Details of the Target
General Information of Target
| Target ID | LDTP04401 | |||||
|---|---|---|---|---|---|---|
| Target Name | Ras-related protein Rab-13 (RAB13) | |||||
| Gene Name | RAB13 | |||||
| Gene ID | 5872 | |||||
| Synonyms |
Ras-related protein Rab-13; Cell growth-inhibiting gene 4 protein |
|||||
| 3D Structure | ||||||
| Sequence |
MAKAYDHLFKLLLIGDSGVGKTCLIIRFAEDNFNNTYISTIGIDFKIRTVDIEGKKIKLQ
VWDTAGQERFKTITTAYYRGAMGIILVYDITDEKSFENIQNWMKSIKENASAGVERLLLG NKCDMEAKRKVQKEQADKLAREHGIRFFETSAKSSMNVDEAFSSLARDILLKSGGRRSGN GNKPPSTDLKTCDKKNTNKCSLG |
|||||
| Target Bioclass |
Enzyme
|
|||||
| Family |
Small GTPase superfamily, Rab family
|
|||||
| Subcellular location |
Cell membrane
|
|||||
| Function |
The small GTPases Rab are key regulators of intracellular membrane trafficking, from the formation of transport vesicles to their fusion with membranes. Rabs cycle between an inactive GDP-bound form and an active GTP-bound form that is able to recruit to membranes different sets of downstream effectors directly responsible for vesicle formation, movement, tethering and fusion. That Rab is involved in endocytic recycling and regulates the transport to the plasma membrane of transmembrane proteins like the tight junction protein OCLN/occludin. Thereby, it regulates the assembly and the activity of tight junctions. Moreover, it may also regulate tight junction assembly by activating the PKA signaling pathway and by reorganizing the actin cytoskeleton through the activation of the downstream effectors PRKACA and MICALL2 respectively. Through its role in tight junction assembly, may play a role in the establishment of Sertoli cell barrier. Plays also a role in angiogenesis through regulation of endothelial cells chemotaxis. Also involved in neurite outgrowth. Has also been proposed to play a role in post-Golgi membrane trafficking from the TGN to the recycling endosome. Finally, it has been involved in insulin-induced transport to the plasma membrane of the glucose transporter GLUT4 and therefore may play a role in glucose homeostasis.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
15.00 | LDD0402 | [1] | |
|
ONAyne Probe Info |
 |
N.A. | LDD0273 | [2] | |
|
OPA-S-S-alkyne Probe Info |
 |
K107(0.52); K172(4.07); K58(13.17) | LDD3494 | [3] | |
|
Alkylaryl probe 2 Probe Info |
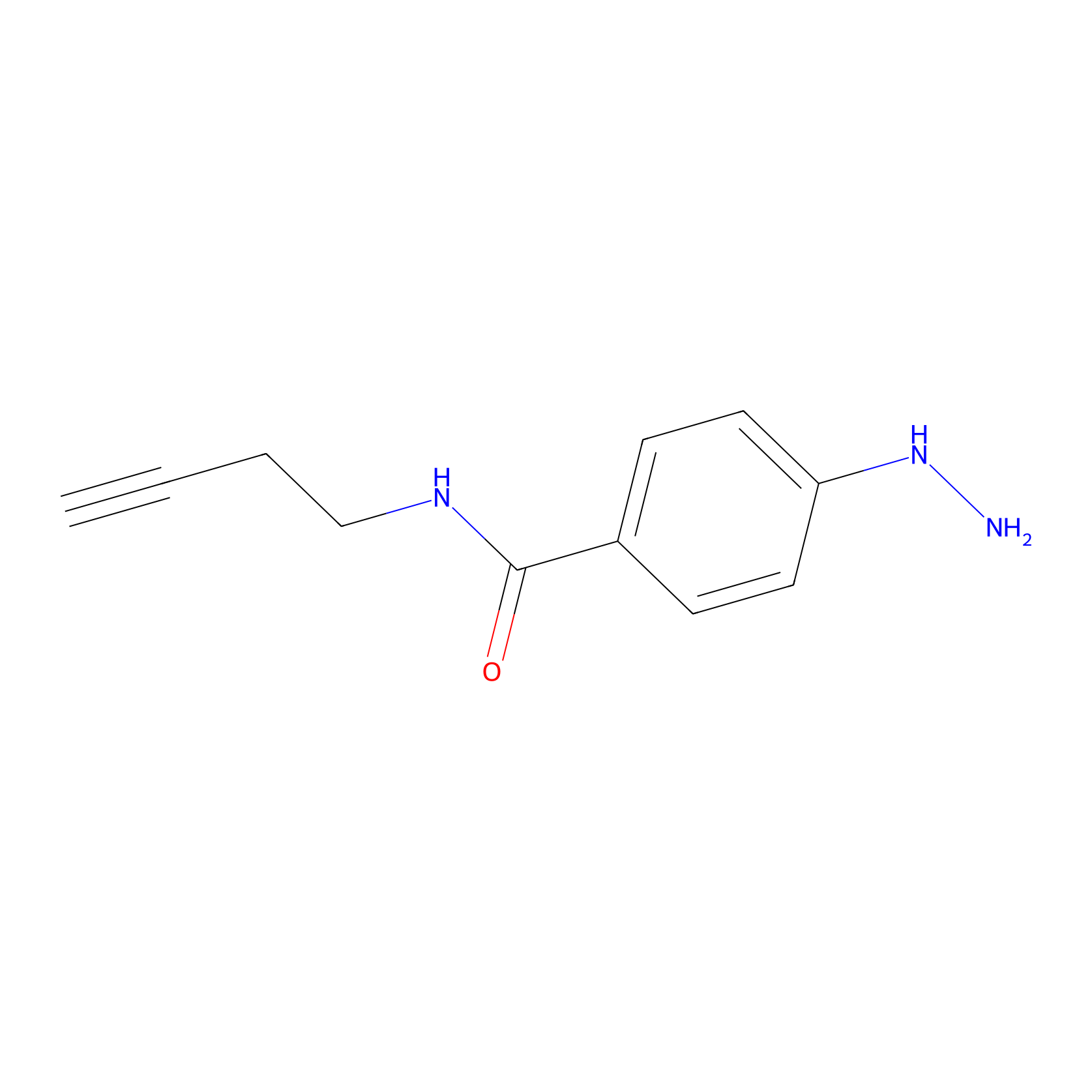 |
3.00 | LDD0390 | [4] | |
|
DBIA Probe Info |
 |
C123(4.09) | LDD3311 | [5] | |
|
JZ128-DTB Probe Info |
 |
N.A. | LDD0462 | [6] | |
|
BTD Probe Info |
 |
C123(1.21) | LDD1700 | [7] | |
|
DA-P3 Probe Info |
 |
4.18 | LDD0180 | [8] | |
|
EA-probe Probe Info |
 |
N.A. | LDD0440 | [9] | |
|
HHS-475 Probe Info |
 |
Y5(0.51) | LDD0264 | [10] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0162 | [11] | |
|
IPM Probe Info |
 |
C192(0.00); C123(0.00) | LDD0025 | [12] | |
|
NAIA_4 Probe Info |
 |
N.A. | LDD2226 | [13] | |
|
NHS Probe Info |
 |
K107(0.00); K58(0.00) | LDD0010 | [14] | |
|
PF-06672131 Probe Info |
 |
N.A. | LDD0152 | [15] | |
|
SF Probe Info |
 |
Y78(0.00); Y5(0.00); Y77(0.00) | LDD0028 | [16] | |
|
STPyne Probe Info |
 |
N.A. | LDD0009 | [14] | |
|
TFBX Probe Info |
 |
N.A. | LDD0148 | [12] | |
|
Phosphinate-6 Probe Info |
 |
N.A. | LDD0018 | [17] | |
|
1c-yne Probe Info |
 |
N.A. | LDD0228 | [18] | |
|
AOyne Probe Info |
 |
13.00 | LDD0443 | [19] | |
|
NAIA_5 Probe Info |
 |
C123(0.00); C192(0.00) | LDD2223 | [13] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0548 | 1-(4-(Benzo[d][1,3]dioxol-5-ylmethyl)piperazin-1-yl)-2-nitroethan-1-one | MDA-MB-231 | C123(0.77) | LDD2142 | [7] |
| LDCM0519 | 1-(6-methoxy-3,4-dihydroquinolin-1(2H)-yl)-2-nitroethan-1-one | MDA-MB-231 | C123(0.78) | LDD2112 | [7] |
| LDCM0502 | 1-(Cyanoacetyl)piperidine | MDA-MB-231 | C123(0.59) | LDD2095 | [7] |
| LDCM0537 | 2-Cyano-N,N-dimethylacetamide | MDA-MB-231 | C123(0.91) | LDD2130 | [7] |
| LDCM0524 | 2-Cyano-N-(2-morpholin-4-yl-ethyl)-acetamide | MDA-MB-231 | C123(0.94) | LDD2117 | [7] |
| LDCM0558 | 2-Cyano-N-phenylacetamide | MDA-MB-231 | C123(1.20) | LDD2152 | [7] |
| LDCM0510 | 3-(4-(Hydroxydiphenylmethyl)piperidin-1-yl)-3-oxopropanenitrile | MDA-MB-231 | C123(1.06) | LDD2103 | [7] |
| LDCM0539 | 3-(4-Isopropylpiperazin-1-yl)-3-oxopropanenitrile | MDA-MB-231 | C123(0.59) | LDD2132 | [7] |
| LDCM0538 | 4-(Cyanoacetyl)morpholine | MDA-MB-231 | C123(0.67) | LDD2131 | [7] |
| LDCM0520 | AKOS000195272 | MDA-MB-231 | C123(0.62) | LDD2113 | [7] |
| LDCM0156 | Aniline | NCI-H1299 | 12.07 | LDD0403 | [1] |
| LDCM0498 | BS-3668 | MDA-MB-231 | C123(0.63) | LDD2091 | [7] |
| LDCM0632 | CL-Sc | Hep-G2 | C192(0.47); C123(0.25) | LDD2227 | [13] |
| LDCM0028 | Dobutamine | HEK-293T | 4.18 | LDD0180 | [8] |
| LDCM0175 | Ethacrynic acid | HeLa | N.A. | LDD0440 | [9] |
| LDCM0116 | HHS-0101 | DM93 | Y5(0.51) | LDD0264 | [10] |
| LDCM0117 | HHS-0201 | DM93 | Y5(0.43) | LDD0265 | [10] |
| LDCM0118 | HHS-0301 | DM93 | Y5(0.85) | LDD0266 | [10] |
| LDCM0119 | HHS-0401 | DM93 | Y5(0.59) | LDD0267 | [10] |
| LDCM0120 | HHS-0701 | DM93 | Y5(1.05) | LDD0268 | [10] |
| LDCM0179 | JZ128 | PC-3 | N.A. | LDD0462 | [6] |
| LDCM0022 | KB02 | 697 | C123(2.40) | LDD2245 | [5] |
| LDCM0023 | KB03 | 697 | C123(3.69) | LDD2662 | [5] |
| LDCM0024 | KB05 | G361 | C123(4.09) | LDD3311 | [5] |
| LDCM0509 | N-(4-bromo-3,5-dimethylphenyl)-2-nitroacetamide | MDA-MB-231 | C123(1.14) | LDD2102 | [7] |
| LDCM0496 | Nucleophilic fragment 11a | MDA-MB-231 | C123(0.63) | LDD2089 | [7] |
| LDCM0497 | Nucleophilic fragment 11b | MDA-MB-231 | C123(0.92) | LDD2090 | [7] |
| LDCM0499 | Nucleophilic fragment 12b | MDA-MB-231 | C123(0.97) | LDD2092 | [7] |
| LDCM0500 | Nucleophilic fragment 13a | MDA-MB-231 | C123(1.17) | LDD2093 | [7] |
| LDCM0501 | Nucleophilic fragment 13b | MDA-MB-231 | C123(1.29) | LDD2094 | [7] |
| LDCM0505 | Nucleophilic fragment 15b | MDA-MB-231 | C123(0.72) | LDD2098 | [7] |
| LDCM0506 | Nucleophilic fragment 16a | MDA-MB-231 | C123(1.05) | LDD2099 | [7] |
| LDCM0507 | Nucleophilic fragment 16b | MDA-MB-231 | C123(0.80) | LDD2100 | [7] |
| LDCM0512 | Nucleophilic fragment 19a | MDA-MB-231 | C123(1.53) | LDD2105 | [7] |
| LDCM0513 | Nucleophilic fragment 19b | MDA-MB-231 | C123(0.84) | LDD2106 | [7] |
| LDCM0514 | Nucleophilic fragment 20a | MDA-MB-231 | C123(0.88) | LDD2107 | [7] |
| LDCM0515 | Nucleophilic fragment 20b | MDA-MB-231 | C123(0.68) | LDD2108 | [7] |
| LDCM0516 | Nucleophilic fragment 21a | MDA-MB-231 | C123(0.87) | LDD2109 | [7] |
| LDCM0518 | Nucleophilic fragment 22a | MDA-MB-231 | C123(1.05) | LDD2111 | [7] |
| LDCM0521 | Nucleophilic fragment 23b | MDA-MB-231 | C123(0.94) | LDD2114 | [7] |
| LDCM0525 | Nucleophilic fragment 25b | MDA-MB-231 | C123(0.75) | LDD2118 | [7] |
| LDCM0526 | Nucleophilic fragment 26a | MDA-MB-231 | C123(1.67) | LDD2119 | [7] |
| LDCM0527 | Nucleophilic fragment 26b | MDA-MB-231 | C123(1.04) | LDD2120 | [7] |
| LDCM0529 | Nucleophilic fragment 27b | MDA-MB-231 | C123(0.33) | LDD2122 | [7] |
| LDCM0531 | Nucleophilic fragment 28b | MDA-MB-231 | C123(1.02) | LDD2124 | [7] |
| LDCM0532 | Nucleophilic fragment 29a | MDA-MB-231 | C123(0.93) | LDD2125 | [7] |
| LDCM0533 | Nucleophilic fragment 29b | MDA-MB-231 | C123(1.20) | LDD2126 | [7] |
| LDCM0534 | Nucleophilic fragment 30a | MDA-MB-231 | C123(1.15) | LDD2127 | [7] |
| LDCM0535 | Nucleophilic fragment 30b | MDA-MB-231 | C123(1.20) | LDD2128 | [7] |
| LDCM0536 | Nucleophilic fragment 31 | MDA-MB-231 | C123(1.02) | LDD2129 | [7] |
| LDCM0540 | Nucleophilic fragment 35 | MDA-MB-231 | C123(0.66) | LDD2133 | [7] |
| LDCM0542 | Nucleophilic fragment 37 | MDA-MB-231 | C123(1.08) | LDD2135 | [7] |
| LDCM0211 | Nucleophilic fragment 3b | MDA-MB-231 | C123(1.21) | LDD1700 | [7] |
| LDCM0546 | Nucleophilic fragment 40 | MDA-MB-231 | C123(0.83) | LDD2140 | [7] |
| LDCM0547 | Nucleophilic fragment 41 | MDA-MB-231 | C123(0.62) | LDD2141 | [7] |
| LDCM0549 | Nucleophilic fragment 43 | MDA-MB-231 | C123(1.04) | LDD2143 | [7] |
| LDCM0550 | Nucleophilic fragment 5a | MDA-MB-231 | C123(2.13) | LDD2144 | [7] |
| LDCM0552 | Nucleophilic fragment 6a | MDA-MB-231 | C123(0.88) | LDD2146 | [7] |
| LDCM0553 | Nucleophilic fragment 6b | MDA-MB-231 | C123(2.58) | LDD2147 | [7] |
| LDCM0556 | Nucleophilic fragment 8a | MDA-MB-231 | C123(0.50) | LDD2150 | [7] |
| LDCM0628 | OTUB2-COV-1 | HEK-293T | C192(1.10); C123(1.07) | LDD2207 | [20] |
| LDCM0099 | Phenelzine | HEK-293T | 3.00 | LDD0390 | [4] |
The Interaction Atlas With This Target
References
