Details of the Target
General Information of Target
| Target ID | LDTP12790 | |||||
|---|---|---|---|---|---|---|
| Target Name | Glutaminyl-peptide cyclotransferase-like protein (QPCTL) | |||||
| Gene Name | QPCTL | |||||
| Gene ID | 54814 | |||||
| Synonyms |
Glutaminyl-peptide cyclotransferase-like protein; EC 2.3.2.5; Golgi-resident glutaminyl-peptide cyclotransferase; isoQC; gQC |
|||||
| 3D Structure | ||||||
| Sequence |
MLYLEDYLEMIEQLPMDLRDRFTEMREMDLQVQNAMDQLEQRVSEFFMNAKKNKPEWREE
QMASIKKDYYKALEDADEKVQLANQIYDLVDRHLRKLDQELAKFKMELEADNAGITEILE RRSLELDTPSQPVNNHHAHSHTPVEKRKYNPTSHHTTTDHIPEKKFKSEALLSTLTSDAS KENTLGCRNNNSTASSNNAYNVNSSQPLGSYNIGSLSSGTGAGAITMAAAQAVQATAQMK EGRRTSSLKASYEAFKNNDFQLGKEFSMARETVGYSSSSALMTTLTQNASSSAADSRSGR KSKNNNKSSSQQSSSSSSSSSLSSCSSSSTVVQEISQQTTVVPESDSNSQVDWTYDPNEP RYCICNQVSYGEMVGCDNQDCPIEWFHYGCVGLTEAPKGKWYCPQCTAAMKRRGSRHK |
|||||
| Target Bioclass |
Enzyme
|
|||||
| Family |
Glutaminyl-peptide cyclotransferase family
|
|||||
| Subcellular location |
Golgi apparatus membrane
|
|||||
| Function | Responsible for the biosynthesis of pyroglutamyl peptides. | |||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Target Site Mutations in Different Cell Lines
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
Acrolein Probe Info |
 |
H252(0.00); H162(0.00) | LDD0227 | [1] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0165 | [2] | |
|
NAIA_5 Probe Info |
 |
N.A. | LDD2223 | [3] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C106 Probe Info |
 |
66.26 | LDD1793 | [4] | |
|
C107 Probe Info |
 |
20.97 | LDD1794 | [4] | |
|
C108 Probe Info |
 |
16.56 | LDD1795 | [4] | |
|
C252 Probe Info |
 |
11.08 | LDD1925 | [4] | |
|
C388 Probe Info |
 |
79.34 | LDD2047 | [4] | |
|
VE-P Probe Info |
 |
N.A. | LDD0396 | [5] | |
|
Staurosporine capture compound Probe Info |
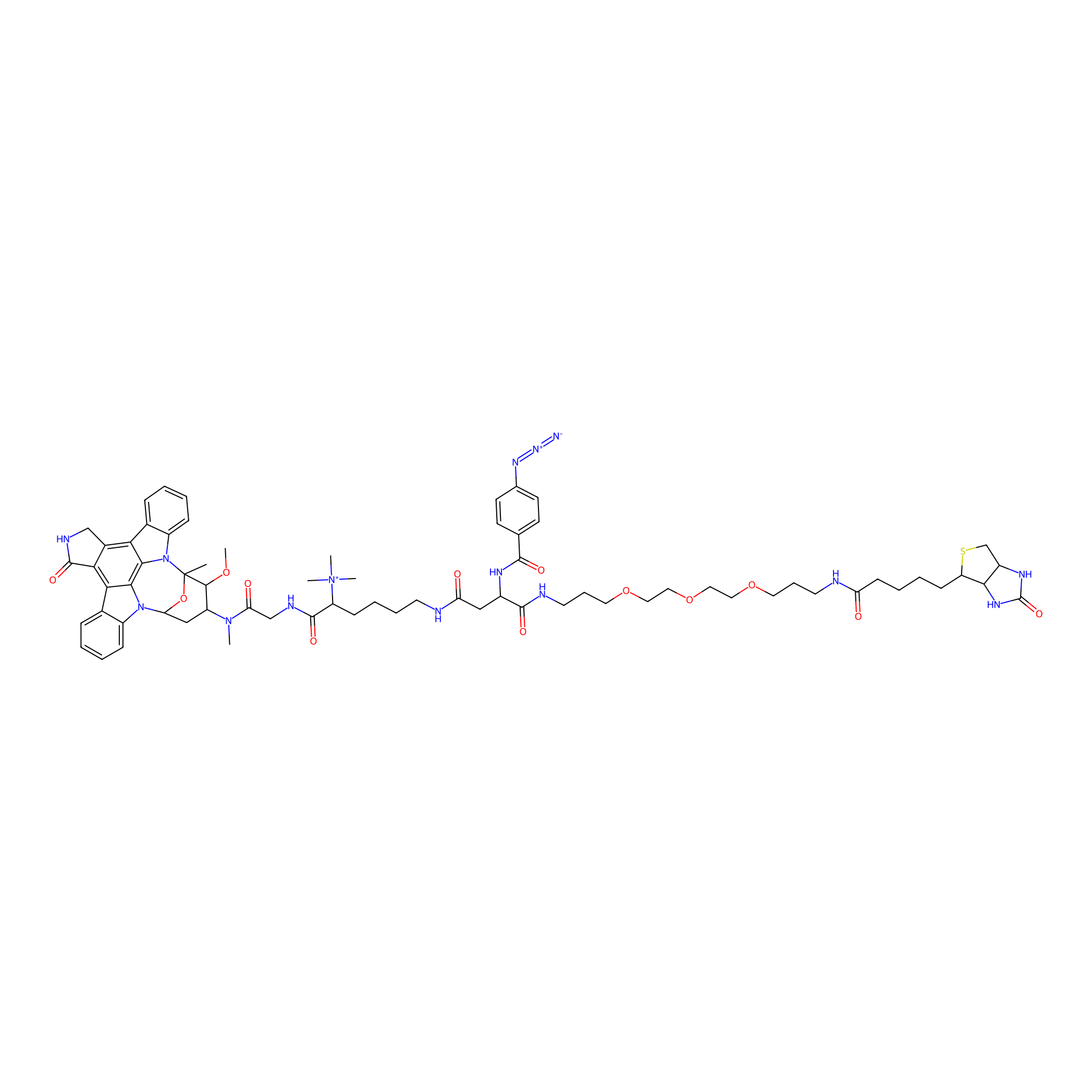 |
N.A. | LDD0083 | [6] | |
|
OEA-DA Probe Info |
 |
20.00 | LDD0046 | [7] | |
Competitor(s) Related to This Target
The Interaction Atlas With This Target
References
