Details of the Target
General Information of Target
| Target ID | LDTP06370 | |||||
|---|---|---|---|---|---|---|
| Target Name | Ephrin type-A receptor 7 (EPHA7) | |||||
| Gene Name | EPHA7 | |||||
| Gene ID | 2045 | |||||
| Synonyms |
EHK3; HEK11; Ephrin type-A receptor 7; EC 2.7.10.1; EPH homology kinase 3; EHK-3; EPH-like kinase 11; EK11; hEK11 |
|||||
| 3D Structure | ||||||
| Sequence |
MVFQTRYPSWIILCYIWLLRFAHTGEAQAAKEVLLLDSKAQQTELEWISSPPNGWEEISG
LDENYTPIRTYQVCQVMEPNQNNWLRTNWISKGNAQRIFVELKFTLRDCNSLPGVLGTCK ETFNLYYYETDYDTGRNIRENLYVKIDTIAADESFTQGDLGERKMKLNTEVREIGPLSKK GFYLAFQDVGACIALVSVKVYYKKCWSIIENLAIFPDTVTGSEFSSLVEVRGTCVSSAEE EAENAPRMHCSAEGEWLVPIGKCICKAGYQQKGDTCEPCGRGFYKSSSQDLQCSRCPTHS FSDKEGSSRCECEDGYYRAPSDPPYVACTRPPSAPQNLIFNINQTTVSLEWSPPADNGGR NDVTYRILCKRCSWEQGECVPCGSNIGYMPQQTGLEDNYVTVMDLLAHANYTFEVEAVNG VSDLSRSQRLFAAVSITTGQAAPSQVSGVMKERVLQRSVELSWQEPEHPNGVITEYEIKY YEKDQRERTYSTVKTKSTSASINNLKPGTVYVFQIRAFTAAGYGNYSPRLDVATLEEATG KMFEATAVSSEQNPVIIIAVVAVAGTIILVFMVFGFIIGRRHCGYSKADQEGDEELYFHF KFPGTKTYIDPETYEDPNRAVHQFAKELDASCIKIERVIGAGEFGEVCSGRLKLPGKRDV AVAIKTLKVGYTEKQRRDFLCEASIMGQFDHPNVVHLEGVVTRGKPVMIVIEFMENGALD AFLRKHDGQFTVIQLVGMLRGIAAGMRYLADMGYVHRDLAARNILVNSNLVCKVSDFGLS RVIEDDPEAVYTTTGGKIPVRWTAPEAIQYRKFTSASDVWSYGIVMWEVMSYGERPYWDM SNQDVIKAIEEGYRLPAPMDCPAGLHQLMLDCWQKERAERPKFEQIVGILDKMIRNPNSL KTPLGTCSRPISPLLDQNTPDFTTFCSVGEWLQAIKMERYKDNFTAAGYNSLESVARMTI EDVMSLGITLVGHQKKIMSSIQTMRAQMLHLHGTGIQV |
|||||
| Target Type |
Literature-reported
|
|||||
| Target Bioclass |
Enzyme
|
|||||
| Family |
Protein kinase superfamily, Tyr protein kinase family, Ephrin receptor subfamily
|
|||||
| Subcellular location |
Cell membrane
|
|||||
| Function |
Receptor tyrosine kinase which binds promiscuously GPI-anchored ephrin-A family ligands residing on adjacent cells, leading to contact-dependent bidirectional signaling into neighboring cells. The signaling pathway downstream of the receptor is referred to as forward signaling while the signaling pathway downstream of the ephrin ligand is referred to as reverse signaling. Among GPI-anchored ephrin-A ligands, EFNA5 is a cognate/functional ligand for EPHA7 and their interaction regulates brain development modulating cell-cell adhesion and repulsion. Has a repellent activity on axons and is for instance involved in the guidance of corticothalamic axons and in the proper topographic mapping of retinal axons to the colliculus. May also regulate brain development through a caspase(CASP3)-dependent proapoptotic activity. Forward signaling may result in activation of components of the ERK signaling pathway including MAP2K1, MAP2K2, MAPK1 and MAPK3 which are phosphorylated upon activation of EPHA7.
|
|||||
| TTD ID | ||||||
| Uniprot ID | ||||||
| DrugMap ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| 22RV1 | Insertion: p.E627GfsTer8 | . | |||
| AU565 | SNV: p.R366T | . | |||
| CCLP1 | SNV: p.R529G | DBIA Probe Info | |||
| CFPAC1 | SNV: p.Q675K | . | |||
| EFO27 | SNV: p.A213V | . | |||
| HS936T | SNV: p.D362N; p.G579R | . | |||
| IGR1 | SNV: p.K203Q | . | |||
| KASUMI1 | SNV: p.M893R | . | |||
| KPL1 | SNV: p.V418L | . | |||
| MCF7 | SNV: p.V418L | . | |||
| MEWO | SNV: p.E162K; p.S462F; p.G905E | . | |||
| MOLM13 | SNV: p.C250Y | . | |||
| NCIH1793 | SNV: p.W55C | . | |||
| NCIH2172 | SNV: p.I339N | . | |||
| P31FUJ | Substitution: p.G656V | . | |||
| SNU1 | SNV: p.K976E | . | |||
| SNU5 | SNV: p.A663V | . | |||
| SSP25 | SNV: p.G394V | . | |||
| TE10 | SNV: p.E56Ter | . | |||
| TYKNU | SNV: p.M893R | . | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
OPA-S-S-alkyne Probe Info |
 |
K272(2.33) | LDD3494 | [1] | |
|
DBIA Probe Info |
 |
C296(1.79) | LDD3406 | [2] | |
|
ATP probe Probe Info |
 |
N.A. | LDD0199 | [3] | |
|
IA-alkyne Probe Info |
 |
C772(0.00); C648(0.00); C583(0.00); C681(0.00) | LDD0165 | [4] | |
|
IPM Probe Info |
 |
C772(0.00); C632(0.00) | LDD2156 | [5] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C170 Probe Info |
 |
16.00 | LDD1850 | [6] | |
|
C197 Probe Info |
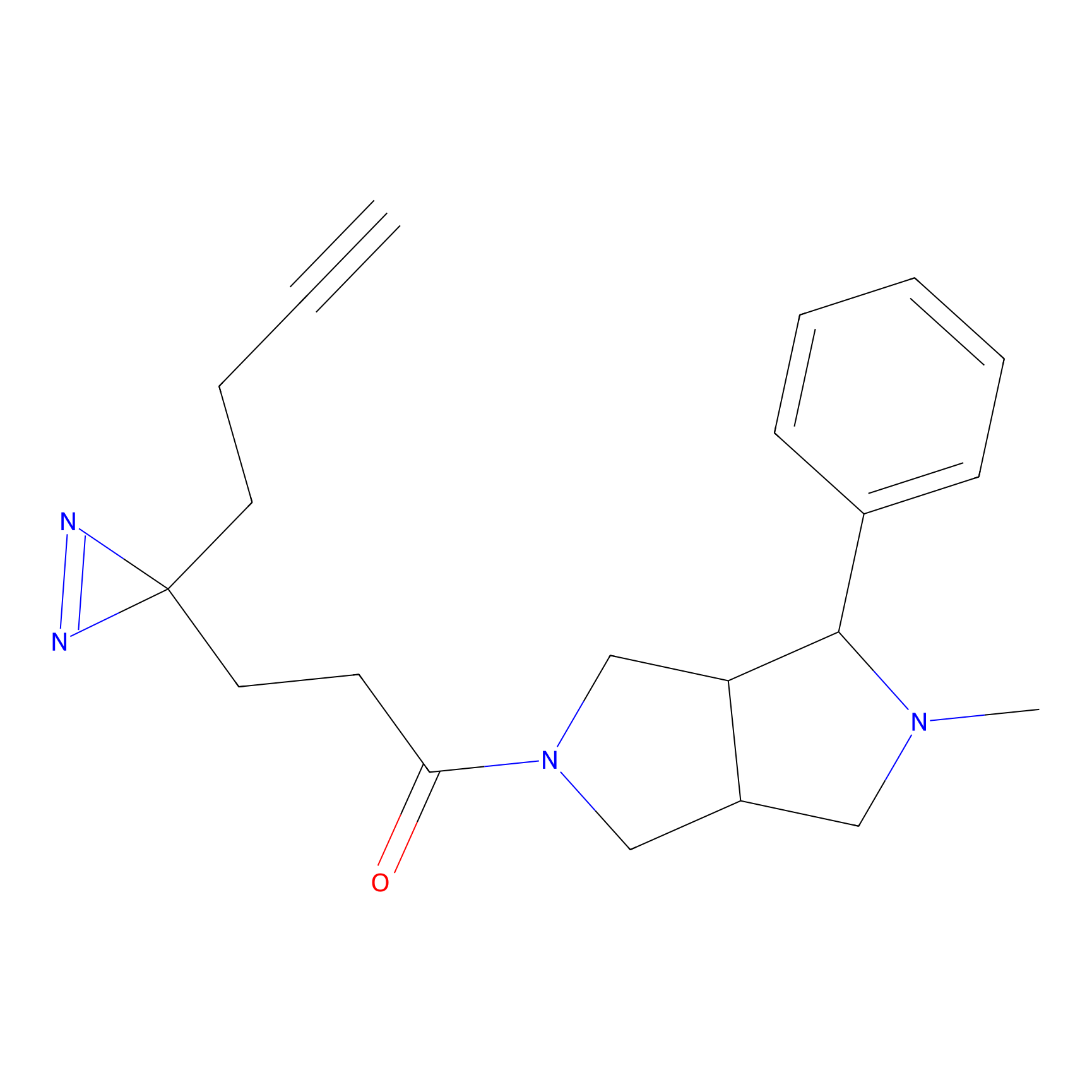 |
6.11 | LDD1873 | [6] | |
|
C208 Probe Info |
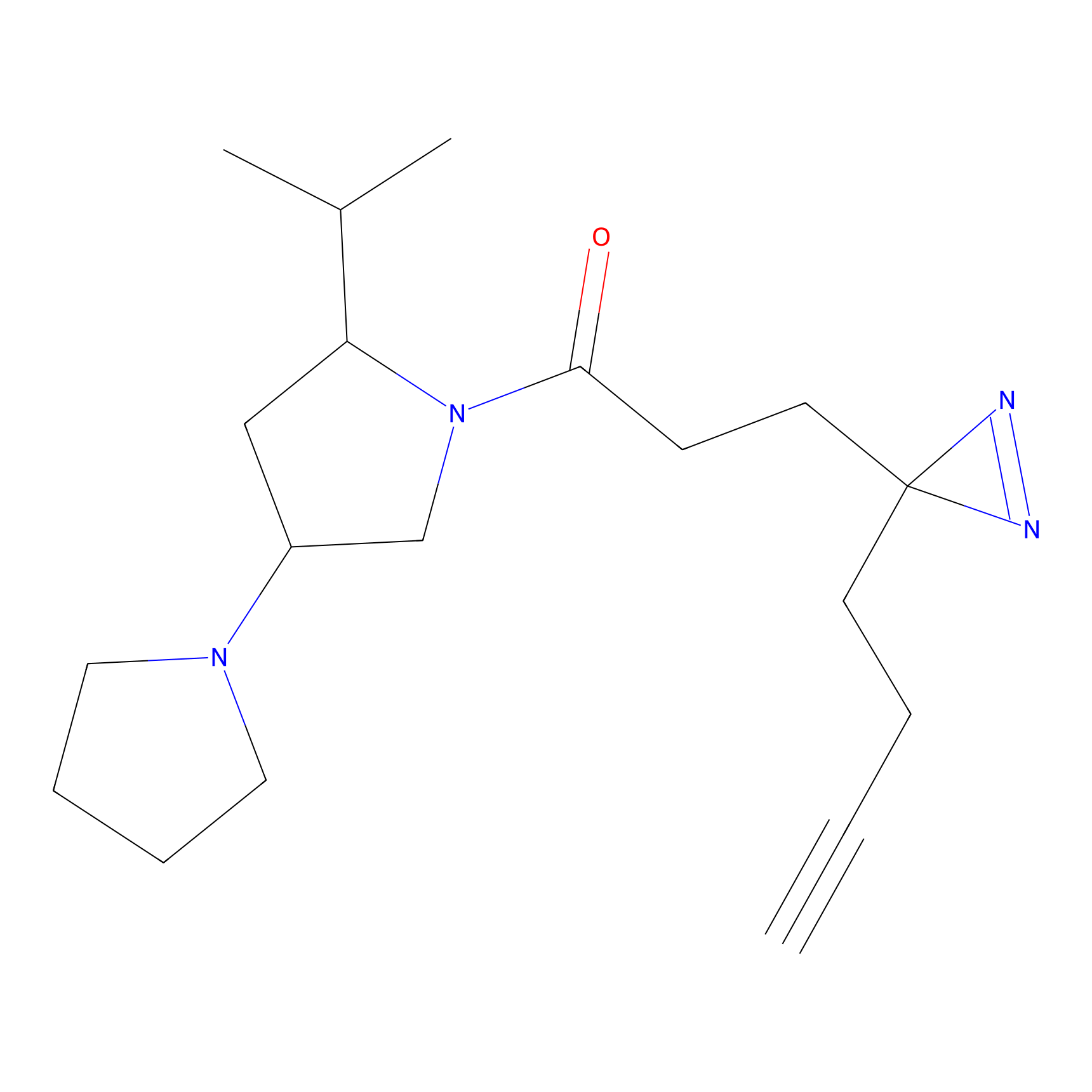 |
6.54 | LDD1883 | [6] | |
|
C237 Probe Info |
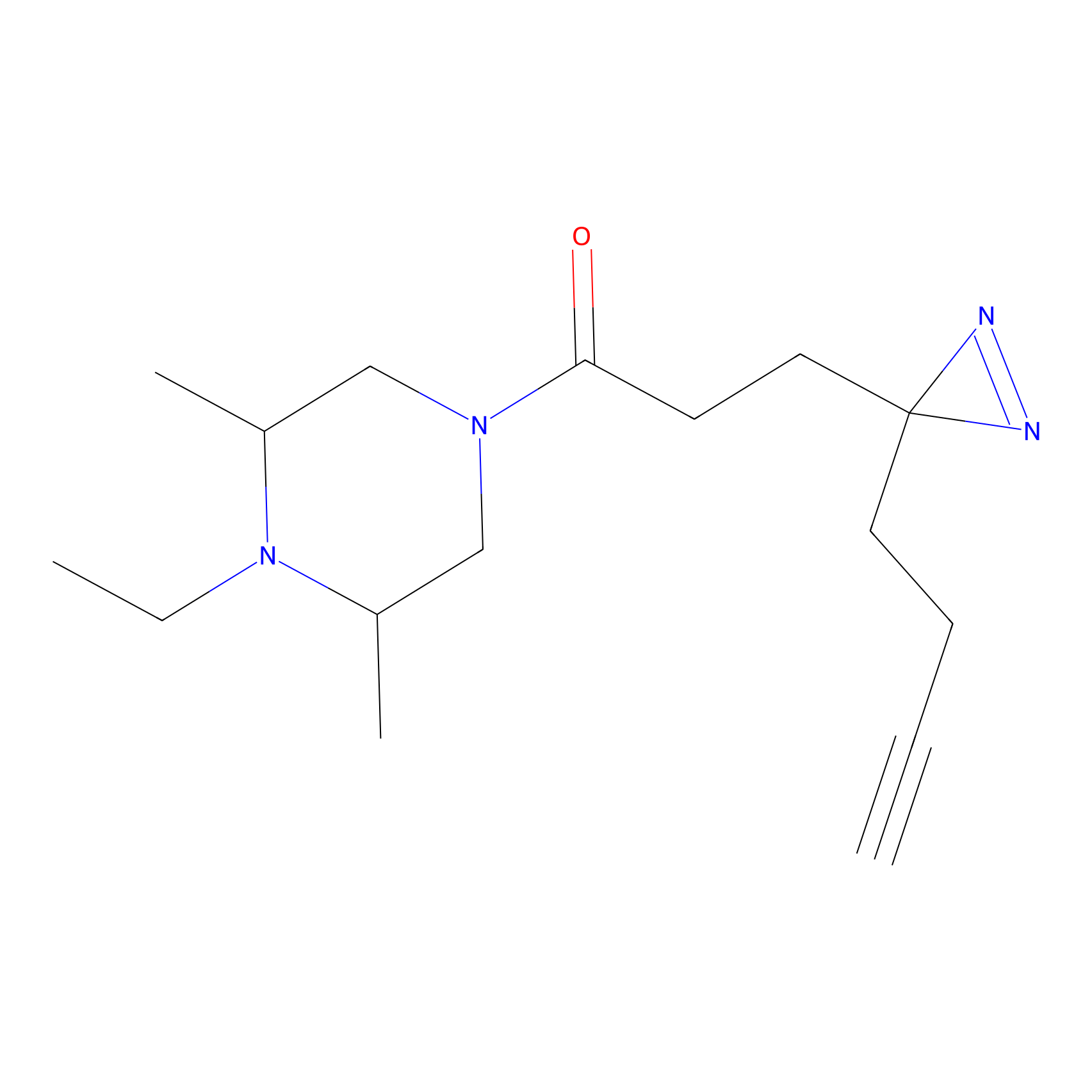 |
8.63 | LDD1910 | [6] | |
|
C310 Probe Info |
 |
30.27 | LDD1977 | [6] | |
|
C348 Probe Info |
 |
20.11 | LDD2009 | [6] | |
|
C349 Probe Info |
 |
13.55 | LDD2010 | [6] | |
|
C350 Probe Info |
 |
36.25 | LDD2011 | [6] | |
Competitor(s) Related to This Target
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
The Drug(s) Related To This Target
Approved
| Drug Name | Drug Type | External ID | |||
|---|---|---|---|---|---|
| Fostamatinib | Small molecular drug | DB12010 | |||
Investigative
References
