Details of the Target
General Information of Target
| Target ID | LDTP07317 | |||||
|---|---|---|---|---|---|---|
| Target Name | Tensin-3 (TNS3) | |||||
| Gene Name | TNS3 | |||||
| Gene ID | 64759 | |||||
| Synonyms |
TEM6; TENS1; TPP; Tensin-3; EC 3.1.3.-; Tensin-like SH2 domain-containing protein 1; Tumor endothelial marker 6 |
|||||
| 3D Structure | ||||||
| Sequence |
MEEGHGLDLTYITERIIAVSFPAGCSEESYLHNLQEVTRMLKSKHGDNYLVLNLSEKRYD
LTKLNPKIMDVGWPELHAPPLDKMCTICKAQESWLNSNLQHVVVIHCRGGKGRIGVVISS YMHFTNVSASADQALDRFAMKKFYDDKVSALMQPSQKRYVQFLSGLLSGSVKMNASPLFL HFVILHGTPNFDTGGVCRPFLKLYQAMQPVYTSGIYNVGPENPSRICIVIEPAQLLKGDV MVKCYHKKYRSATRDVIFRLQFHTGAVQGYGLVFGKEDLDNASKDDRFPDYGKVELVFSA TPEKIQGSEHLYNDHGVIVDYNTTDPLIRWDSYENLSADGEVLHTQGPVDGSLYAKVRKK SSSDPGIPGGPQAIPATNSPDHSDHTLSVSSDSGHSTASARTDKTEERLAPGTRRGLSAQ EKAELDQLLSGFGLEDPGSSLKEMTDARSKYSGTRHVVPAQVHVNGDAALKDRETDILDD EMPHHDLHSVDSLGTLSSSEGPQSAHLGPFTCHKSSQNSLLSDGFGSNVGEDPQGTLVPD LGLGMDGPYERERTFGSREPKQPQPLLRKPSVSAQMQAYGQSSYSTQTWVRQQQMVVAHQ YSFAPDGEARLVSRCPADNPGLVQAQPRVPLTPTRGTSSRVAVQRGVGSGPHPPDTQQPS PSKAFKPRFPGDQVVNGAGPELSTGPSPGSPTLDIDQSIEQLNRLILELDPTFEPIPTHM NALGSQANGSVSPDSVGGGLRASSRLPDTGEGPSRATGRQGSSAEQPLGGRLRKLSLGQY DNDAGGQLPFSKCAWGKAGVDYAPNLPPFPSPADVKETMTPGYPQDLDIIDGRILSSKES MCSTPAFPVSPETPYVKTALRHPPFSPPEPPLSSPASQHKGGREPRSCPETLTHAVGMSE SPIGPKSTMLRADASSTPSFQQAFASSCTISSNGPGQRRESSSSAERQWVESSPKPMVSL LGSGRPTGSPLSAEFSGTRKDSPVLSCFPPSELQAPFHSHELSLAEPPDSLAPPSSQAFL GFGTAPVGSGLPPEEDLGALLANSHGASPTPSIPLTATGAADNGFLSHNFLTVAPGHSSH HSPGLQGQGVTLPGQPPLPEKKRASEGDRSLGSVSPSSSGFSSPHSGSTISIPFPNVLPD FSKASEAASPLPDSPGDKLVIVKFVQDTSKFWYKADISREQAIAMLKDKEPGSFIVRDSH SFRGAYGLAMKVATPPPSVLQLNKKAGDLANELVRHFLIECTPKGVRLKGCSNEPYFGSL TALVCQHSITPLALPCKLLIPERDPLEEIAESSPQTAANSAAELLKQGAACNVWYLNSVE MESLTGHQAIQKALSITLVQEPPPVSTVVHFKVSAQGITLTDNQRKLFFRRHYPVNSVIF CALDPQDRKWIKDGPSSKVFGFVARKQGSATDNVCHLFAEHDPEQPASAIVNFVSKVMIG SPKKV |
|||||
| Target Bioclass |
Enzyme
|
|||||
| Family |
PTEN phosphatase protein family
|
|||||
| Subcellular location |
Cell junction, focal adhesion
|
|||||
| Function |
May act as a protein phosphatase and/or a lipid phosphatase (Probable). Involved in the dissociation of the integrin-tensin-actin complex. EGF activates TNS4 and down-regulates TNS3 which results in capping the tail of ITGB1. Increases DOCK5 guanine nucleotide exchange activity towards Rac and plays a role in osteoclast podosome organization. Enhances RHOA activation in the presence of DLC1. Required for growth factor-induced epithelial cell migration; growth factor stimulation induces TNS3 phosphorylation which changes its binding preference from DLC1 to the p85 regulatory subunit of the PI3K kinase complex, displacing PI3K inhibitor PTEN and resulting in translocation of the TNS3-p85 complex to the leading edge of migrating cells to promote RAC1 activation. Meanwhile, PTEN switches binding preference from p85 to DLC1 and the PTEN-DLC1 complex translocates to the posterior of migrating cells to activate RHOA. Acts as an adapter protein by bridging the association of scaffolding protein PEAK1 with integrins ITGB1, ITGB3 and ITGB5 which contributes to the promotion of cell migration. Controls tonsil-derived mesenchymal stem cell proliferation and differentiation by regulating the activity of integrin ITGB1.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| 8305C | SNV: p.C1251F | . | |||
| AU565 | SNV: p.S944C | DBIA Probe Info | |||
| CCLP1 | SNV: p.L1031H | DBIA Probe Info | |||
| CORL23 | SNV: p.R668S | DBIA Probe Info | |||
| HCT15 | SNV: p.P918S | . | |||
| HT1376 | SNV: p.H1068P | DBIA Probe Info | |||
| IGR1 | Substitution: p.P767F | DBIA Probe Info | |||
| JURKAT | SNV: p.S29Y | . | |||
| KMCH1 | SNV: p.S1122C | DBIA Probe Info | |||
| MELJUSO | SNV: p.P365L | DBIA Probe Info | |||
| NCIH2286 | SNV: p.A1427S | DBIA Probe Info | |||
| SISO | SNV: p.Q921Ter; p.Q1166E | DBIA Probe Info | |||
| SKN | Deletion: p.P747HfsTer17 | DBIA Probe Info | |||
| TF1 | SNV: p.N1043D | DBIA Probe Info | |||
| TOV21G | SNV: p.S452N | DBIA Probe Info | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
STPyne Probe Info |
 |
K1158(1.15); K1170(5.88); K147(5.88); K792(10.00) | LDD0277 | [1] | |
|
m-APA Probe Info |
 |
13.04 | LDD0403 | [2] | |
|
BTD Probe Info |
 |
C615(0.65) | LDD2116 | [3] | |
|
Sulforaphane-probe2 Probe Info |
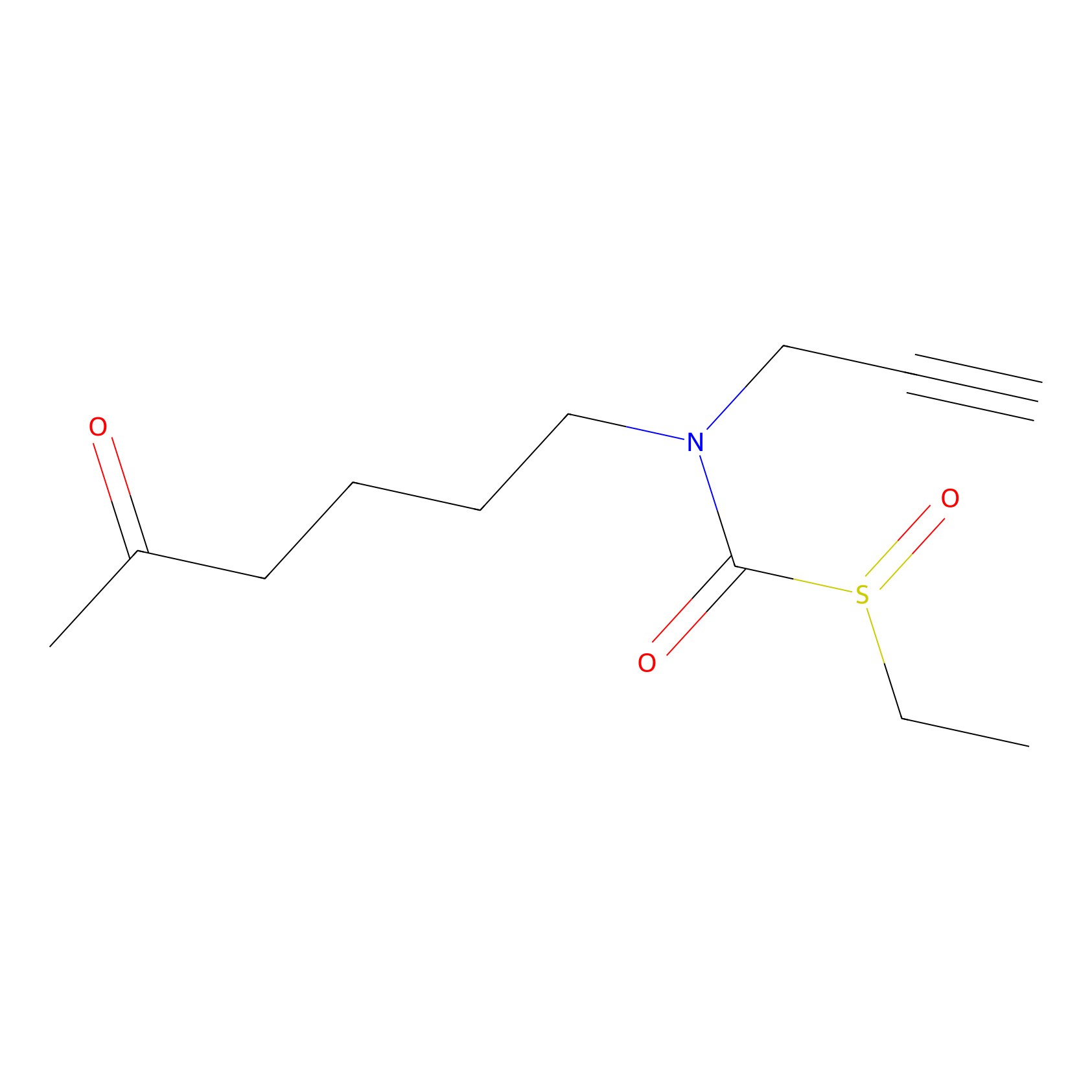 |
2.35 | LDD0160 | [4] | |
|
AHL-Pu-1 Probe Info |
 |
C842(4.52) | LDD0170 | [5] | |
|
DBIA Probe Info |
 |
C928(1.10); C842(1.04); C1241(1.02); C615(0.95) | LDD0078 | [6] | |
|
JW-RF-010 Probe Info |
 |
C615(0.00); C25(0.00) | LDD0026 | [7] | |
|
NAIA_4 Probe Info |
 |
C25(0.00); C615(0.00); C842(0.00); C888(0.00) | LDD2226 | [8] | |
|
TFBX Probe Info |
 |
N.A. | LDD0027 | [7] | |
|
WYneN Probe Info |
 |
C615(0.00); C888(0.00); C1241(0.00) | LDD0021 | [9] | |
|
WYneO Probe Info |
 |
C888(0.00); C615(0.00) | LDD0022 | [9] | |
|
IA-alkyne Probe Info |
 |
C615(0.00); C1241(0.00) | LDD0151 | [10] | |
|
IPM Probe Info |
 |
N.A. | LDD0005 | [9] | |
|
Phosphinate-6 Probe Info |
 |
C1241(0.00); C615(0.00) | LDD0018 | [11] | |
|
Acrolein Probe Info |
 |
C842(0.00); C25(0.00); C1241(0.00) | LDD0217 | [12] | |
|
Methacrolein Probe Info |
 |
C842(0.00); C1241(0.00) | LDD0218 | [12] | |
|
W1 Probe Info |
 |
N.A. | LDD0236 | [13] | |
|
NAIA_5 Probe Info |
 |
C888(0.00); C1241(0.00); C197(0.00); C1311(0.00) | LDD2223 | [8] | |
|
HHS-482 Probe Info |
 |
Y855(4.42) | LDD2239 | [14] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0524 | 2-Cyano-N-(2-morpholin-4-yl-ethyl)-acetamide | MDA-MB-231 | C1241(1.10) | LDD2117 | [3] |
| LDCM0025 | 4SU-RNA | DM93 | C842(4.52) | LDD0170 | [5] |
| LDCM0215 | AC10 | HEK-293T | C615(1.08) | LDD1508 | [15] |
| LDCM0216 | AC100 | HCT 116 | C615(0.84) | LDD0533 | [6] |
| LDCM0217 | AC101 | HCT 116 | C615(0.97) | LDD0534 | [6] |
| LDCM0218 | AC102 | HCT 116 | C615(0.76) | LDD0535 | [6] |
| LDCM0219 | AC103 | HCT 116 | C615(0.86) | LDD0536 | [6] |
| LDCM0220 | AC104 | HCT 116 | C615(0.91) | LDD0537 | [6] |
| LDCM0221 | AC105 | HCT 116 | C615(0.96) | LDD0538 | [6] |
| LDCM0222 | AC106 | HCT 116 | C615(0.94) | LDD0539 | [6] |
| LDCM0223 | AC107 | HCT 116 | C615(0.97) | LDD0540 | [6] |
| LDCM0224 | AC108 | HCT 116 | C615(0.70) | LDD0541 | [6] |
| LDCM0225 | AC109 | HCT 116 | C615(0.56) | LDD0542 | [6] |
| LDCM0227 | AC110 | HCT 116 | C615(0.70) | LDD0544 | [6] |
| LDCM0228 | AC111 | HCT 116 | C615(0.76) | LDD0545 | [6] |
| LDCM0229 | AC112 | HCT 116 | C615(0.95) | LDD0546 | [6] |
| LDCM0230 | AC113 | HCT 116 | C615(0.84) | LDD0547 | [6] |
| LDCM0231 | AC114 | HCT 116 | C615(0.79) | LDD0548 | [6] |
| LDCM0232 | AC115 | HCT 116 | C615(1.03) | LDD0549 | [6] |
| LDCM0233 | AC116 | HCT 116 | C615(1.43) | LDD0550 | [6] |
| LDCM0234 | AC117 | HCT 116 | C615(1.08) | LDD0551 | [6] |
| LDCM0235 | AC118 | HCT 116 | C615(1.01) | LDD0552 | [6] |
| LDCM0236 | AC119 | HCT 116 | C615(0.90) | LDD0553 | [6] |
| LDCM0237 | AC12 | HEK-293T | C615(0.99) | LDD1510 | [15] |
| LDCM0238 | AC120 | HCT 116 | C615(0.90) | LDD0555 | [6] |
| LDCM0239 | AC121 | HCT 116 | C615(1.59) | LDD0556 | [6] |
| LDCM0240 | AC122 | HCT 116 | C615(0.80) | LDD0557 | [6] |
| LDCM0241 | AC123 | HCT 116 | C615(0.94) | LDD0558 | [6] |
| LDCM0242 | AC124 | HCT 116 | C615(1.16) | LDD0559 | [6] |
| LDCM0243 | AC125 | HCT 116 | C615(1.04) | LDD0560 | [6] |
| LDCM0244 | AC126 | HCT 116 | C615(1.38) | LDD0561 | [6] |
| LDCM0245 | AC127 | HCT 116 | C615(1.57) | LDD0562 | [6] |
| LDCM0246 | AC128 | HCT 116 | C615(1.05) | LDD0563 | [6] |
| LDCM0247 | AC129 | HCT 116 | C615(0.82) | LDD0564 | [6] |
| LDCM0249 | AC130 | HCT 116 | C615(1.25) | LDD0566 | [6] |
| LDCM0250 | AC131 | HCT 116 | C615(1.12) | LDD0567 | [6] |
| LDCM0251 | AC132 | HCT 116 | C615(0.68) | LDD0568 | [6] |
| LDCM0252 | AC133 | HCT 116 | C615(1.08) | LDD0569 | [6] |
| LDCM0253 | AC134 | HCT 116 | C615(1.01) | LDD0570 | [6] |
| LDCM0254 | AC135 | HCT 116 | C615(1.27) | LDD0571 | [6] |
| LDCM0255 | AC136 | HCT 116 | C615(0.93) | LDD0572 | [6] |
| LDCM0256 | AC137 | HCT 116 | C615(0.78) | LDD0573 | [6] |
| LDCM0257 | AC138 | HCT 116 | C615(0.87) | LDD0574 | [6] |
| LDCM0258 | AC139 | HCT 116 | C615(1.14) | LDD0575 | [6] |
| LDCM0259 | AC14 | HEK-293T | C615(1.25) | LDD1512 | [15] |
| LDCM0260 | AC140 | HCT 116 | C615(0.90) | LDD0577 | [6] |
| LDCM0261 | AC141 | HCT 116 | C615(0.79) | LDD0578 | [6] |
| LDCM0262 | AC142 | HCT 116 | C615(0.79) | LDD0579 | [6] |
| LDCM0263 | AC143 | HCT 116 | C615(1.19) | LDD0580 | [6] |
| LDCM0264 | AC144 | HCT 116 | C615(1.18) | LDD0581 | [6] |
| LDCM0265 | AC145 | HCT 116 | C615(1.23) | LDD0582 | [6] |
| LDCM0266 | AC146 | HCT 116 | C615(1.33) | LDD0583 | [6] |
| LDCM0267 | AC147 | HCT 116 | C615(1.30) | LDD0584 | [6] |
| LDCM0268 | AC148 | HCT 116 | C615(1.67) | LDD0585 | [6] |
| LDCM0269 | AC149 | HCT 116 | C615(1.32) | LDD0586 | [6] |
| LDCM0271 | AC150 | HCT 116 | C615(1.30) | LDD0588 | [6] |
| LDCM0272 | AC151 | HCT 116 | C615(1.16) | LDD0589 | [6] |
| LDCM0273 | AC152 | HCT 116 | C615(1.19) | LDD0590 | [6] |
| LDCM0274 | AC153 | HCT 116 | C615(1.70) | LDD0591 | [6] |
| LDCM0621 | AC154 | HCT 116 | C615(1.17) | LDD2158 | [6] |
| LDCM0622 | AC155 | HCT 116 | C615(1.39) | LDD2159 | [6] |
| LDCM0623 | AC156 | HCT 116 | C615(1.14) | LDD2160 | [6] |
| LDCM0624 | AC157 | HCT 116 | C615(1.25) | LDD2161 | [6] |
| LDCM0277 | AC18 | HEK-293T | C615(1.13) | LDD1516 | [15] |
| LDCM0279 | AC2 | HEK-293T | C615(1.26) | LDD1518 | [15] |
| LDCM0280 | AC20 | HEK-293T | C615(1.01) | LDD1519 | [15] |
| LDCM0281 | AC21 | HEK-293T | C1241(1.10) | LDD1520 | [15] |
| LDCM0282 | AC22 | HEK-293T | C615(0.94) | LDD1521 | [15] |
| LDCM0286 | AC26 | HEK-293T | C615(1.10) | LDD1525 | [15] |
| LDCM0288 | AC28 | HEK-293T | C615(0.67) | LDD1527 | [15] |
| LDCM0289 | AC29 | HEK-293T | C1241(1.00) | LDD1528 | [15] |
| LDCM0291 | AC30 | HEK-293T | C615(1.23) | LDD1530 | [15] |
| LDCM0295 | AC34 | HEK-293T | C615(1.03) | LDD1534 | [15] |
| LDCM0297 | AC36 | HEK-293T | C615(0.81) | LDD1536 | [15] |
| LDCM0298 | AC37 | HEK-293T | C1241(1.10) | LDD1537 | [15] |
| LDCM0299 | AC38 | HEK-293T | C615(1.23) | LDD1538 | [15] |
| LDCM0301 | AC4 | HEK-293T | C615(0.91) | LDD1540 | [15] |
| LDCM0304 | AC42 | HEK-293T | C615(1.20) | LDD1543 | [15] |
| LDCM0306 | AC44 | HEK-293T | C615(0.85) | LDD1545 | [15] |
| LDCM0307 | AC45 | HEK-293T | C1241(0.93) | LDD1546 | [15] |
| LDCM0308 | AC46 | HEK-293T | C615(1.17) | LDD1547 | [15] |
| LDCM0312 | AC5 | HEK-293T | C1241(0.98) | LDD1551 | [15] |
| LDCM0313 | AC50 | HEK-293T | C615(1.60) | LDD1552 | [15] |
| LDCM0315 | AC52 | HEK-293T | C615(1.09) | LDD1554 | [15] |
| LDCM0316 | AC53 | HEK-293T | C1241(1.07) | LDD1555 | [15] |
| LDCM0317 | AC54 | HEK-293T | C615(1.08) | LDD1556 | [15] |
| LDCM0321 | AC58 | HEK-293T | C615(1.00) | LDD1560 | [15] |
| LDCM0323 | AC6 | HEK-293T | C615(0.86) | LDD1562 | [15] |
| LDCM0324 | AC60 | HEK-293T | C615(1.09) | LDD1563 | [15] |
| LDCM0325 | AC61 | HEK-293T | C1241(1.01) | LDD1564 | [15] |
| LDCM0326 | AC62 | HEK-293T | C615(1.26) | LDD1565 | [15] |
| LDCM0365 | AC98 | HCT 116 | C615(1.04) | LDD0682 | [6] |
| LDCM0366 | AC99 | HCT 116 | C615(0.78) | LDD0683 | [6] |
| LDCM0248 | AKOS034007472 | HEK-293T | C1241(1.03) | LDD1511 | [15] |
| LDCM0156 | Aniline | NCI-H1299 | 13.04 | LDD0403 | [2] |
| LDCM0020 | ARS-1620 | HCC44 | C928(1.10); C842(1.04); C1241(1.02); C615(0.95) | LDD0078 | [6] |
| LDCM0630 | CCW28-3 | 231MFP | C928(1.36); C1241(1.35); C888(0.99) | LDD2214 | [16] |
| LDCM0108 | Chloroacetamide | HeLa | C842(0.00); C1241(0.00) | LDD0222 | [12] |
| LDCM0368 | CL10 | HEK-293T | C615(0.89) | LDD1572 | [15] |
| LDCM0405 | CL18 | HEK-293T | C615(0.98) | LDD1609 | [15] |
| LDCM0408 | CL20 | HEK-293T | C615(1.35) | LDD1612 | [15] |
| LDCM0409 | CL21 | HEK-293T | C1241(0.96) | LDD1613 | [15] |
| LDCM0410 | CL22 | HEK-293T | C615(0.83) | LDD1614 | [15] |
| LDCM0419 | CL30 | HEK-293T | C615(1.16) | LDD1623 | [15] |
| LDCM0421 | CL32 | HEK-293T | C615(1.01) | LDD1625 | [15] |
| LDCM0422 | CL33 | HEK-293T | C1241(1.31) | LDD1626 | [15] |
| LDCM0423 | CL34 | HEK-293T | C615(0.71) | LDD1627 | [15] |
| LDCM0432 | CL42 | HEK-293T | C615(1.33) | LDD1636 | [15] |
| LDCM0434 | CL44 | HEK-293T | C615(1.04) | LDD1638 | [15] |
| LDCM0435 | CL45 | HEK-293T | C1241(0.98) | LDD1639 | [15] |
| LDCM0436 | CL46 | HEK-293T | C615(1.03) | LDD1640 | [15] |
| LDCM0445 | CL54 | HEK-293T | C615(1.02) | LDD1648 | [15] |
| LDCM0447 | CL56 | HEK-293T | C615(0.94) | LDD1650 | [15] |
| LDCM0448 | CL57 | HEK-293T | C1241(0.95) | LDD1651 | [15] |
| LDCM0449 | CL58 | HEK-293T | C615(0.89) | LDD1652 | [15] |
| LDCM0451 | CL6 | HEK-293T | C615(1.04) | LDD1654 | [15] |
| LDCM0453 | CL61 | HCT 116 | C615(0.47) | LDD0770 | [6] |
| LDCM0454 | CL62 | HCT 116 | C615(0.59) | LDD0771 | [6] |
| LDCM0455 | CL63 | HCT 116 | C615(1.07) | LDD0772 | [6] |
| LDCM0456 | CL64 | HCT 116 | C615(1.48) | LDD0773 | [6] |
| LDCM0457 | CL65 | HCT 116 | C615(1.15) | LDD0774 | [6] |
| LDCM0458 | CL66 | HCT 116 | C615(1.52) | LDD0775 | [6] |
| LDCM0459 | CL67 | HCT 116 | C615(1.18) | LDD0776 | [6] |
| LDCM0460 | CL68 | HCT 116 | C615(0.94) | LDD0777 | [6] |
| LDCM0461 | CL69 | HCT 116 | C615(0.95) | LDD0778 | [6] |
| LDCM0463 | CL70 | HCT 116 | C615(1.01) | LDD0780 | [6] |
| LDCM0464 | CL71 | HCT 116 | C615(1.13) | LDD0781 | [6] |
| LDCM0465 | CL72 | HCT 116 | C615(0.99) | LDD0782 | [6] |
| LDCM0466 | CL73 | HCT 116 | C615(1.23) | LDD0783 | [6] |
| LDCM0467 | CL74 | HCT 116 | C615(1.00) | LDD0784 | [6] |
| LDCM0471 | CL78 | HEK-293T | C615(1.29) | LDD1674 | [15] |
| LDCM0473 | CL8 | HEK-293T | C615(0.79) | LDD1676 | [15] |
| LDCM0474 | CL80 | HEK-293T | C615(1.12) | LDD1677 | [15] |
| LDCM0475 | CL81 | HEK-293T | C1241(0.98) | LDD1678 | [15] |
| LDCM0476 | CL82 | HEK-293T | C615(0.91) | LDD1679 | [15] |
| LDCM0484 | CL9 | HEK-293T | C1241(1.02) | LDD1687 | [15] |
| LDCM0485 | CL90 | HEK-293T | C615(0.97) | LDD1688 | [15] |
| LDCM0487 | CL92 | HEK-293T | C615(1.04) | LDD1690 | [15] |
| LDCM0488 | CL93 | HEK-293T | C1241(0.98) | LDD1691 | [15] |
| LDCM0489 | CL94 | HEK-293T | C615(1.08) | LDD1692 | [15] |
| LDCM0213 | Electrophilic fragment 2 | MDA-MB-231 | C1241(1.26); C888(1.07) | LDD1702 | [3] |
| LDCM0575 | Fragment13 | MDA-MB-231 | C888(1.42) | LDD1395 | [17] |
| LDCM0576 | Fragment14 | MDA-MB-231 | C888(1.89) | LDD1397 | [17] |
| LDCM0579 | Fragment20 | MDA-MB-231 | C888(2.82) | LDD1402 | [17] |
| LDCM0580 | Fragment21 | MDA-MB-231 | C888(0.89) | LDD1404 | [17] |
| LDCM0578 | Fragment27 | MDA-MB-231 | C888(0.40) | LDD1401 | [17] |
| LDCM0588 | Fragment30 | MDA-MB-231 | C888(20.00) | LDD1419 | [17] |
| LDCM0590 | Fragment32 | MDA-MB-231 | C888(4.01) | LDD1423 | [17] |
| LDCM0468 | Fragment33 | HCT 116 | C615(1.25) | LDD0785 | [6] |
| LDCM0596 | Fragment38 | MDA-MB-231 | C888(1.14) | LDD1433 | [17] |
| LDCM0599 | Fragment41 | MDA-MB-231 | C888(1.32) | LDD1438 | [17] |
| LDCM0601 | Fragment43 | MDA-MB-231 | C888(2.16) | LDD1441 | [17] |
| LDCM0604 | Fragment46 | MDA-MB-231 | C888(1.01) | LDD1445 | [17] |
| LDCM0605 | Fragment47 | MDA-MB-231 | C888(1.01) | LDD1446 | [17] |
| LDCM0607 | Fragment49 | MDA-MB-231 | C888(2.76) | LDD1448 | [17] |
| LDCM0614 | Fragment56 | MDA-MB-231 | C888(0.96) | LDD1458 | [17] |
| LDCM0107 | IAA | HeLa | N.A. | LDD0221 | [12] |
| LDCM0022 | KB02 | MDA-MB-231 | C888(2.07) | LDD1374 | [17] |
| LDCM0023 | KB03 | MDA-MB-231 | C888(2.53) | LDD1701 | [3] |
| LDCM0024 | KB05 | G361 | C615(2.62); C842(1.58); C227(1.50) | LDD3311 | [18] |
| LDCM0109 | NEM | HeLa | N.A. | LDD0223 | [12] |
| LDCM0523 | Nucleophilic fragment 24b | MDA-MB-231 | C615(0.65) | LDD2116 | [3] |
| LDCM0527 | Nucleophilic fragment 26b | MDA-MB-231 | C615(0.73) | LDD2120 | [3] |
| LDCM0549 | Nucleophilic fragment 43 | MDA-MB-231 | C615(0.96) | LDD2143 | [3] |
| LDCM0131 | RA190 | MM1.R | C928(3.89); C842(3.42); C1251(3.31); C888(3.07) | LDD0304 | [19] |
| LDCM0003 | Sulforaphane | MDA-MB-231 | 2.35 | LDD0160 | [4] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transporter and channel
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Cadherin-1 (CDH1) | . | P12830 | |||
Other
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Breast cancer anti-estrogen resistance protein 1 (BCAR1) | CAS family | P56945 | |||
References
