Details of the Target
General Information of Target
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
CY-1 Probe Info |
 |
100.00 | LDD0243 | [1] | |
|
CY4 Probe Info |
 |
100.00 | LDD0244 | [1] | |
|
FBP2 Probe Info |
 |
2.74 | LDD0323 | [2] | |
|
YN-1 Probe Info |
 |
100.00 | LDD0444 | [3] | |
|
m-APA Probe Info |
 |
14.85 | LDD0403 | [4] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0227 | [5] | |
|
5E-2FA Probe Info |
 |
N.A. | LDD2235 | [6] | |
|
NHS Probe Info |
 |
N.A. | LDD0010 | [7] | |
|
TER-AC Probe Info |
 |
N.A. | LDD0426 | [8] | |
|
TPP-AC Probe Info |
 |
N.A. | LDD0427 | [8] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C070 Probe Info |
 |
13.55 | LDD1766 | [9] | |
|
C095 Probe Info |
 |
5.43 | LDD1786 | [9] | |
|
C271 Probe Info |
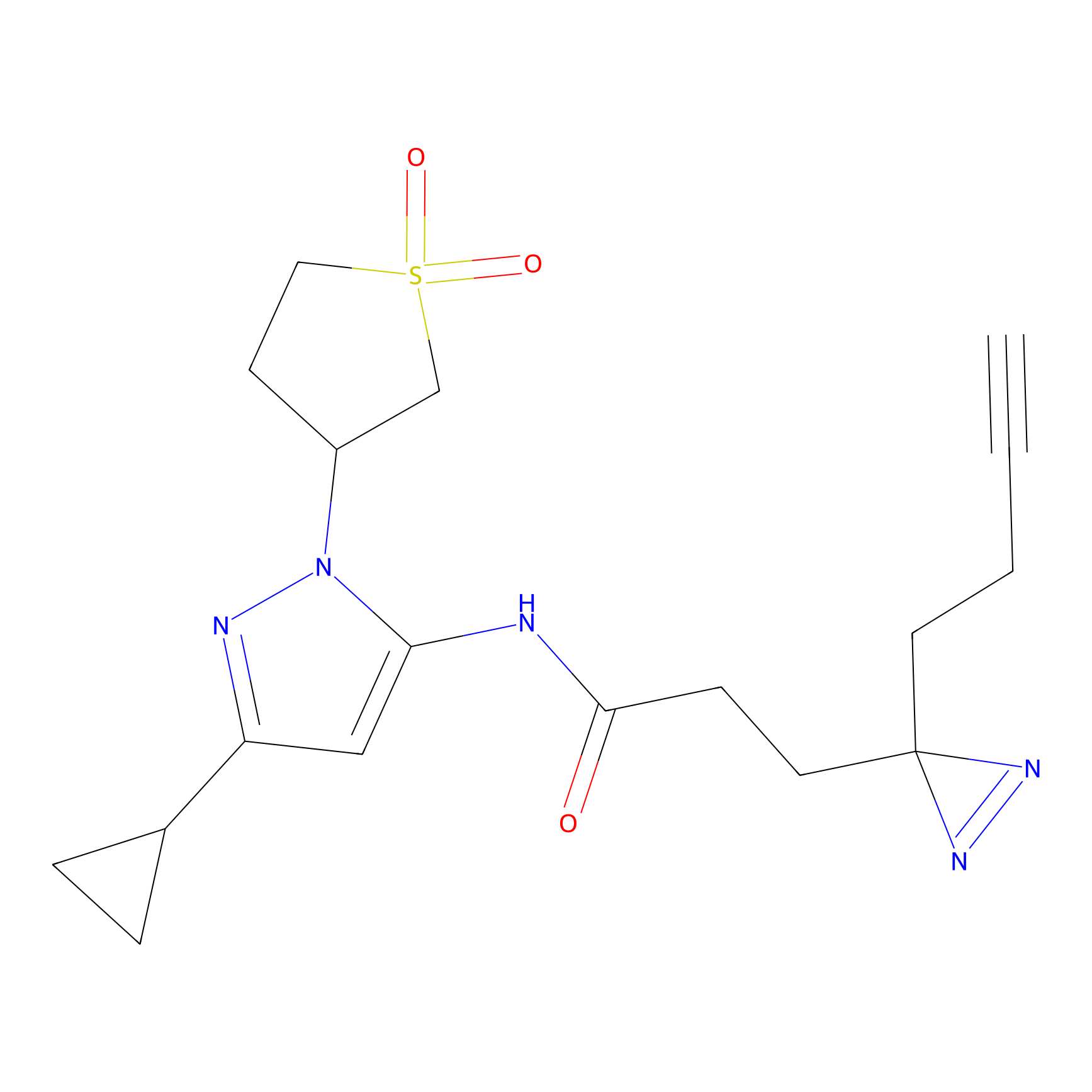 |
7.16 | LDD1941 | [9] | |
|
C278 Probe Info |
 |
59.71 | LDD1948 | [9] | |
|
C426 Probe Info |
 |
13.45 | LDD2081 | [9] | |
|
FFF probe11 Probe Info |
 |
14.28 | LDD0471 | [10] | |
|
FFF probe13 Probe Info |
 |
20.00 | LDD0475 | [10] | |
|
FFF probe14 Probe Info |
 |
13.35 | LDD0477 | [10] | |
|
FFF probe2 Probe Info |
 |
17.16 | LDD0463 | [10] | |
|
FFF probe3 Probe Info |
 |
5.63 | LDD0480 | [10] | |
Competitor(s) Related to This Target
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Other
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Complex III assembly factor LYRM7 (LYRM7) | Complex I LYR family | Q5U5X0 | |||
The Drug(s) Related To This Target
Investigative
References
