Details of the Target
General Information of Target
| Target ID | LDTP00014 | |||||
|---|---|---|---|---|---|---|
| Target Name | Extended synaptotagmin-2 (ESYT2) | |||||
| Gene Name | ESYT2 | |||||
| Gene ID | 57488 | |||||
| Synonyms |
FAM62B; KIAA1228; Extended synaptotagmin-2; E-Syt2; Chr2Syt |
|||||
| 3D Structure | ||||||
| Sequence |
MTANRDAALSSHRHPGCAQRPRTPTFASSSQRRSAFGFDDGNFPGLGERSHAPGSRLGAR
RRAKTARGLRGHRQRGAGAGLSRPGSARAPSPPRPGGPENPGGVLSVELPGLLAQLARSF ALLLPVYALGYLGLSFSWVLLALALLAWCRRSRGLKALRLCRALALLEDEERVVRLGVRA CDLPAWVHFPDTERAEWLNKTVKHMWPFICQFIEKLFRETIEPAVRGANTHLSTFSFTKV DVGQQPLRINGVKVYTENVDKRQIILDLQISFVGNCEIDLEIKRYFCRAGVKSIQIHGTM RVILEPLIGDMPLVGALSIFFLRKPLLEINWTGLTNLLDVPGLNGLSDTIILDIISNYLV LPNRITVPLVSEVQIAQLRFPVPKGVLRIHFIEAQDLQGKDTYLKGLVKGKSDPYGIIRV GNQIFQSRVIKENLSPKWNEVYEALVYEHPGQELEIELFDEDPDKDDFLGSLMIDLIEVE KERLLDEWFTLDEVPKGKLHLRLEWLTLMPNASNLDKVLTDIKADKDQANDGLSSALLIL YLDSARNLPSGKKISSNPNPVVQMSVGHKAQESKIRYKTNEPVWEENFTFFIHNPKRQDL EVEVRDEQHQCSLGNLKVPLSQLLTSEDMTVSQRFQLSNSGPNSTIKMKIALRVLHLEKR ERPPDHQHSAQVKRPSVSKEGRKTSIKSHMSGSPGPGGSNTAPSTPVIGGSDKPGMEEKA QPPEAGPQGLHDLGRSSSSLLASPGHISVKEPTPSIASDISLPIATQELRQRLRQLENGT TLGQSPLGQIQLTIRHSSQRNKLIVVVHACRNLIAFSEDGSDPYVRMYLLPDKRRSGRRK THVSKKTLNPVFDQSFDFSVSLPEVQRRTLDVAVKNSGGFLSKDKGLLGKVLVALASEEL AKGWTQWYDLTEDGTRPQAMT |
|||||
| Target Bioclass |
Transporter and channel
|
|||||
| Family |
Extended synaptotagmin family
|
|||||
| Subcellular location |
Cell membrane
|
|||||
| Function |
Tethers the endoplasmic reticulum to the cell membrane and promotes the formation of appositions between the endoplasmic reticulum and the cell membrane. Binds glycerophospholipids in a barrel-like domain and may play a role in cellular lipid transport. Plays a role in FGF signaling via its role in the rapid internalization of FGFR1 that has been activated by FGF1 binding; this occurs most likely via the AP-2 complex. Promotes the localization of SACM1L at endoplasmic reticulum-plasma membrane contact sites (EPCS).
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
11.20 | LDD0402 | [1] | |
|
HDSF-alk Probe Info |
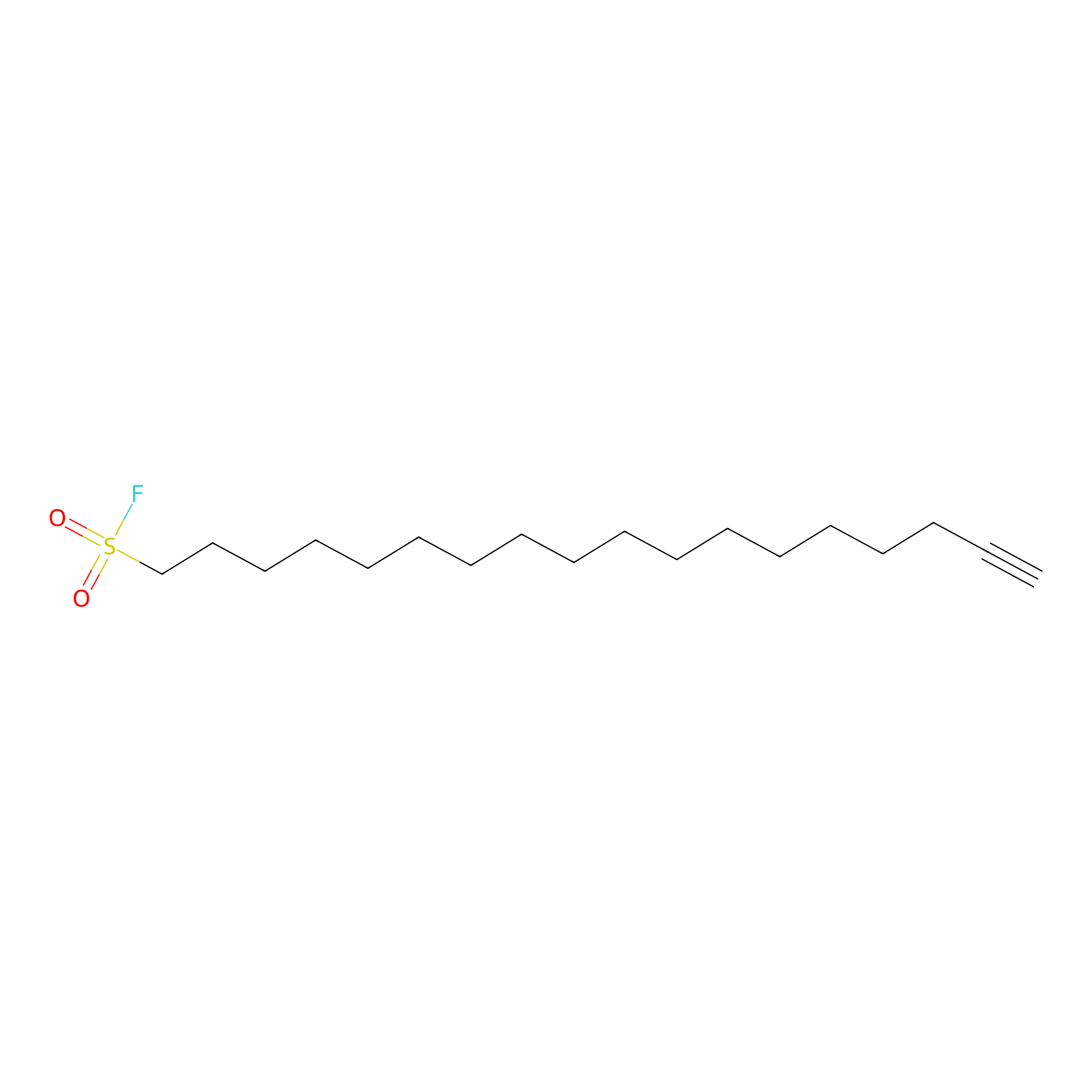 |
2.84 | LDD0197 | [2] | |
|
CHEMBL5175495 Probe Info |
 |
5.76 | LDD0196 | [3] | |
|
TG42 Probe Info |
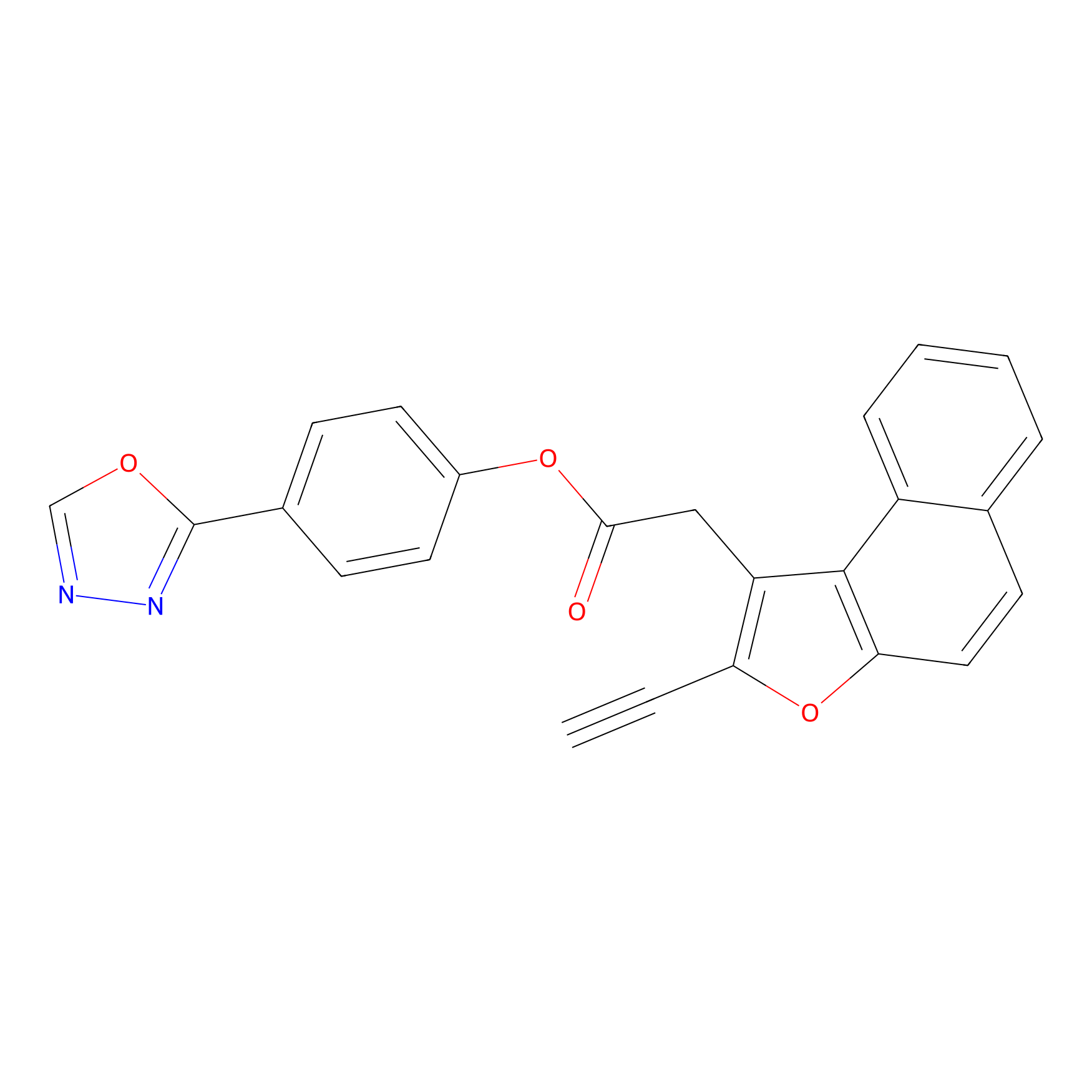 |
4.33 | LDD0326 | [4] | |
|
W1 Probe Info |
 |
16.82 | LDD0235 | [5] | |
|
STPyne Probe Info |
 |
K261(6.48); K384(10.00); K405(5.88); K411(4.79) | LDD0277 | [6] | |
|
DBIA Probe Info |
 |
C831(2.42) | LDD3310 | [7] | |
|
DA-P3 Probe Info |
 |
4.59 | LDD0182 | [8] | |
|
AHL-Pu-1 Probe Info |
 |
C181(3.71) | LDD0168 | [9] | |
|
5E-2FA Probe Info |
 |
H609(0.00); H689(0.00) | LDD2235 | [10] | |
|
AMP probe Probe Info |
 |
K875(0.00); K883(0.00) | LDD0200 | [11] | |
|
ATP probe Probe Info |
 |
K875(0.00); K883(0.00) | LDD0199 | [11] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
C210(0.00); C181(0.00); C810(0.00); C611(0.00) | LDD0038 | [12] | |
|
IA-alkyne Probe Info |
 |
C181(0.00); C210(0.00); C810(0.00) | LDD0036 | [12] | |
|
IPIAA_H Probe Info |
 |
N.A. | LDD0030 | [13] | |
|
IPIAA_L Probe Info |
 |
C181(0.00); C611(0.00) | LDD0031 | [13] | |
|
Lodoacetamide azide Probe Info |
 |
C181(0.00); C611(0.00); C210(0.00); C810(0.00) | LDD0037 | [12] | |
|
NAIA_4 Probe Info |
 |
C181(0.00); C611(0.00) | LDD2226 | [14] | |
|
Compound 10 Probe Info |
 |
N.A. | LDD2216 | [15] | |
|
Compound 11 Probe Info |
 |
N.A. | LDD2213 | [15] | |
|
IPM Probe Info |
 |
N.A. | LDD0005 | [16] | |
|
TFBX Probe Info |
 |
C611(0.00); C181(0.00) | LDD0148 | [17] | |
|
VSF Probe Info |
 |
N.A. | LDD0007 | [16] | |
|
Phosphinate-6 Probe Info |
 |
N.A. | LDD0018 | [18] | |
|
Acrolein Probe Info |
 |
C611(0.00); C181(0.00) | LDD0217 | [19] | |
|
Crotonaldehyde Probe Info |
 |
C611(0.00); H666(0.00) | LDD0219 | [19] | |
|
Methacrolein Probe Info |
 |
N.A. | LDD0218 | [19] | |
|
AOyne Probe Info |
 |
6.90 | LDD0443 | [20] | |
|
NAIA_5 Probe Info |
 |
N.A. | LDD2223 | [14] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C289 Probe Info |
 |
25.28 | LDD1959 | [21] | |
|
FFF probe11 Probe Info |
 |
20.00 | LDD0471 | [22] | |
|
FFF probe13 Probe Info |
 |
20.00 | LDD0475 | [22] | |
|
FFF probe14 Probe Info |
 |
20.00 | LDD0477 | [22] | |
|
FFF probe3 Probe Info |
 |
20.00 | LDD0464 | [22] | |
|
STS-1 Probe Info |
 |
1.00 | LDD0137 | [23] | |
|
STS-2 Probe Info |
 |
N.A. | LDD0138 | [23] | |
|
Staurosporine capture compound Probe Info |
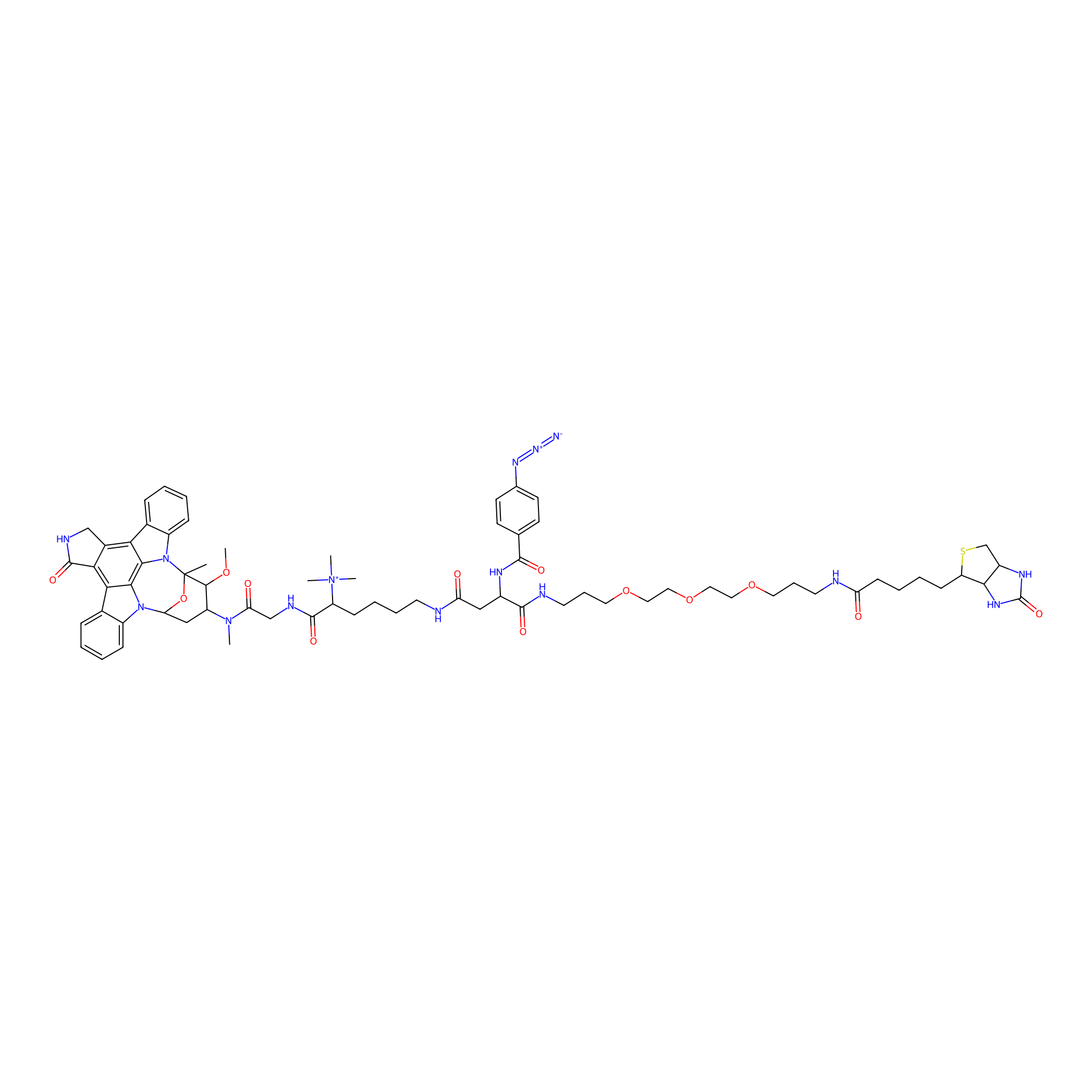 |
N.A. | LDD0083 | [24] | |
|
OEA-DA Probe Info |
 |
5.08 | LDD0046 | [25] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0025 | 4SU-RNA | HEK-293T | C181(3.71) | LDD0168 | [9] |
| LDCM0026 | 4SU-RNA+native RNA | HEK-293T | C181(3.75) | LDD0169 | [9] |
| LDCM0108 | Chloroacetamide | HeLa | C611(0.00); C181(0.00) | LDD0222 | [19] |
| LDCM0632 | CL-Sc | Hep-G2 | C181(0.79) | LDD2227 | [14] |
| LDCM0213 | Electrophilic fragment 2 | MDA-MB-231 | C181(15.38); C611(1.68) | LDD1702 | [26] |
| LDCM0625 | F8 | Ramos | C181(0.84) | LDD2187 | [27] |
| LDCM0573 | Fragment11 | Ramos | C181(0.54) | LDD2190 | [27] |
| LDCM0575 | Fragment13 | Ramos | C181(0.82) | LDD2192 | [27] |
| LDCM0576 | Fragment14 | Ramos | C181(1.17) | LDD2193 | [27] |
| LDCM0580 | Fragment21 | Ramos | C181(0.99) | LDD2195 | [27] |
| LDCM0582 | Fragment23 | Ramos | C181(0.58) | LDD2196 | [27] |
| LDCM0578 | Fragment27 | Ramos | C181(0.91) | LDD2197 | [27] |
| LDCM0586 | Fragment28 | Ramos | C181(1.66) | LDD2198 | [27] |
| LDCM0588 | Fragment30 | Ramos | C181(0.93) | LDD2199 | [27] |
| LDCM0589 | Fragment31 | Ramos | C181(1.09) | LDD2200 | [27] |
| LDCM0468 | Fragment33 | Ramos | C181(1.81) | LDD2202 | [27] |
| LDCM0596 | Fragment38 | Ramos | C181(1.02) | LDD2203 | [27] |
| LDCM0566 | Fragment4 | Ramos | C181(1.81) | LDD2184 | [27] |
| LDCM0614 | Fragment56 | Ramos | C181(0.78) | LDD2205 | [27] |
| LDCM0569 | Fragment7 | Ramos | C181(1.27) | LDD2186 | [27] |
| LDCM0107 | IAA | HeLa | H746(0.00); H188(0.00) | LDD0221 | [19] |
| LDCM0022 | KB02 | HEK-293T | C611(0.96); C181(0.92); C210(1.05) | LDD1492 | [28] |
| LDCM0023 | KB03 | HEK-293T | C611(0.97); C181(0.99); C210(0.91) | LDD1497 | [28] |
| LDCM0024 | KB05 | COLO792 | C831(2.42) | LDD3310 | [7] |
| LDCM0030 | Luteolin | HEK-293T | 4.59 | LDD0182 | [8] |
| LDCM0109 | NEM | HeLa | H746(0.00); C810(0.00); H731(0.00) | LDD0223 | [19] |
| LDCM0628 | OTUB2-COV-1 | HEK-293T | C181(1.27) | LDD2207 | [29] |
| LDCM0131 | RA190 | MM1.R | C181(1.62) | LDD0304 | [30] |
| LDCM0019 | Staurosporine | Hep-G2 | N.A. | LDD0083 | [24] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Transporter and channel
References
