Details of the Target
General Information of Target
| Target ID | LDTP13919 | |||||
|---|---|---|---|---|---|---|
| Target Name | Thyroid hormone receptor-associated protein 3 (THRAP3) | |||||
| Gene Name | THRAP3 | |||||
| Gene ID | 9967 | |||||
| Synonyms |
BCLAF2; TRAP150; Thyroid hormone receptor-associated protein 3; BCLAF1 and THRAP3 family member 2; Thyroid hormone receptor-associated protein complex 150 kDa component; Trap150 |
|||||
| 3D Structure | ||||||
| Sequence |
MAEGSGEVVAVSATGAANGLNNGAGGTSATTCNPLSRKLHKILETRLDNDKEMLEALKAL
STFFVENSLRTRRNLRGDIERKSLAINEEFVSIFKEVKEELESISEDVQAMSNCCQDMTS RLQAAKEQTQDLIVKTTKLQSESQKLEIRAQVADAFLSKFQLTSDEMSLLRGTREGPITE DFFKALGRVKQIHNDVKVLLRTNQQTAGLEIMEQMALLQETAYERLYRWAQSECRTLTQE SCDVSPVLTQAMEALQDRPVLYKYTLDEFGTARRSTVVRGFIDALTRGGPGGTPRPIEMH SHDPLRYVGDMLAWLHQATASEKEHLEALLKHVTTQGVEENIQEVVGHITEGVCRPLKVR IEQVIVAEPGAVLLYKISNLLKFYHHTISGIVGNSATALLTTIEEMHLLSKKIFFNSLSL HASKLMDKVELPPPDLGPSSALNQTLMLLREVLASHDSSVVPLDARQADFVQVLSCVLDP LLQMCTVSASNLGTADMATFMVNSLYMMKTTLALFEFTDRRLEMLQFQIEAHLDTLINEQ ASYVLTRVGLSYIYNTVQQHKPEQGSLANMPNLDSVTLKAAMVQFDRYLSAPDNLLIPQL NFLLSATVKEQIVKQSTELVCRAYGEVYAAVMNPINEYKDPENILHRSPQQVQTLLS |
|||||
| Target Bioclass |
Other
|
|||||
| Family |
BCLAF1/THRAP3 family
|
|||||
| Subcellular location |
Nucleus
|
|||||
| Function |
Involved in pre-mRNA splicing. Remains associated with spliced mRNA after splicing which probably involves interactions with the exon junction complex (EJC). Can trigger mRNA decay which seems to be independent of nonsense-mediated decay involving premature stop codons (PTC) recognition. May be involved in nuclear mRNA decay. Involved in regulation of signal-induced alternative splicing. During splicing of PTPRC/CD45 is proposed to sequester phosphorylated SFPQ from PTPRC/CD45 pre-mRNA in resting T-cells. Involved in cyclin-D1/CCND1 mRNA stability probably by acting as component of the SNARP complex which associates with both the 3'end of the CCND1 gene and its mRNA. Involved in response to DNA damage. Is excluced from DNA damage sites in a manner that parallels transcription inhibition; the function may involve the SNARP complex. Initially thought to play a role in transcriptional coactivation through its association with the TRAP complex; however, it is not regarded as a stable Mediator complex subunit. Cooperatively with HELZ2, enhances the transcriptional activation mediated by PPARG, maybe through the stabilization of the PPARG binding to DNA in presence of ligand. May play a role in the terminal stage of adipocyte differentiation. Plays a role in the positive regulation of the circadian clock. Acts as a coactivator of the CLOCK-BMAL1 heterodimer and promotes its transcriptional activator activity and binding to circadian target genes.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
11.07 | LDD0402 | [1] | |
|
P3 Probe Info |
 |
1.77 | LDD0450 | [2] | |
|
A-EBA Probe Info |
 |
3.28 | LDD0215 | [3] | |
|
CY-1 Probe Info |
 |
100.00 | LDD0243 | [4] | |
|
TH211 Probe Info |
 |
Y54(9.30) | LDD0260 | [5] | |
|
C-Sul Probe Info |
 |
2.73 | LDD0066 | [6] | |
|
ONAyne Probe Info |
 |
N.A. | LDD0273 | [7] | |
|
OPA-S-S-alkyne Probe Info |
 |
K486(1.70); K711(1.73); K346(2.43); K519(3.80) | LDD3494 | [8] | |
|
Probe 1 Probe Info |
 |
Y344(24.66); Y710(33.62) | LDD3495 | [9] | |
|
DBIA Probe Info |
 |
C476(1.54) | LDD2248 | [10] | |
|
HHS-475 Probe Info |
 |
Y710(0.49); Y880(0.74); Y228(2.26) | LDD0264 | [11] | |
|
HHS-465 Probe Info |
 |
Y228(10.00) | LDD2237 | [12] | |
|
5E-2FA Probe Info |
 |
N.A. | LDD2235 | [13] | |
|
ATP probe Probe Info |
 |
K551(0.00); K221(0.00); K697(0.00); K527(0.00) | LDD0199 | [14] | |
|
NHS Probe Info |
 |
K811(0.00); K709(0.00); K221(0.00); K688(0.00) | LDD0010 | [15] | |
|
SF Probe Info |
 |
Y228(0.00); Y710(0.00); Y873(0.00); Y344(0.00) | LDD0028 | [16] | |
|
STPyne Probe Info |
 |
N.A. | LDD0009 | [15] | |
|
Phosphinate-6 Probe Info |
 |
N.A. | LDD0018 | [17] | |
|
Ox-W18 Probe Info |
 |
W223(0.00); W471(0.00); W869(0.00) | LDD2175 | [18] | |
|
1c-yne Probe Info |
 |
K215(0.00); K221(0.00) | LDD0228 | [19] | |
|
1d-yne Probe Info |
 |
N.A. | LDD0357 | [19] | |
|
Acrolein Probe Info |
 |
H637(0.00); H883(0.00); H689(0.00); H636(0.00) | LDD0217 | [20] | |
|
Crotonaldehyde Probe Info |
 |
H613(0.00); H58(0.00); H651(0.00); H883(0.00) | LDD0219 | [20] | |
|
HHS-482 Probe Info |
 |
Y228(1.37); Y344(0.83); Y710(1.07); Y873(0.77) | LDD2239 | [12] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C187 Probe Info |
 |
18.90 | LDD1865 | [21] | |
|
C191 Probe Info |
 |
13.00 | LDD1868 | [21] | |
|
C193 Probe Info |
 |
5.58 | LDD1869 | [21] | |
|
FFF probe13 Probe Info |
 |
20.00 | LDD0475 | [22] | |
|
Diazir Probe Info |
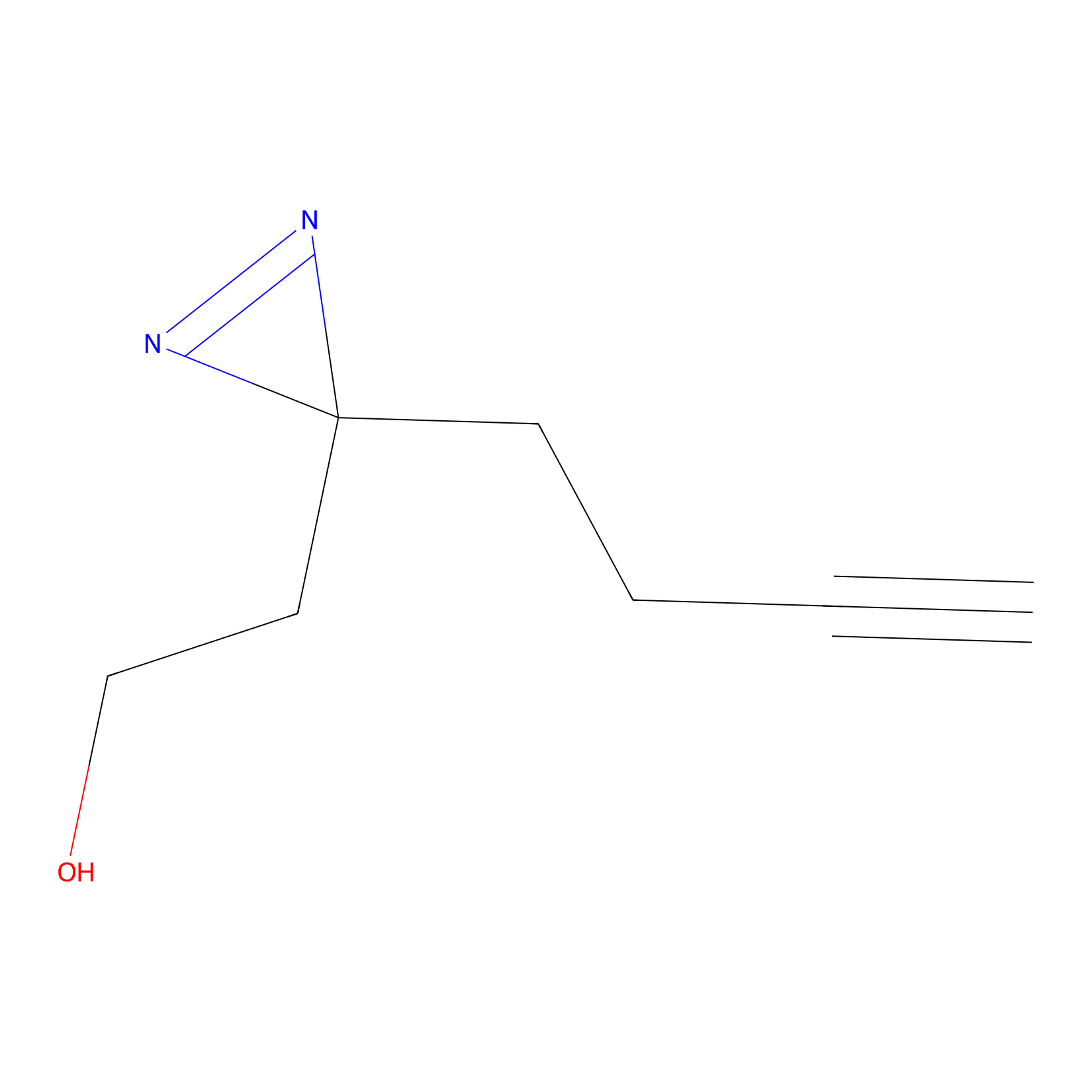 |
N.A. | LDD0011 | [15] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0156 | Aniline | NCI-H1299 | 11.39 | LDD0403 | [1] |
| LDCM0108 | Chloroacetamide | HeLa | H689(0.00); H637(0.00); H641(0.00); H613(0.00) | LDD0222 | [20] |
| LDCM0116 | HHS-0101 | DM93 | Y710(0.49); Y880(0.74); Y228(2.26) | LDD0264 | [11] |
| LDCM0117 | HHS-0201 | DM93 | Y228(1.80); Y344(20.00) | LDD0265 | [11] |
| LDCM0118 | HHS-0301 | DM93 | Y880(1.20); Y344(7.69) | LDD0266 | [11] |
| LDCM0119 | HHS-0401 | DM93 | Y880(0.51); Y344(0.67); Y228(2.28) | LDD0267 | [11] |
| LDCM0120 | HHS-0701 | DM93 | Y880(0.56); Y228(1.05) | LDD0268 | [11] |
| LDCM0107 | IAA | HeLa | H613(0.00); H637(0.00); H689(0.00); H636(0.00) | LDD0221 | [20] |
| LDCM0022 | KB02 | 8305C | C476(1.54) | LDD2248 | [10] |
| LDCM0023 | KB03 | 8505C | C476(1.58) | LDD2666 | [10] |
| LDCM0024 | KB05 | CAL-62 | C476(2.64) | LDD3123 | [10] |
| LDCM0109 | NEM | HeLa | H637(0.00); H613(0.00); H58(0.00); H883(0.00) | LDD0223 | [20] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Eukaryotic initiation factor 4A-III (EIF4A3) | DEAD box helicase family | P38919 | |||
| Casein kinase II subunit alpha (CSNK2A1) | Ser/Thr protein kinase family | P68400 | |||
Transporter and channel
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Nuclear RNA export factor 1 (NXF1) | NXF family | Q9UBU9 | |||
Transcription factor
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Splicing factor, proline- and glutamine-rich (SFPQ) | . | P23246 | |||
Other
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| SNW domain-containing protein 1 (SNW1) | SNW family | Q13573 | |||
References
