Details of the Target
General Information of Target
| Target ID | LDTP02870 | |||||
|---|---|---|---|---|---|---|
| Target Name | Y-box-binding protein 3 (YBX3) | |||||
| Gene Name | YBX3 | |||||
| Gene ID | 8531 | |||||
| Synonyms |
CSDA; DBPA; Y-box-binding protein 3; Cold shock domain-containing protein A; DNA-binding protein A; Single-strand DNA-binding protein NF-GMB |
|||||
| 3D Structure | ||||||
| Sequence |
MSEAGEATTTTTTTLPQAPTEAAAAAPQDPAPKSPVGSGAPQAAAPAPAAHVAGNPGGDA
APAATGTAAAASLATAAGSEDAEKKVLATKVLGTVKWFNVRNGYGFINRNDTKEDVFVHQ TAIKKNNPRKYLRSVGDGETVEFDVVEGEKGAEAANVTGPDGVPVEGSRYAADRRRYRRG YYGRRRGPPRNYAGEEEEEGSGSSEGFDPPATDRQFSGARNQLRRPQYRPQYRQRRFPPY HVGQTFDRRSRVLPHPNRIQAGEIGEMKDGVPEGAQLQGPVHRNPTYRPRYRSRGPPRPR PAPAVGEAEDKENQQATSGPNQPSVRRGYRRPYNYRRRPRPPNAPSQDGKEAKAGEAPTE NPAPPTQQSSAE |
|||||
| Target Bioclass |
Transcription factor
|
|||||
| Subcellular location |
Cytoplasm
|
|||||
| Function |
Binds to the GM-CSF promoter. Seems to act as a repressor. Binds also to full-length mRNA and to short RNA sequences containing the consensus site 5'-UCCAUCA-3'. May have a role in translation repression.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Target Site Mutations in Different Cell Lines
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C-Sul Probe Info |
 |
68.24 | LDD0066 | [1] | |
|
TH211 Probe Info |
 |
Y104(6.69) | LDD0257 | [2] | |
|
Acrolein Probe Info |
 |
H282(0.00); H51(0.00) | LDD0223 | [3] | |
|
5E-2FA Probe Info |
 |
H282(0.00); H119(0.00) | LDD2235 | [4] | |
|
ATP probe Probe Info |
 |
K124(0.00); K125(0.00); K268(0.00); K311(0.00) | LDD0199 | [5] | |
|
m-APA Probe Info |
 |
H282(0.00); H119(0.00); H241(0.00) | LDD2231 | [4] | |
|
1d-yne Probe Info |
 |
N.A. | LDD0358 | [6] | |
|
ATP probe Probe Info |
 |
N.A. | LDD0035 | [7] | |
|
SF Probe Info |
 |
K113(0.00); K125(0.00); K124(0.00) | LDD0028 | [8] | |
|
1c-yne Probe Info |
 |
K311(0.00); K268(0.00) | LDD0228 | [6] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C346 Probe Info |
 |
11.24 | LDD2007 | [9] | |
|
C347 Probe Info |
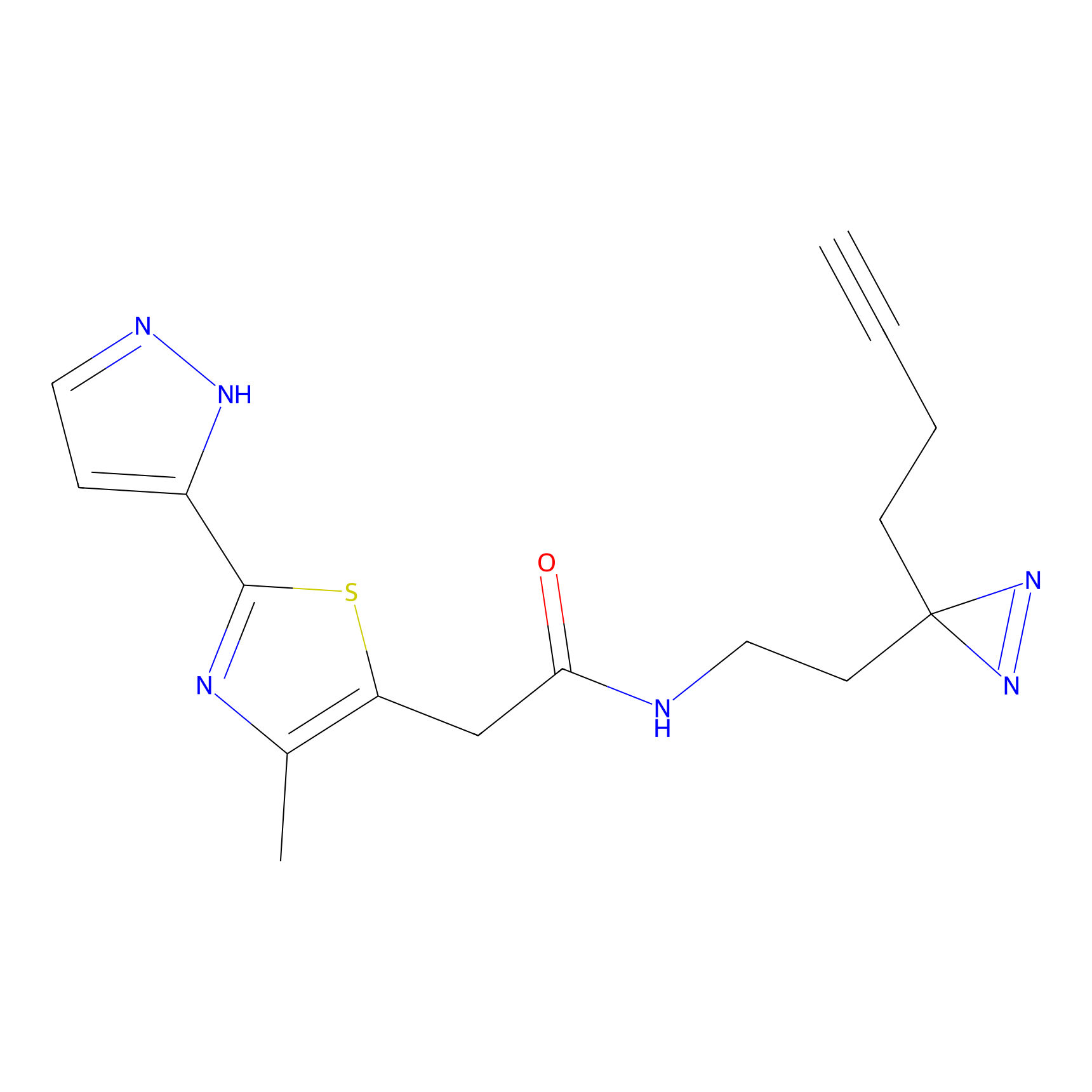 |
5.21 | LDD2008 | [9] | |
|
C364 Probe Info |
 |
20.39 | LDD2025 | [9] | |
|
FFF probe13 Probe Info |
 |
20.00 | LDD0475 | [10] | |
|
FFF probe2 Probe Info |
 |
17.89 | LDD0463 | [10] | |
|
FFF probe3 Probe Info |
 |
18.27 | LDD0464 | [10] | |
|
FFF probe4 Probe Info |
 |
20.00 | LDD0466 | [10] | |
|
FFF probe6 Probe Info |
 |
5.79 | LDD0468 | [10] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0191 | Compound 21 | HEK-293T | 8.96 | LDD0508 | [10] |
| LDCM0190 | Compound 34 | HEK-293T | 10.32 | LDD0510 | [10] |
| LDCM0192 | Compound 35 | HEK-293T | 5.55 | LDD0509 | [10] |
| LDCM0193 | Compound 36 | HEK-293T | 16.61 | LDD0511 | [10] |
| LDCM0109 | NEM | HeLa | H282(0.00); H51(0.00) | LDD0223 | [3] |
The Interaction Atlas With This Target
References
