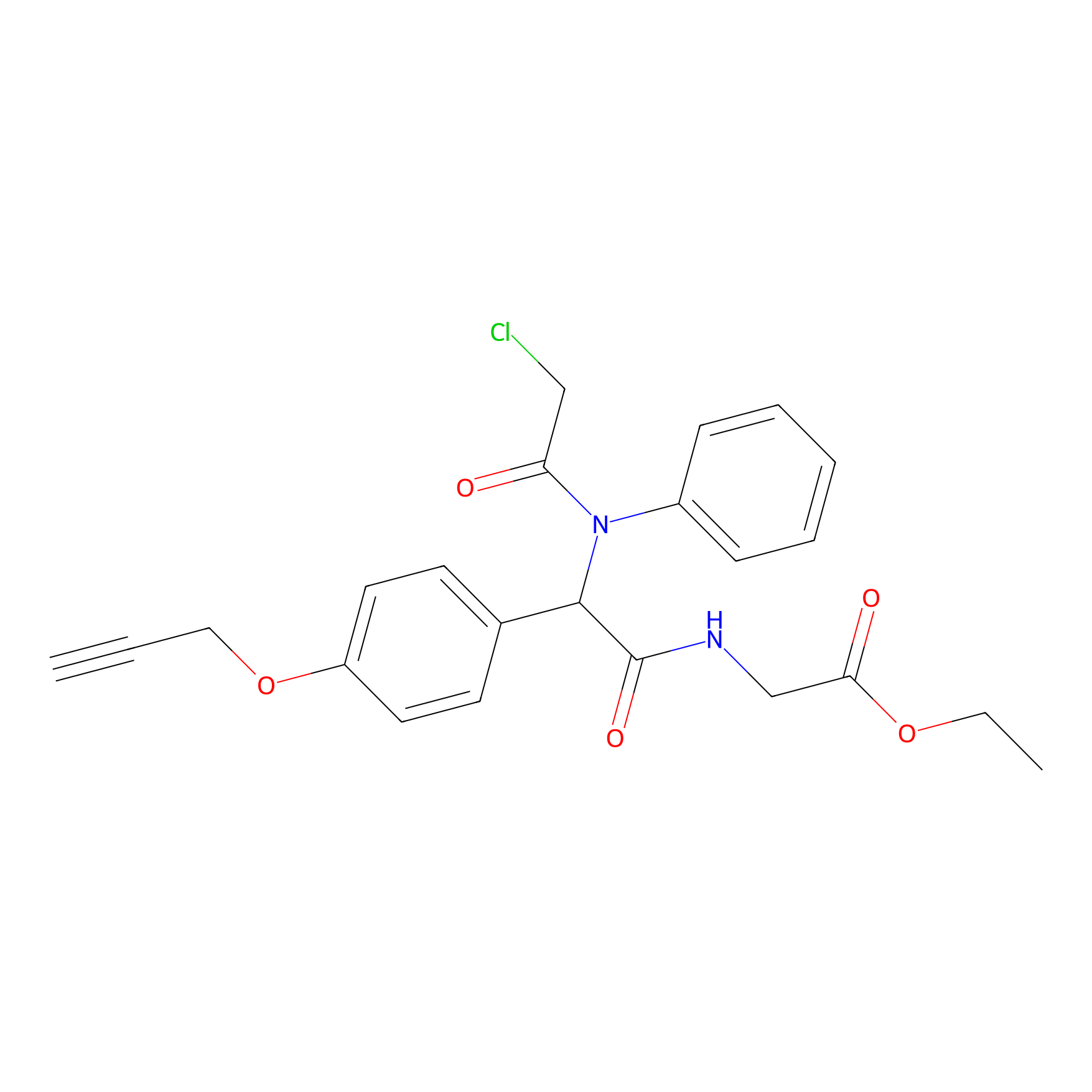 |
1.81 | LDD0409 | [5] | |
|
BTD Probe Info |
 |
C12(1.71) | LDD2096 | [6] | |
|
NAIA_5 Probe Info |
 |
C127(0.09) | LDD2227 | [7] | |
|
AHL-Pu-1 Probe Info |
 |
C129(2.47); C303(2.15) | LDD0168 | [8] | |
|
Alkyne-RA190 Probe Info |
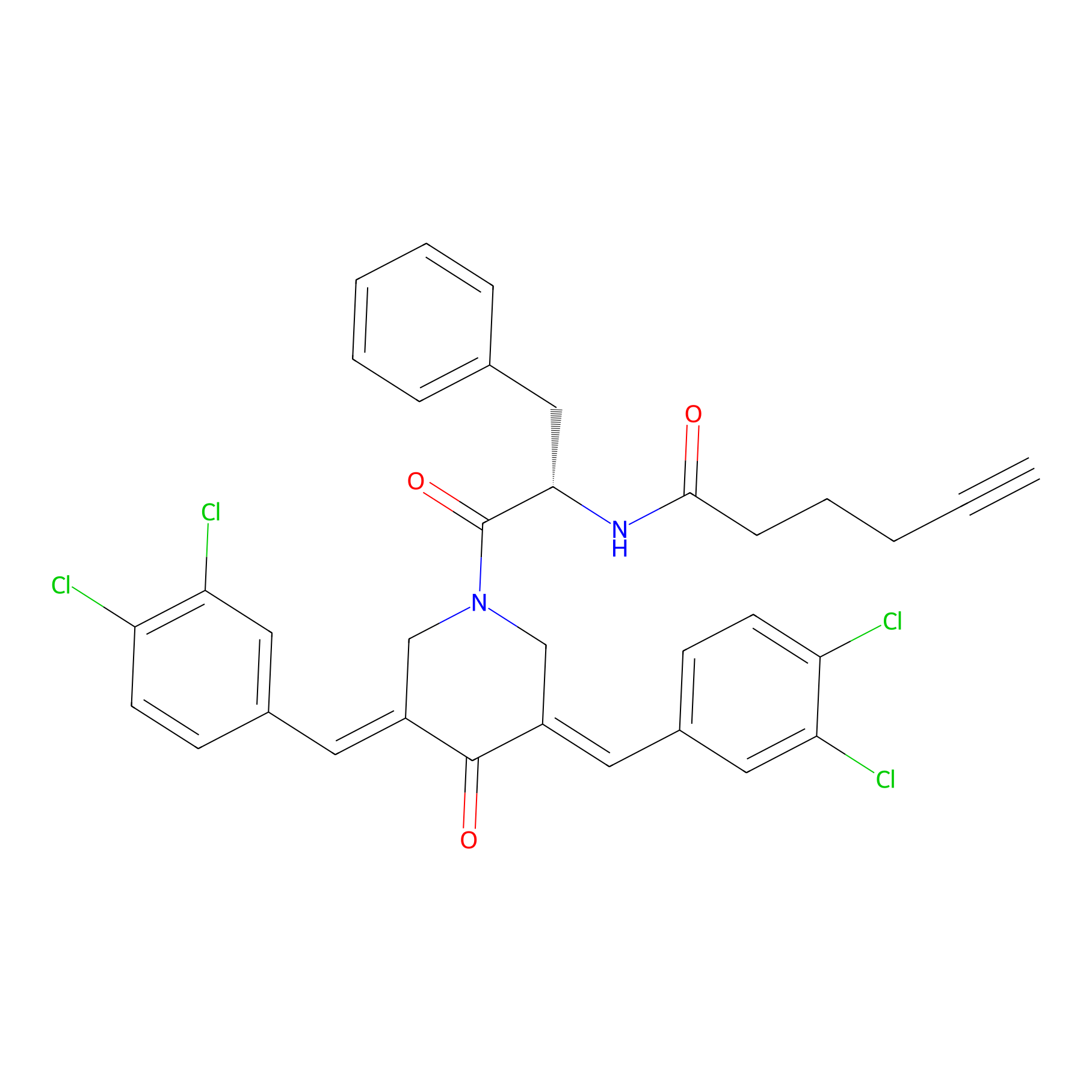 |
2.10 | LDD0299 | [9] | |
|
HHS-475 Probe Info |
 |
Y281(0.79) | LDD0264 | [10] | |
|
HHS-465 Probe Info |
 |
Y159(8.69); Y222(6.05); Y281(7.61) | LDD2237 | [11] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
N.A. | LDD0038 | [12] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0036 | [12] | |
|
IPIAA_H Probe Info |
 |
N.A. | LDD0030 | [13] | |
|
Lodoacetamide azide Probe Info |
 |
N.A. | LDD0037 | [12] | |
|
ATP probe Probe Info |
 |
K362(0.00); K252(0.00) | LDD0035 | [14] | |
|
WYneO Probe Info |
 |
C303(0.00); C354(0.00); C12(0.00) | LDD0022 | [15] | |
|
ENE Probe Info |
 |
C12(0.00); C303(0.00); C354(0.00) | LDD0006 | [15] | |
|
IPM Probe Info |
 |
C12(0.00); C303(0.00); C239(0.00); C354(0.00) | LDD0005 | [15] | |
|
TFBX Probe Info |
 |
C303(0.00); C95(0.00); C354(0.00) | LDD0148 | [16] | |
|
VSF Probe Info |
 |
N.A. | LDD0007 | [15] | |
|
1c-yne Probe Info |
 |
N.A. | LDD0228 | [17] | |
|
Acrolein Probe Info |
 |
C354(0.00); C303(0.00); C129(0.00); C127(0.00) | LDD0217 | [18] | |
|
Cinnamaldehyde Probe Info |
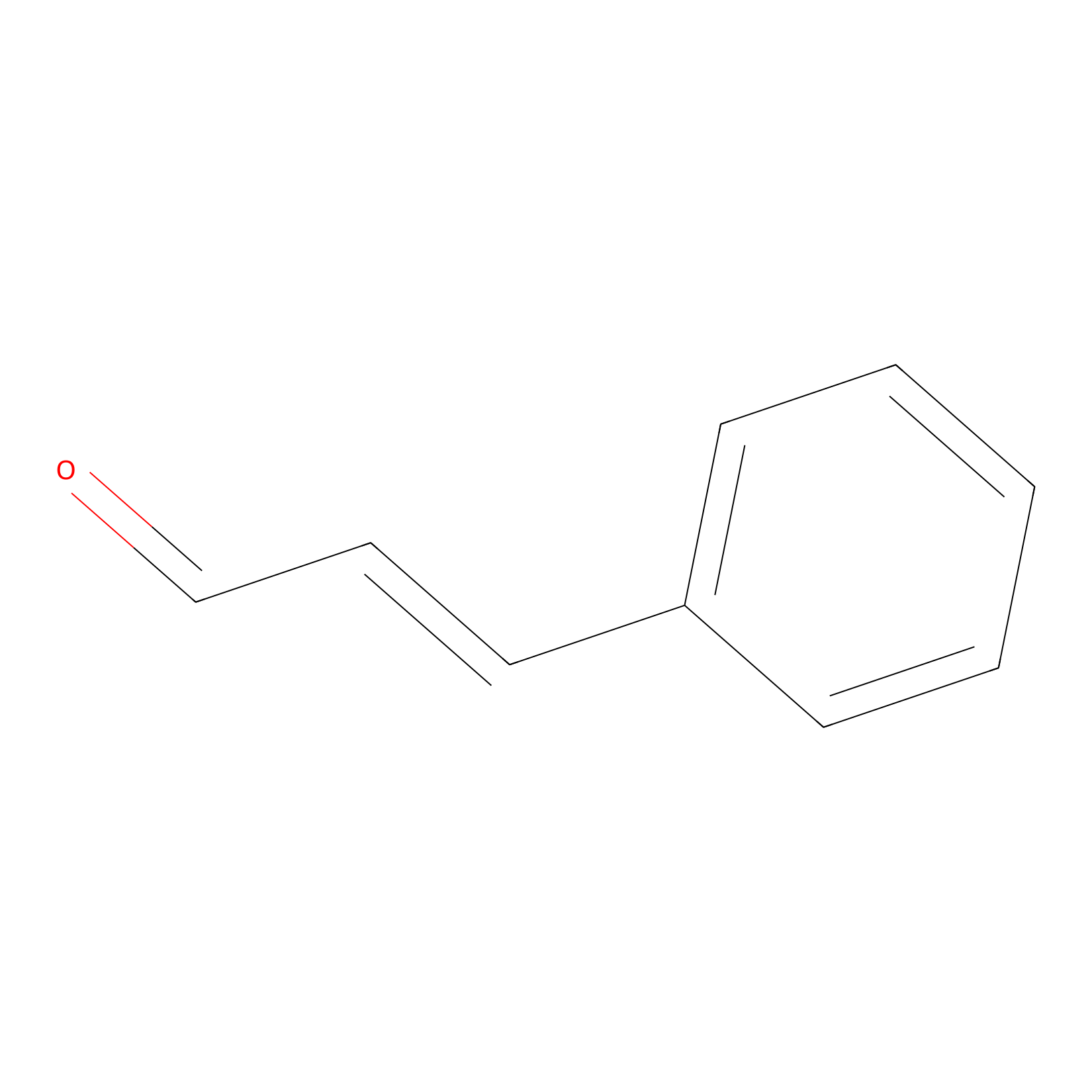 |
C354(0.00); C303(0.00) | LDD0220 | [18] | |
|
Crotonaldehyde Probe Info |
 |
C354(0.00); C303(0.00) | LDD0219 | [18] | |
|
Methacrolein Probe Info |
 |
C127(0.00); C354(0.00); C303(0.00) | LDD0218 | [18] | |
|
AOyne Probe Info |
 |
15.00 | LDD0443 | [19] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
FFF probe11 Probe Info |
 |
20.00 | LDD0472 | [20] | |
|
FFF probe3 Probe Info |
 |
8.61 | LDD0464 | [20] | |
|
FFF probe6 Probe Info |
 |
8.79 | LDD0467 | [20] | |
|
STS-2 Probe Info |
 |
N.A. | LDD0138 | [21] | |
|
DA-2 Probe Info |
 |
N.A. | LDD0070 | [22] | |
|
STS-1 Probe Info |
 |
N.A. | LDD0069 | [23] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0025 | 4SU-RNA | HEK-293T | C129(2.47); C303(2.15) | LDD0168 | [8] |
| LDCM0108 | Chloroacetamide | HeLa | H264(0.00); C354(0.00); C303(0.00); C129(0.00) | LDD0222 | [18] |
| LDCM0632 | CL-Sc | Hep-G2 | C127(0.09) | LDD2227 | [7] |
| LDCM0182 | Compound 18 | HEK-293T | 9.95 | LDD0501 | [20] |
| LDCM0191 | Compound 21 | HEK-293T | 4.44 | LDD0508 | [20] |
| LDCM0181 | Compound 41 | HEK-293T | 8.85 | LDD0502 | [20] |
| LDCM0183 | Compound 42 | HEK-293T | 5.15 | LDD0500 | [20] |
| LDCM0116 | HHS-0101 | DM93 | Y281(0.79) | LDD0264 | [10] |
| LDCM0117 | HHS-0201 | DM93 | Y281(0.80) | LDD0265 | [10] |
| LDCM0118 | HHS-0301 | DM93 | Y281(0.85) | LDD0266 | [10] |
| LDCM0119 | HHS-0401 | DM93 | Y281(0.91) | LDD0267 | [10] |
| LDCM0120 | HHS-0701 | DM93 | Y281(0.76) | LDD0268 | [10] |
| LDCM0107 | IAA | HeLa | C354(0.00); C303(0.00); H137(0.00) | LDD0221 | [18] |
| LDCM0022 | KB02 | A-375 | C406(1.70) | LDD2255 | [4] |
| LDCM0023 | KB03 | A-375 | C406(2.62) | LDD2672 | [4] |
| LDCM0024 | KB05 | IGR37 | C406(1.32) | LDD3314 | [4] |
| LDCM0109 | NEM | HeLa | H264(0.00); H137(0.00) | LDD0223 | [18] |
| LDCM0503 | Nucleophilic fragment 14b | MDA-MB-231 | C12(1.71) | LDD2096 | [6] |
| LDCM0159 | P28 | BxPC-3 | 1.81 | LDD0409 | [5] |
| LDCM0131 | RA190 | MM1.R | 2.10 | LDD0299 | [9] |
References
