Details of the Target
General Information of Target
| Target ID | LDTP00398 | |||||
|---|---|---|---|---|---|---|
| Target Name | Insulin receptor substrate 4 (IRS4) | |||||
| Gene Name | IRS4 | |||||
| Gene ID | 8471 | |||||
| Synonyms |
Insulin receptor substrate 4; IRS-4; 160 kDa phosphotyrosine protein; py160; Phosphoprotein of 160 kDa; pp160 |
|||||
| 3D Structure | ||||||
| Sequence |
MASCSFTRDQATRRLRGAAAAAAAALAAVVTTPLLSSGTPTALIGTGSSCPGAMWLSTAT
GSRSDSESEEEDLPVGEEVCKRGYLRKQKHGHRRYFVLKLETADAPARLEYYENARKFRH SVRAAAAAAAAAASGAAIPPLIPPRRVITLYQCFSVSQRADARYRHLIALFTQDEYFAMV AENESEQESWYLLLSRLILESKRRRCGTLGAQPDGEPAALAAAAAAEPPFYKDVWQVIVK PRGLGHRKELSGVFRLCLTDEEVVFVRLNTEVASVVVQLLSIRRCGHSEQYFFLEVGRST VIGPGELWMQVDDCVVAQNMHELFLEKMRALCADEYRARCRSYSISIGAHLLTLLSARRH LGLVPLEPGGWLRRSRFEQFCHLRAIGDGEDEMLFTRRFVTPSEPVAHSRRGRLHLPRGR RSRRAVSVPASFFRRLAPSPARPRHPAEAPNNGARLSSEVSGSGSGNFGEEGNPQGKEDQ EGSGGDYMPMNNWGSGNGRGSGGGQGSNGQGSSSHSSGGNQCSGEGQGSRGGQGSNGQGS GGNQCSRDGQGTAGGHGSGGGQRPGGGHGSGGGQGPGDGHGSGGGKNSGGGKGSGSGKGS DGDGERGKSLKKRSYFGKLTQSKQQQMPPPPPPPPPPPPAGGTGGKGKSGGRFRLYFCVD RGATKECKEAKEVKDAEIPEGAARGPHRARAFDEDEDDPYVPMRPGVATPLVSSSDYMPM APQNVSASKKRHSRSPFEDSRGYMMMFPRVSPPPAPSPPKAPDTNKEDDSKDNDSESDYM FMAPGAGAIPKNPRNPQGGSSSKSWSSYFSLPNPFRSSPLGQNDNSEYVPMLPGKFLGRG LDKEVSYNWDPKDAASKPSGEGSFSKPGDGGSPSKPSDHEPPKNKAKRPNRLSFITKGYK IKPKPQKPTHEQREADSSSDYVNMDFTKRESNTPAPSTQGLPDSWGIIAEPRQSAFSNYV NVEFGVPFPNPANDLSDLLRAIPRANPLSLDSARWPLPPLPLSATGSNAIEEEGDYIEVI FNSAMTPAMALADSAIRYDAETGRIYVVDPFSECCMDISLSPSRCSEPPPVARLLQEEEQ ERRRPQSRSQSFFAAARAAVSAFPTDSLERDLSPSSAPAVASAAEPTLALSQVVAAASAL AAAPGIGAAAAAAGFDSASARWFQPVANAADAEAVRGAQDVAGGSNPGAHNPSANLARGD NQAGGAAAAAAAPEPPPRSRRVPRPPEREDSDNDDDTHVRMDFARRDNQFDSPKRGR |
|||||
| Target Bioclass |
Other
|
|||||
| Subcellular location |
Cell membrane
|
|||||
| Function |
Acts as an interface between multiple growth factor receptors possessing tyrosine kinase activity, such as insulin receptor, IGF1R and FGFR1, and a complex network of intracellular signaling molecules containing SH2 domains. Involved in the IGF1R mitogenic signaling pathway. Promotes the AKT1 signaling pathway and BAD phosphorylation during insulin stimulation without activation of RPS6KB1 or the inhibition of apoptosis. Interaction with GRB2 enhances insulin-stimulated mitogen-activated protein kinase activity. May be involved in nonreceptor tyrosine kinase signaling in myoblasts. Plays a pivotal role in the proliferation/differentiation of hepatoblastoma cell through EPHB2 activation upon IGF1 stimulation. May play a role in the signal transduction in response to insulin and to a lesser extent in response to IL4 and GH on mitogenesis. Plays a role in growth, reproduction and glucose homeostasis. May act as negative regulators of the IGF1 signaling pathway by suppressing the function of IRS1 and IRS2.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
Alkylaryl probe 1 Probe Info |
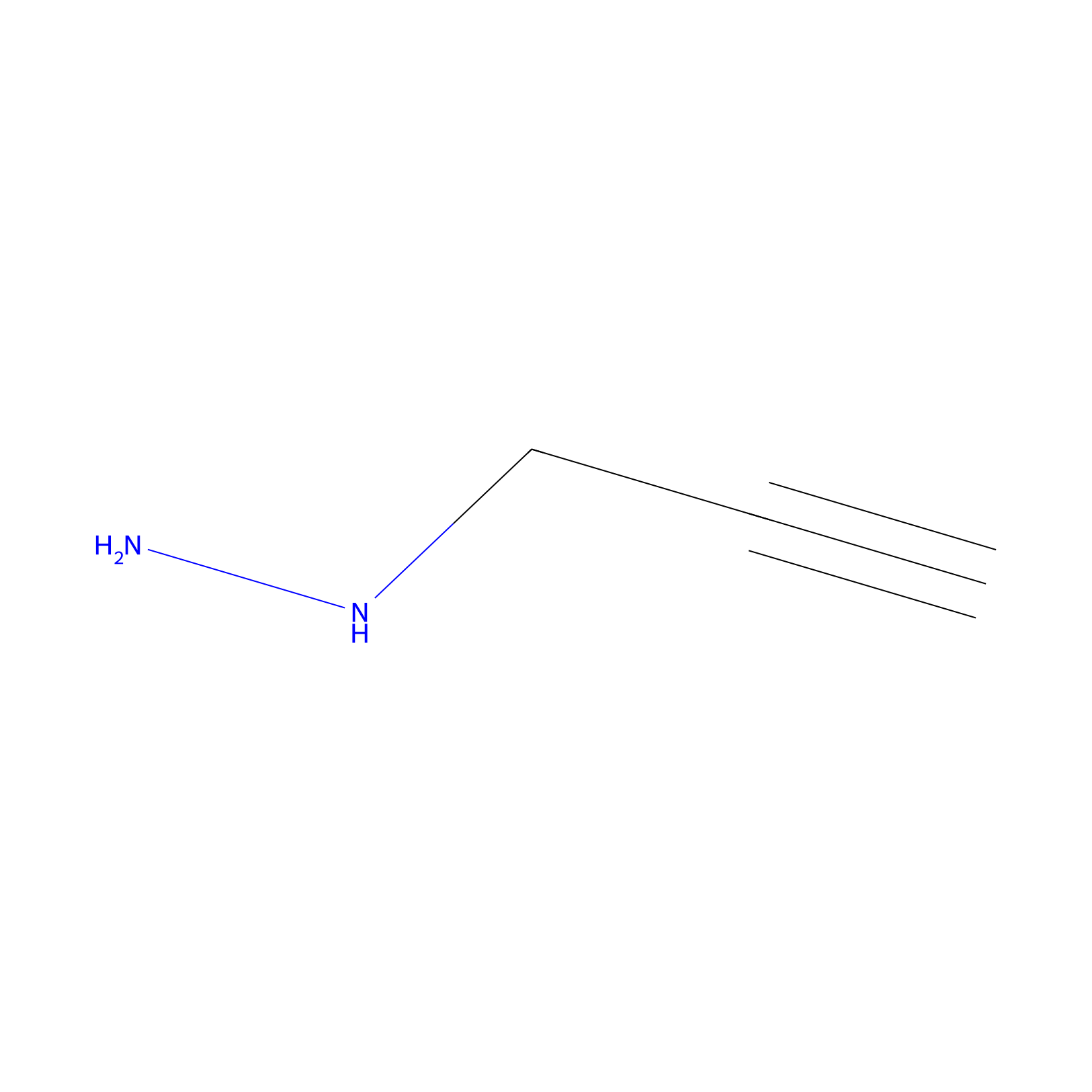 |
20.00 | LDD0385 | [1] | |
|
CHEMBL5175495 Probe Info |
 |
7.46 | LDD0196 | [2] | |
|
TH211 Probe Info |
 |
Y1038(20.00); Y743(20.00); Y899(20.00); Y1046(19.61) | LDD0257 | [3] | |
|
TH214 Probe Info |
 |
Y743(20.00); Y808(20.00) | LDD0258 | [3] | |
|
TH216 Probe Info |
 |
Y656(20.00); Y1038(19.59); Y828(18.66); Y112(10.94) | LDD0259 | [3] | |
|
ONAyne Probe Info |
 |
N.A. | LDD0273 | [4] | |
|
AHL-Pu-1 Probe Info |
 |
C285(3.06) | LDD0168 | [5] | |
|
HHS-465 Probe Info |
 |
Y808(9.91); Y828(9.90) | LDD2237 | [6] | |
|
DBIA Probe Info |
 |
C1054(6.91); C332(2.58); C1065(3.47); C1055(3.60) | LDD0080 | [7] | |
|
ATP probe Probe Info |
 |
K618(0.00); K240(0.00); K99(0.00) | LDD0199 | [8] | |
|
IA-alkyne Probe Info |
 |
C332(0.00); C1054(0.00); C545(0.00) | LDD0032 | [9] | |
|
DA-P3 Probe Info |
 |
C522(0.00); C545(0.00) | LDD0178 | [10] | |
|
ENE Probe Info |
 |
C545(0.00); C1065(0.00); C522(0.00) | LDD0006 | [11] | |
|
IPM Probe Info |
 |
C332(0.00); C1054(0.00); C1065(0.00); C285(0.00) | LDD0005 | [11] | |
|
NHS Probe Info |
 |
N.A. | LDD0010 | [11] | |
|
NPM Probe Info |
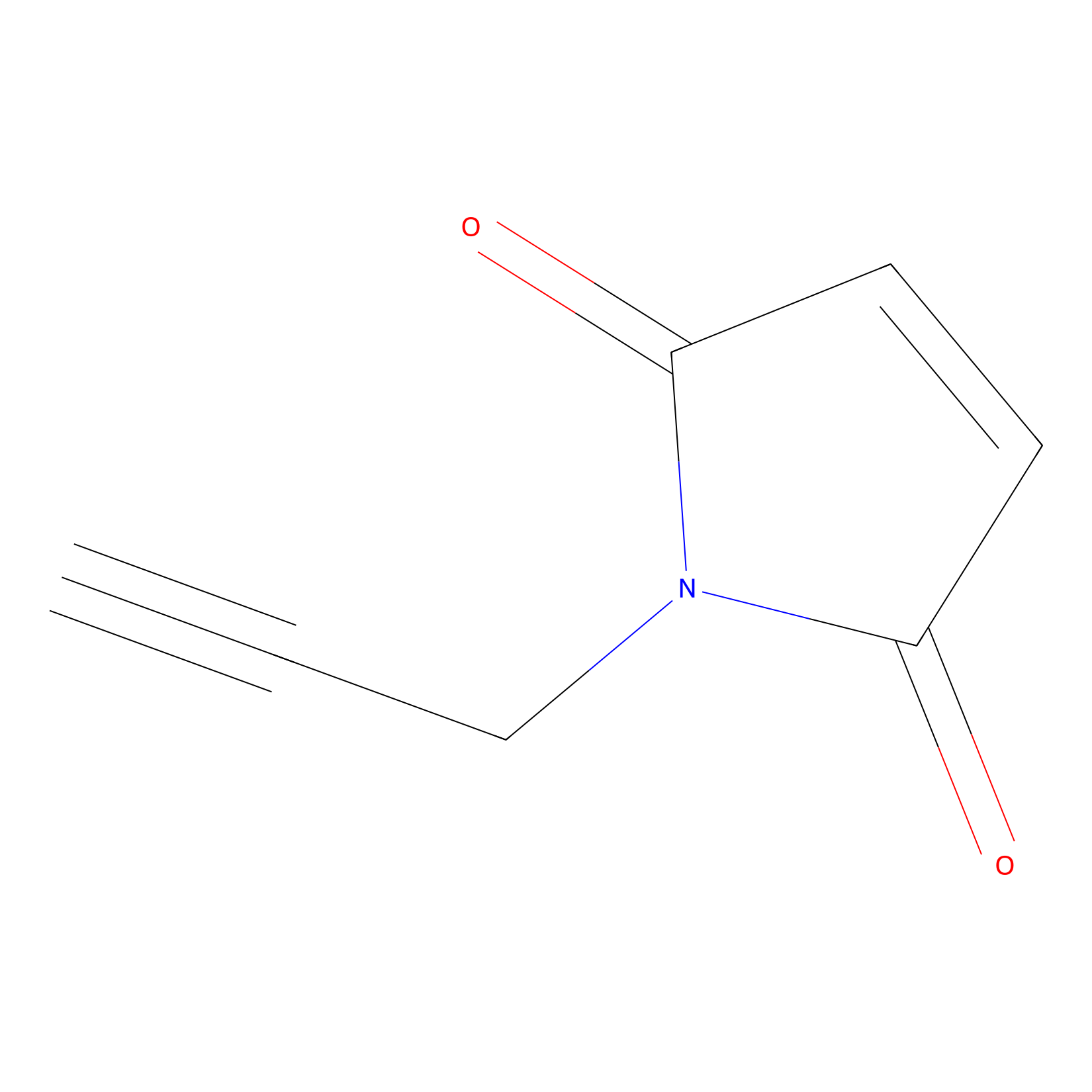 |
N.A. | LDD0016 | [11] | |
|
PPMS Probe Info |
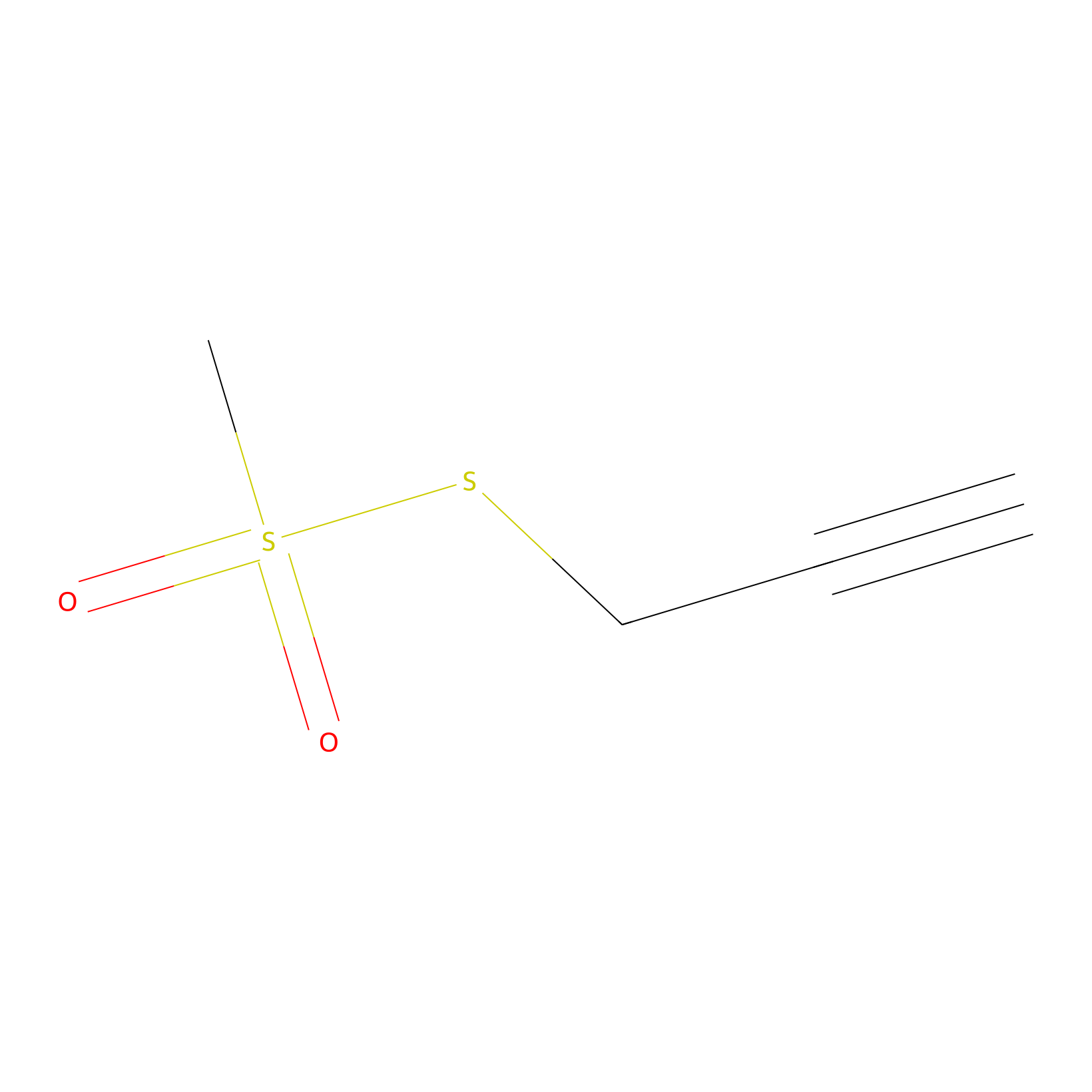 |
C545(0.00); C522(0.00) | LDD0008 | [11] | |
|
SF Probe Info |
 |
Y1038(0.00); Y111(0.00) | LDD0028 | [12] | |
|
VSF Probe Info |
 |
C206(0.00); C1054(0.00); C522(0.00); C1055(0.00) | LDD0007 | [11] | |
|
Phosphinate-6 Probe Info |
 |
C667(0.00); C522(0.00); C1065(0.00); C658(0.00) | LDD0018 | [13] | |
|
1c-yne Probe Info |
 |
K843(0.00); K803(0.00) | LDD0228 | [14] | |
|
W1 Probe Info |
 |
E113(0.00); Y112(0.00); N114(0.00) | LDD0236 | [15] | |
|
AOyne Probe Info |
 |
9.90 | LDD0443 | [16] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
FFF probe11 Probe Info |
 |
14.66 | LDD0471 | [17] | |
|
FFF probe13 Probe Info |
 |
17.36 | LDD0475 | [17] | |
|
FFF probe14 Probe Info |
 |
12.50 | LDD0477 | [17] | |
|
FFF probe2 Probe Info |
 |
9.46 | LDD0463 | [17] | |
|
FFF probe3 Probe Info |
 |
17.65 | LDD0464 | [17] | |
|
OEA-DA Probe Info |
 |
3.33 | LDD0046 | [18] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0025 | 4SU-RNA | HEK-293T | C285(3.06) | LDD0168 | [5] |
| LDCM0026 | 4SU-RNA+native RNA | HEK-293T | C153(2.76); C1055(2.10); C658(2.13); C545(2.03) | LDD0169 | [5] |
| LDCM0214 | AC1 | HEK-293T | C545(1.12); C1065(0.95); C332(1.09); C381(1.02) | LDD1507 | [19] |
| LDCM0215 | AC10 | HEK-293T | C545(1.01); C332(0.97); C381(0.97); C285(0.94) | LDD1508 | [19] |
| LDCM0226 | AC11 | HEK-293T | C545(1.08); C1065(1.03); C332(0.94); C381(1.08) | LDD1509 | [19] |
| LDCM0237 | AC12 | HEK-293T | C545(0.95); C1065(1.01); C332(0.91); C381(1.06) | LDD1510 | [19] |
| LDCM0259 | AC14 | HEK-293T | C545(1.00); C1065(1.09); C332(0.94); C381(1.05) | LDD1512 | [19] |
| LDCM0270 | AC15 | HEK-293T | C545(0.95); C1065(0.98); C332(0.92); C381(1.10) | LDD1513 | [19] |
| LDCM0276 | AC17 | HEK-293T | C545(1.03); C1065(1.05); C332(0.92); C381(1.01) | LDD1515 | [19] |
| LDCM0277 | AC18 | HEK-293T | C545(1.05); C332(1.01); C381(0.96); C285(0.97) | LDD1516 | [19] |
| LDCM0278 | AC19 | HEK-293T | C545(2.25); C1065(1.35); C332(1.07); C381(1.10) | LDD1517 | [19] |
| LDCM0279 | AC2 | HEK-293T | C545(1.06); C332(1.05); C381(0.98); C285(0.94) | LDD1518 | [19] |
| LDCM0280 | AC20 | HEK-293T | C545(1.02); C1065(0.97); C332(0.94); C381(1.02) | LDD1519 | [19] |
| LDCM0281 | AC21 | HEK-293T | C545(0.99); C1065(1.02); C332(0.93); C381(0.96) | LDD1520 | [19] |
| LDCM0282 | AC22 | HEK-293T | C545(0.95); C1065(1.02); C332(1.02); C381(0.98) | LDD1521 | [19] |
| LDCM0283 | AC23 | HEK-293T | C545(0.97); C1065(0.91); C332(0.96); C381(1.02) | LDD1522 | [19] |
| LDCM0284 | AC24 | HEK-293T | C545(0.98); C1065(0.98); C332(1.03); C381(0.97) | LDD1523 | [19] |
| LDCM0285 | AC25 | HEK-293T | C545(1.04); C1065(0.83); C332(0.93); C381(1.10) | LDD1524 | [19] |
| LDCM0286 | AC26 | HEK-293T | C545(1.02); C332(1.00); C381(1.05); C285(0.95) | LDD1525 | [19] |
| LDCM0287 | AC27 | HEK-293T | C545(1.14); C1065(0.99); C332(0.94); C381(1.13) | LDD1526 | [19] |
| LDCM0288 | AC28 | HEK-293T | C545(0.99); C1065(0.99); C332(0.94); C381(0.98) | LDD1527 | [19] |
| LDCM0289 | AC29 | HEK-293T | C545(0.98); C1065(1.12); C332(0.93); C381(0.97) | LDD1528 | [19] |
| LDCM0290 | AC3 | HEK-293T | C545(1.08); C1065(1.07); C332(0.96); C381(1.02) | LDD1529 | [19] |
| LDCM0291 | AC30 | HEK-293T | C545(0.98); C1065(1.02); C332(0.98); C381(1.03) | LDD1530 | [19] |
| LDCM0292 | AC31 | HEK-293T | C545(0.89); C1065(0.87); C332(0.94); C381(1.01) | LDD1531 | [19] |
| LDCM0293 | AC32 | HEK-293T | C545(0.85); C1065(0.95); C332(0.93); C381(1.02) | LDD1532 | [19] |
| LDCM0294 | AC33 | HEK-293T | C545(1.08); C1065(0.92); C332(0.97); C381(1.09) | LDD1533 | [19] |
| LDCM0295 | AC34 | HEK-293T | C545(1.06); C332(0.97); C381(0.99); C285(0.96) | LDD1534 | [19] |
| LDCM0296 | AC35 | HEK-293T | C545(1.09); C1065(0.96); C332(0.92); C381(1.00) | LDD1535 | [19] |
| LDCM0297 | AC36 | HEK-293T | C545(0.97); C1065(1.01); C332(0.82); C381(0.97) | LDD1536 | [19] |
| LDCM0298 | AC37 | HEK-293T | C545(0.97); C1065(1.05); C332(0.88); C381(0.89) | LDD1537 | [19] |
| LDCM0299 | AC38 | HEK-293T | C545(0.99); C1065(0.96); C332(0.93); C381(1.06) | LDD1538 | [19] |
| LDCM0300 | AC39 | HEK-293T | C545(1.03); C1065(1.54); C332(0.91); C381(1.04) | LDD1539 | [19] |
| LDCM0301 | AC4 | HEK-293T | C545(1.04); C1065(1.00); C332(0.85); C381(0.97) | LDD1540 | [19] |
| LDCM0302 | AC40 | HEK-293T | C545(0.84); C1065(0.98); C332(0.89); C381(1.06) | LDD1541 | [19] |
| LDCM0303 | AC41 | HEK-293T | C545(1.14); C1065(0.93); C332(0.96); C381(1.14) | LDD1542 | [19] |
| LDCM0304 | AC42 | HEK-293T | C545(1.14); C332(0.97); C381(1.02); C285(0.85) | LDD1543 | [19] |
| LDCM0305 | AC43 | HEK-293T | C545(0.98); C1065(0.93); C332(0.93); C381(1.04) | LDD1544 | [19] |
| LDCM0306 | AC44 | HEK-293T | C545(1.02); C1065(1.06); C332(0.91); C381(1.13) | LDD1545 | [19] |
| LDCM0307 | AC45 | HEK-293T | C545(1.03); C1065(1.05); C332(0.88); C381(0.90) | LDD1546 | [19] |
| LDCM0308 | AC46 | HEK-293T | C545(1.05); C1065(1.05); C332(0.92); C381(1.00) | LDD1547 | [19] |
| LDCM0309 | AC47 | HEK-293T | C545(0.97); C1065(0.91); C332(0.86); C381(1.03) | LDD1548 | [19] |
| LDCM0310 | AC48 | HEK-293T | C545(0.97); C1065(0.97); C332(0.84); C381(0.98) | LDD1549 | [19] |
| LDCM0311 | AC49 | HEK-293T | C545(1.06); C1065(1.02); C332(1.00); C381(1.08) | LDD1550 | [19] |
| LDCM0312 | AC5 | HEK-293T | C545(0.98); C1065(1.05); C332(0.93); C381(0.90) | LDD1551 | [19] |
| LDCM0313 | AC50 | HEK-293T | C545(1.02); C332(1.01); C381(0.98); C285(0.88) | LDD1552 | [19] |
| LDCM0314 | AC51 | HEK-293T | C545(1.04); C1065(1.11); C332(0.91); C381(1.06) | LDD1553 | [19] |
| LDCM0315 | AC52 | HEK-293T | C545(1.00); C1065(1.04); C332(0.92); C381(1.04) | LDD1554 | [19] |
| LDCM0316 | AC53 | HEK-293T | C545(1.00); C1065(1.04); C332(1.01); C381(0.92) | LDD1555 | [19] |
| LDCM0317 | AC54 | HEK-293T | C545(0.94); C1065(1.03); C332(0.98); C381(1.02) | LDD1556 | [19] |
| LDCM0318 | AC55 | HEK-293T | C545(0.94); C1065(0.93); C332(0.93); C381(1.04) | LDD1557 | [19] |
| LDCM0319 | AC56 | HEK-293T | C545(0.95); C1065(0.96); C332(0.94); C381(0.98) | LDD1558 | [19] |
| LDCM0320 | AC57 | HEK-293T | C545(1.13); C1065(1.09); C332(0.90); C381(1.04) | LDD1559 | [19] |
| LDCM0321 | AC58 | HEK-293T | C545(1.07); C332(0.97); C381(0.99); C285(0.86) | LDD1560 | [19] |
| LDCM0322 | AC59 | HEK-293T | C545(1.06); C1065(0.98); C332(0.92); C381(1.03) | LDD1561 | [19] |
| LDCM0323 | AC6 | HEK-293T | C545(1.07); C1065(0.95); C332(0.90); C381(1.02) | LDD1562 | [19] |
| LDCM0324 | AC60 | HEK-293T | C545(1.07); C1065(0.99); C332(0.87); C381(1.05) | LDD1563 | [19] |
| LDCM0325 | AC61 | HEK-293T | C545(1.02); C1065(1.04); C332(0.98); C381(0.96) | LDD1564 | [19] |
| LDCM0326 | AC62 | HEK-293T | C545(1.10); C1065(1.02); C332(0.97); C381(0.98) | LDD1565 | [19] |
| LDCM0327 | AC63 | HEK-293T | C545(1.04); C1065(1.17); C332(0.98); C381(1.00) | LDD1566 | [19] |
| LDCM0328 | AC64 | HEK-293T | C545(0.96); C1065(0.95); C332(0.90); C381(0.98) | LDD1567 | [19] |
| LDCM0334 | AC7 | HEK-293T | C545(1.03); C1065(0.86); C332(0.94); C381(1.04) | LDD1568 | [19] |
| LDCM0345 | AC8 | HEK-293T | C545(1.05); C1065(0.93); C332(0.97); C381(1.02) | LDD1569 | [19] |
| LDCM0248 | AKOS034007472 | HEK-293T | C545(0.96); C1065(1.03); C332(0.88); C381(0.95) | LDD1511 | [19] |
| LDCM0356 | AKOS034007680 | HEK-293T | C545(1.10); C1065(1.20); C332(0.93); C381(1.06) | LDD1570 | [19] |
| LDCM0275 | AKOS034007705 | HEK-293T | C545(0.97); C1065(0.91); C332(0.97); C381(0.98) | LDD1514 | [19] |
| LDCM0367 | CL1 | HEK-293T | C545(1.03); C1065(0.83); C332(1.00); C381(1.02) | LDD1571 | [19] |
| LDCM0368 | CL10 | HEK-293T | C545(0.86); C1065(1.15); C332(1.15); C381(1.84) | LDD1572 | [19] |
| LDCM0369 | CL100 | HEK-293T | C545(1.05); C1065(1.04); C332(0.89); C381(0.98) | LDD1573 | [19] |
| LDCM0370 | CL101 | HEK-293T | C545(1.04); C1065(1.06); C332(0.95); C381(0.94) | LDD1574 | [19] |
| LDCM0371 | CL102 | HEK-293T | C545(1.12); C1065(1.04); C332(0.95); C381(1.07) | LDD1575 | [19] |
| LDCM0372 | CL103 | HEK-293T | C545(1.03); C1065(0.95); C381(1.05); C285(1.04) | LDD1576 | [19] |
| LDCM0373 | CL104 | HEK-293T | C545(1.00); C1065(1.05); C332(0.93); C381(0.95) | LDD1577 | [19] |
| LDCM0374 | CL105 | HEK-293T | C545(1.03); C1065(1.01); C332(0.97); C381(1.02) | LDD1578 | [19] |
| LDCM0375 | CL106 | HEK-293T | C545(1.09); C1065(0.93); C332(1.00); C381(1.09) | LDD1579 | [19] |
| LDCM0376 | CL107 | HEK-293T | C545(1.00); C1065(1.14); C381(1.01); C285(1.05) | LDD1580 | [19] |
| LDCM0377 | CL108 | HEK-293T | C545(1.03); C1065(1.06); C332(0.93); C381(1.01) | LDD1581 | [19] |
| LDCM0378 | CL109 | HEK-293T | C545(0.95); C1065(1.06); C332(0.93); C381(1.09) | LDD1582 | [19] |
| LDCM0379 | CL11 | HEK-293T | C545(0.79); C1065(0.89); C332(0.97); C381(1.39) | LDD1583 | [19] |
| LDCM0380 | CL110 | HEK-293T | C545(1.00); C1065(1.07); C332(0.98); C381(1.12) | LDD1584 | [19] |
| LDCM0381 | CL111 | HEK-293T | C545(1.00); C1065(0.98); C381(1.15); C285(1.07) | LDD1585 | [19] |
| LDCM0382 | CL112 | HEK-293T | C545(0.98); C1065(1.18); C332(0.96); C381(1.02) | LDD1586 | [19] |
| LDCM0383 | CL113 | HEK-293T | C545(1.15); C1065(0.99); C332(0.98); C381(1.04) | LDD1587 | [19] |
| LDCM0384 | CL114 | HEK-293T | C545(1.09); C1065(1.14); C332(0.99); C381(1.17) | LDD1588 | [19] |
| LDCM0385 | CL115 | HEK-293T | C545(1.02); C1065(1.00); C381(1.07); C285(1.04) | LDD1589 | [19] |
| LDCM0386 | CL116 | HEK-293T | C545(1.04); C1065(1.16); C332(0.92); C381(0.93) | LDD1590 | [19] |
| LDCM0387 | CL117 | HEK-293T | C545(1.08); C1065(1.04); C332(0.89); C381(0.96) | LDD1591 | [19] |
| LDCM0388 | CL118 | HEK-293T | C545(1.19); C1065(1.06); C332(0.96); C381(1.00) | LDD1592 | [19] |
| LDCM0389 | CL119 | HEK-293T | C545(0.99); C1065(1.14); C381(0.97); C285(1.00) | LDD1593 | [19] |
| LDCM0390 | CL12 | HEK-293T | C545(0.83); C1065(0.97); C332(0.88); C381(1.00) | LDD1594 | [19] |
| LDCM0391 | CL120 | HEK-293T | C545(1.07); C1065(1.07); C332(0.93); C381(1.01) | LDD1595 | [19] |
| LDCM0392 | CL121 | HEK-293T | C545(1.09); C1065(1.06); C332(0.91); C381(1.08) | LDD1596 | [19] |
| LDCM0393 | CL122 | HEK-293T | C545(1.11); C1065(1.02); C332(0.98); C381(0.97) | LDD1597 | [19] |
| LDCM0394 | CL123 | HEK-293T | C545(0.94); C1065(1.21); C381(1.17); C285(1.03) | LDD1598 | [19] |
| LDCM0395 | CL124 | HEK-293T | C545(1.01); C1065(1.20); C332(0.97); C381(1.12) | LDD1599 | [19] |
| LDCM0396 | CL125 | HEK-293T | C545(1.09); C1065(1.05); C332(0.97); C381(0.93) | LDD1600 | [19] |
| LDCM0397 | CL126 | HEK-293T | C545(1.05); C1065(0.96); C332(0.93); C381(0.97) | LDD1601 | [19] |
| LDCM0398 | CL127 | HEK-293T | C545(1.07); C1065(0.99); C381(1.01); C285(0.97) | LDD1602 | [19] |
| LDCM0399 | CL128 | HEK-293T | C545(1.08); C1065(1.09); C332(0.97); C381(0.98) | LDD1603 | [19] |
| LDCM0400 | CL13 | HEK-293T | C545(1.28); C1065(1.03); C332(1.00); C381(1.16) | LDD1604 | [19] |
| LDCM0401 | CL14 | HEK-293T | C545(1.09); C1065(0.90); C332(0.92); C381(0.93) | LDD1605 | [19] |
| LDCM0402 | CL15 | HEK-293T | C545(1.10); C1065(1.05); C381(1.23); C285(1.12) | LDD1606 | [19] |
| LDCM0403 | CL16 | HEK-293T | C545(0.99); C1065(1.11); C332(0.89); C381(1.04) | LDD1607 | [19] |
| LDCM0404 | CL17 | HEK-293T | C545(1.18); C1065(1.08); C332(1.37); C381(1.38) | LDD1608 | [19] |
| LDCM0405 | CL18 | HEK-293T | C545(0.88); C332(0.99); C381(1.02); C285(0.96) | LDD1609 | [19] |
| LDCM0406 | CL19 | HEK-293T | C545(0.86); C1065(1.06); C332(0.92); C381(1.11) | LDD1610 | [19] |
| LDCM0407 | CL2 | HEK-293T | C545(1.13); C1065(0.84); C332(1.01); C381(0.95) | LDD1611 | [19] |
| LDCM0408 | CL20 | HEK-293T | C545(0.85); C1065(0.95); C332(0.89); C381(1.22) | LDD1612 | [19] |
| LDCM0409 | CL21 | HEK-293T | C545(0.91); C1065(0.96); C332(1.00); C381(1.38) | LDD1613 | [19] |
| LDCM0410 | CL22 | HEK-293T | C545(0.88); C1065(1.08); C332(1.06); C381(1.26) | LDD1614 | [19] |
| LDCM0411 | CL23 | HEK-293T | C545(0.69); C1065(0.98); C332(1.01); C381(1.19) | LDD1615 | [19] |
| LDCM0412 | CL24 | HEK-293T | C545(0.84); C1065(1.13); C332(0.97); C381(0.97) | LDD1616 | [19] |
| LDCM0413 | CL25 | HEK-293T | C545(1.04); C1065(0.99); C332(0.99); C381(1.06) | LDD1617 | [19] |
| LDCM0414 | CL26 | HEK-293T | C545(0.97); C1065(0.94); C332(1.01); C381(0.95) | LDD1618 | [19] |
| LDCM0415 | CL27 | HEK-293T | C545(0.93); C1065(1.00); C381(1.03); C285(1.11) | LDD1619 | [19] |
| LDCM0416 | CL28 | HEK-293T | C545(0.94); C1065(1.06); C332(0.97); C381(1.06) | LDD1620 | [19] |
| LDCM0417 | CL29 | HEK-293T | C545(1.00); C1065(0.78); C332(1.04); C381(1.09) | LDD1621 | [19] |
| LDCM0418 | CL3 | HEK-293T | C545(0.98); C1065(0.98); C381(1.05); C285(1.21) | LDD1622 | [19] |
| LDCM0419 | CL30 | HEK-293T | C545(0.91); C332(1.01); C381(0.98); C285(0.95) | LDD1623 | [19] |
| LDCM0420 | CL31 | HEK-293T | C545(0.94); C1065(1.36); C332(0.98); C381(1.14) | LDD1624 | [19] |
| LDCM0421 | CL32 | HEK-293T | C545(0.85); C1065(0.93); C332(0.95); C381(1.13) | LDD1625 | [19] |
| LDCM0422 | CL33 | HEK-293T | C545(0.80); C1065(1.32); C332(1.05); C381(1.61) | LDD1626 | [19] |
| LDCM0423 | CL34 | HEK-293T | C545(0.83); C1065(1.07); C332(1.02); C381(1.30) | LDD1627 | [19] |
| LDCM0424 | CL35 | HEK-293T | C545(0.72); C1065(0.90); C332(0.96); C381(1.33) | LDD1628 | [19] |
| LDCM0425 | CL36 | HEK-293T | C545(0.81); C1065(0.99); C332(0.99); C381(1.16) | LDD1629 | [19] |
| LDCM0426 | CL37 | HEK-293T | C545(1.02); C1065(0.94); C332(0.87); C381(0.92) | LDD1630 | [19] |
| LDCM0428 | CL39 | HEK-293T | C545(0.89); C1065(1.02); C381(1.15); C285(1.21) | LDD1632 | [19] |
| LDCM0429 | CL4 | HEK-293T | C545(1.03); C1065(1.00); C332(0.98); C381(0.97) | LDD1633 | [19] |
| LDCM0430 | CL40 | HEK-293T | C545(0.91); C1065(1.01); C332(0.86); C381(0.97) | LDD1634 | [19] |
| LDCM0431 | CL41 | HEK-293T | C545(0.97); C1065(1.13); C332(1.02); C381(1.22) | LDD1635 | [19] |
| LDCM0432 | CL42 | HEK-293T | C545(0.94); C332(0.96); C381(1.01); C285(1.01) | LDD1636 | [19] |
| LDCM0433 | CL43 | HEK-293T | C545(0.84); C1065(0.85); C332(0.99); C381(1.00) | LDD1637 | [19] |
| LDCM0434 | CL44 | HEK-293T | C545(0.95); C1065(1.04); C332(0.97); C381(1.12) | LDD1638 | [19] |
| LDCM0435 | CL45 | HEK-293T | C545(0.83); C1065(1.16); C332(0.96); C381(1.19) | LDD1639 | [19] |
| LDCM0436 | CL46 | HEK-293T | C545(0.85); C1065(0.99); C332(1.12); C381(1.34) | LDD1640 | [19] |
| LDCM0437 | CL47 | HEK-293T | C545(0.86); C1065(0.82); C332(0.97); C381(1.40) | LDD1641 | [19] |
| LDCM0438 | CL48 | HEK-293T | C545(0.84); C1065(1.08); C332(1.01); C381(1.09) | LDD1642 | [19] |
| LDCM0439 | CL49 | HEK-293T | C545(1.01); C1065(0.90); C332(0.84); C381(0.98) | LDD1643 | [19] |
| LDCM0440 | CL5 | HEK-293T | C545(1.16); C1065(1.01); C332(0.86); C381(1.05) | LDD1644 | [19] |
| LDCM0441 | CL50 | HEK-293T | C545(0.92); C1065(0.89); C332(0.94); C381(1.00) | LDD1645 | [19] |
| LDCM0443 | CL52 | HEK-293T | C545(0.97); C1065(1.11); C332(0.86); C381(0.99) | LDD1646 | [19] |
| LDCM0444 | CL53 | HEK-293T | C545(0.99); C1065(1.07); C332(0.81); C381(1.23) | LDD1647 | [19] |
| LDCM0445 | CL54 | HEK-293T | C545(0.95); C332(0.98); C381(1.19); C285(1.05) | LDD1648 | [19] |
| LDCM0446 | CL55 | HEK-293T | C545(0.99); C1065(0.94); C332(0.97); C381(1.04) | LDD1649 | [19] |
| LDCM0447 | CL56 | HEK-293T | C545(0.88); C1065(1.02); C332(0.92); C381(1.15) | LDD1650 | [19] |
| LDCM0448 | CL57 | HEK-293T | C545(1.01); C1065(1.16); C332(1.00); C381(1.27) | LDD1651 | [19] |
| LDCM0449 | CL58 | HEK-293T | C545(0.80); C1065(1.01); C332(0.96); C381(1.41) | LDD1652 | [19] |
| LDCM0450 | CL59 | HEK-293T | C545(0.82); C1065(1.11); C332(0.96); C381(1.39) | LDD1653 | [19] |
| LDCM0451 | CL6 | HEK-293T | C545(0.99); C332(0.94); C381(1.19); C285(1.07) | LDD1654 | [19] |
| LDCM0452 | CL60 | HEK-293T | C545(0.80); C1065(0.99); C332(0.98); C381(1.06) | LDD1655 | [19] |
| LDCM0453 | CL61 | HEK-293T | C545(0.97); C1065(0.88); C332(0.92); C381(0.93) | LDD1656 | [19] |
| LDCM0454 | CL62 | HEK-293T | C545(1.01); C1065(0.93); C332(0.94); C381(0.96) | LDD1657 | [19] |
| LDCM0455 | CL63 | HEK-293T | C545(0.99); C1065(1.05); C381(1.04); C285(0.92) | LDD1658 | [19] |
| LDCM0456 | CL64 | HEK-293T | C545(0.99); C1065(1.08); C332(0.98); C381(1.07) | LDD1659 | [19] |
| LDCM0457 | CL65 | HEK-293T | C545(0.95); C1065(1.24); C332(0.89); C381(1.09) | LDD1660 | [19] |
| LDCM0458 | CL66 | HEK-293T | C545(0.96); C332(1.00); C381(1.02); C285(1.02) | LDD1661 | [19] |
| LDCM0459 | CL67 | HEK-293T | C545(1.01); C1065(0.98); C332(0.92); C381(1.03) | LDD1662 | [19] |
| LDCM0460 | CL68 | HEK-293T | C545(0.86); C1065(0.99); C332(0.98); C381(1.12) | LDD1663 | [19] |
| LDCM0461 | CL69 | HEK-293T | C545(0.84); C1065(1.08); C332(0.96); C381(1.05) | LDD1664 | [19] |
| LDCM0462 | CL7 | HEK-293T | C545(0.97); C1065(0.91); C332(0.96); C381(1.11) | LDD1665 | [19] |
| LDCM0463 | CL70 | HEK-293T | C545(0.84); C1065(1.02); C332(0.97); C381(1.22) | LDD1666 | [19] |
| LDCM0464 | CL71 | HEK-293T | C545(0.86); C1065(1.11); C332(0.99); C381(1.33) | LDD1667 | [19] |
| LDCM0465 | CL72 | HEK-293T | C545(0.77); C1065(1.02); C332(1.00); C381(1.14) | LDD1668 | [19] |
| LDCM0466 | CL73 | HEK-293T | C545(0.99); C1065(1.01); C332(0.94); C381(1.05) | LDD1669 | [19] |
| LDCM0467 | CL74 | HEK-293T | C545(1.02); C1065(0.93); C332(0.91); C381(0.92) | LDD1670 | [19] |
| LDCM0469 | CL76 | HEK-293T | C545(1.03); C1065(1.00); C332(0.93); C381(1.00) | LDD1672 | [19] |
| LDCM0470 | CL77 | HEK-293T | C545(0.94); C1065(1.16); C332(1.00); C381(1.34) | LDD1673 | [19] |
| LDCM0471 | CL78 | HEK-293T | C545(0.95); C332(0.94); C381(1.02); C285(0.92) | LDD1674 | [19] |
| LDCM0472 | CL79 | HEK-293T | C545(0.96); C1065(1.04); C332(0.90); C381(1.01) | LDD1675 | [19] |
| LDCM0473 | CL8 | HEK-293T | C545(0.85); C1065(1.12); C332(0.93); C381(1.95) | LDD1676 | [19] |
| LDCM0474 | CL80 | HEK-293T | C545(0.85); C1065(1.02); C332(0.92); C381(1.02) | LDD1677 | [19] |
| LDCM0475 | CL81 | HEK-293T | C545(0.89); C1065(1.07); C332(0.94); C381(1.10) | LDD1678 | [19] |
| LDCM0476 | CL82 | HEK-293T | C545(0.90); C1065(1.09); C332(0.97); C381(1.18) | LDD1679 | [19] |
| LDCM0477 | CL83 | HEK-293T | C545(0.83); C1065(1.05); C332(1.01); C381(1.27) | LDD1680 | [19] |
| LDCM0478 | CL84 | HEK-293T | C545(0.83); C1065(1.05); C332(1.01); C381(1.09) | LDD1681 | [19] |
| LDCM0479 | CL85 | HEK-293T | C545(1.05); C1065(0.91); C332(0.87); C381(0.89) | LDD1682 | [19] |
| LDCM0480 | CL86 | HEK-293T | C545(1.02); C1065(0.84); C332(0.88); C381(0.87) | LDD1683 | [19] |
| LDCM0481 | CL87 | HEK-293T | C545(1.04); C1065(1.13); C381(1.06); C285(1.05) | LDD1684 | [19] |
| LDCM0482 | CL88 | HEK-293T | C545(1.01); C1065(1.05); C332(0.88); C381(0.89) | LDD1685 | [19] |
| LDCM0483 | CL89 | HEK-293T | C545(0.95); C1065(0.98); C332(1.04); C381(1.02) | LDD1686 | [19] |
| LDCM0484 | CL9 | HEK-293T | C545(0.95); C1065(1.03); C332(0.95); C381(1.07) | LDD1687 | [19] |
| LDCM0485 | CL90 | HEK-293T | C545(1.07); C332(1.18); C381(1.74); C285(1.17) | LDD1688 | [19] |
| LDCM0486 | CL91 | HEK-293T | C545(1.01); C1065(1.06); C332(0.96); C381(1.05) | LDD1689 | [19] |
| LDCM0487 | CL92 | HEK-293T | C545(0.95); C1065(1.08); C332(0.85); C381(1.08) | LDD1690 | [19] |
| LDCM0488 | CL93 | HEK-293T | C545(0.89); C1065(0.99); C332(0.89); C381(1.09) | LDD1691 | [19] |
| LDCM0489 | CL94 | HEK-293T | C545(0.87); C1065(1.05); C332(0.97); C381(1.04) | LDD1692 | [19] |
| LDCM0490 | CL95 | HEK-293T | C545(0.86); C1065(1.39); C332(1.01); C381(1.50) | LDD1693 | [19] |
| LDCM0491 | CL96 | HEK-293T | C545(1.01); C1065(0.98); C332(0.96); C381(1.05) | LDD1694 | [19] |
| LDCM0492 | CL97 | HEK-293T | C545(1.08); C1065(0.94); C332(0.86); C381(0.99) | LDD1695 | [19] |
| LDCM0493 | CL98 | HEK-293T | C545(1.04); C1065(0.90); C332(0.97); C381(1.03) | LDD1696 | [19] |
| LDCM0494 | CL99 | HEK-293T | C545(1.03); C1065(1.04); C381(1.03); C285(1.01) | LDD1697 | [19] |
| LDCM0495 | E2913 | HEK-293T | C545(0.92); C1065(0.98); C381(1.11); C285(0.97) | LDD1698 | [19] |
| LDCM0468 | Fragment33 | HEK-293T | C545(0.94); C1065(0.97); C381(1.05); C285(1.02) | LDD1671 | [19] |
| LDCM0427 | Fragment51 | HEK-293T | C545(1.09); C1065(0.90); C332(0.98); C381(0.94) | LDD1631 | [19] |
| LDCM0022 | KB02 | HCT 116 | C1054(6.91); C332(2.58); C1065(3.47); C1055(3.60) | LDD0080 | [7] |
| LDCM0023 | KB03 | HCT 116 | C1054(6.83); C332(2.29); C1065(4.07); C1055(4.11) | LDD0081 | [7] |
| LDCM0024 | KB05 | HCT 116 | C1054(9.52); C332(4.53); C1065(4.47); C1055(4.01) | LDD0082 | [7] |
| LDCM0627 | NUDT7-COV-1 | HEK-293T | C257(1.06); C1065(0.61); C332(0.49); C206(0.48) | LDD2206 | [20] |
| LDCM0628 | OTUB2-COV-1 | HEK-293T | C381(1.57); C206(0.83); C332(0.78); C1065(0.72) | LDD2207 | [20] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Prostaglandin E synthase 3 (PTGES3) | P23/wos2 family | Q15185 | |||
| Epidermal growth factor receptor (EGFR) | Tyr protein kinase family | P00533 | |||
Other
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Microtubule-associated proteins 1A/1B light chain 3A (MAP1LC3A) | ATG8 family | Q9H492 | |||
| Adapter molecule crk (CRK) | CRK family | P46108 | |||
References
