Details of the Target
General Information of Target
| Target ID | LDTP02959 | |||||
|---|---|---|---|---|---|---|
| Target Name | Protein SON (SON) | |||||
| Gene Name | SON | |||||
| Gene ID | 6651 | |||||
| Synonyms |
C21orf50; DBP5; KIAA1019; NREBP; Protein SON; Bax antagonist selected in saccharomyces 1; BASS1; Negative regulatory element-binding protein; NRE-binding protein; Protein DBP-5; SON3 |
|||||
| 3D Structure | ||||||
| Sequence |
MATNIEQIFRSFVVSKFREIQQELSSGRNEGQLNGETNTPIEGNQAGDAAASARSLPNEE
IVQKIEEVLSGVLDTELRYKPDLKEGSRKSRCVSVQTDPTDEIPTKKSKKHKKHKNKKKK KKKEKEKKYKRQPEESESKTKSHDDGNIDLESDSFLKFDSEPSAVALELPTRAFGPSETN ESPAVVLEPPVVSMEVSEPHILETLKPATKTAELSVVSTSVISEQSEQSVAVMPEPSMTK ILDSFAAAPVPTTTLVLKSSEPVVTMSVEYQMKSVLKSVESTSPEPSKIMLVEPPVAKVL EPSETLVVSSETPTEVYPEPSTSTTMDFPESSAIEALRLPEQPVDVPSEIADSSMTRPQE LPELPKTTALELQESSVASAMELPGPPATSMPELQGPPVTPVLELPGPSATPVPELPGPL STPVPELPGPPATAVPELPGPSVTPVPQLSQELPGLPAPSMGLEPPQEVPEPPVMAQELP GLPLVTAAVELPEQPAVTVAMELTEQPVTTTELEQPVGMTTVEHPGHPEVTTATGLLGQP EATMVLELPGQPVATTALELPGQPSVTGVPELPGLPSATRALELSGQPVATGALELPGPL MAAGALEFSGQSGAAGALELLGQPLATGVLELPGQPGAPELPGQPVATVALEISVQSVVT TSELSTMTVSQSLEVPSTTALESYNTVAQELPTTLVGETSVTVGVDPLMAPESHILASNT METHILASNTMDSQMLASNTMDSQMLASNTMDSQMLASSTMDSQMLATSSMDSQMLATSS MDSQMLATSTMDSQMLATSSMDSQMLATSSMDSQMLATSSMDSQMLATSSMDSQMLATST MDSQMLATSTMDSQMLATSSMDSQMLASGTMDSQMLASGTMDAQMLASGTMDAQMLASST QDSAMLGSKSPDPYRLAQDPYRLAQDPYRLGHDPYRLGHDAYRLGQDPYRLGHDPYRLTP DPYRMSPRPYRIAPRSYRIAPRPYRLAPRPLMLASRRSMMMSYAAERSMMSSYERSMMSY ERSMMSPMAERSMMSAYERSMMSAYERSMMSPMAERSMMSAYERSMMSAYERSMMSPMAD RSMMSMGADRSMMSSYSAADRSMMSSYSAADRSMMSSYTADRSMMSMAADSYTDSYTDTY TEAYMVPPLPPEEPPTMPPLPPEEPPMTPPLPPEEPPEGPALPTEQSALTAENTWPTEVP SSPSEESVSQPEPPVSQSEISEPSAVPTDYSVSASDPSVLVSEAAVTVPEPPPEPESSIT LTPVESAVVAEEHEVVPERPVTCMVSETPAMSAEPTVLASEPPVMSETAETFDSMRASGH VASEVSTSLLVPAVTTPVLAESILEPPAMAAPESSAMAVLESSAVTVLESSTVTVLESST VTVLEPSVVTVPEPPVVAEPDYVTIPVPVVSALEPSVPVLEPAVSVLQPSMIVSEPSVSV QESTVTVSEPAVTVSEQTQVIPTEVAIESTPMILESSIMSSHVMKGINLSSGDQNLAPEI GMQEIALHSGEEPHAEEHLKGDFYESEHGINIDLNINNHLIAKEMEHNTVCAAGTSPVGE IGEEKILPTSETKQRTVLDTYPGVSEADAGETLSSTGPFALEPDATGTSKGIEFTTASTL SLVNKYDVDLSLTTQDTEHDMVISTSPSGGSEADIEGPLPAKDIHLDLPSNNNLVSKDTE EPLPVKESDQTLAALLSPKESSGGEKEVPPPPKETLPDSGFSANIEDINEADLVRPLLPK DMERLTSLRAGIEGPLLASDVGRDRSAASPVVSSMPERASESSSEEKDDYEIFVKVKDTH EKSKKNKNRDKGEKEKKRDSSLRSRSKRSKSSEHKSRKRTSESRSRARKRSSKSKSHRSQ TRSRSRSRRRRRSSRSRSKSRGRRSVSKEKRKRSPKHRSKSRERKRKRSSSRDNRKTVRA RSRTPSRRSRSHTPSRRRRSRSVGRRRSFSISPSRRSRTPSRRSRTPSRRSRTPSRRSRT PSRRSRTPSRRSRTPSRRRRSRSVVRRRSFSISPVRLRRSRTPLRRRFSRSPIRRKRSRS SERGRSPKRLTDLDKAQLLEIAKANAAAMCAKAGVPLPPNLKPAPPPTIEEKVAKKSGGA TIEELTEKCKQIAQSKEDDDVIVNKPHVSDEEEEEPPFYHHPFKLSEPKPIFFNLNIAAA KPTPPKSQVTLTKEFPVSSGSQHRKKEADSVYGEWVPVEKNGEENKDDDNVFSSNLPSEP VDISTAMSERALAQKRLSENAFDLEAMSMLNRAQERIDAWAQLNSIPGQFTGSTGVQVLT QEQLANTGAQAWIKKDQFLRAAPVTGGMGAVLMRKMGWREGEGLGKNKEGNKEPILVDFK TDRKGLVAVGERAQKRSGNFSAAMKDLSGKHPVSALMEICNKRRWQPPEFLLVHDSGPDH RKHFLFRVLRNGALTRPNCMFFLNRY |
|||||
| Target Bioclass |
Transporter and channel
|
|||||
| Subcellular location |
Nucleus speckle
|
|||||
| Function |
RNA-binding protein that acts as a mRNA splicing cofactor by promoting efficient splicing of transcripts that possess weak splice sites. Specifically promotes splicing of many cell-cycle and DNA-repair transcripts that possess weak splice sites, such as TUBG1, KATNB1, TUBGCP2, AURKB, PCNT, AKT1, RAD23A, and FANCG. Probably acts by facilitating the interaction between Serine/arginine-rich proteins such as SRSF2 and the RNA polymerase II. Also binds to DNA; binds to the consensus DNA sequence: 5'-GA[GT]AN[CG][AG]CC-3'. May indirectly repress hepatitis B virus (HBV) core promoter activity and transcription of HBV genes and production of HBV virions. Essential for correct RNA splicing of multiple genes critical for brain development, neuronal migration and metabolism, including TUBG1, FLNA, PNKP, WDR62, PSMD3, PCK2, PFKL, IDH2, and ACY1.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| DEL | SNV: p.P1698R | DBIA Probe Info | |||
| HCT15 | SNV: p.G2288D | DBIA Probe Info | |||
| HEC1B | SNV: p.T1374I | . | |||
| Ishikawa (Heraklio) 02 ER | SNV: p.Q477H | DBIA Probe Info | |||
| JURKAT | SNV: p.L1745I | Compound 10 Probe Info | |||
| KMRC1 | SNV: p.T678M | DBIA Probe Info | |||
| KYSE30 | Deletion: p.S770_S789del | DBIA Probe Info | |||
| L428 | SNV: p.K1787N | . | |||
| LN229 | SNV: p.P1711R | DBIA Probe Info | |||
| LNCaP clone FGC | SNV: p.G2268S | DBIA Probe Info | |||
| MDST8 | SNV: p.V185L | DBIA Probe Info | |||
| MFE319 | SNV: p.A1542V | DBIA Probe Info | |||
| REH | SNV: p.A458P; p.S1102F | DBIA Probe Info | |||
| RL952 | SNV: p.P1169Q; p.R1963H | DBIA Probe Info | |||
| SKMEL24 | SNV: p.T798I | DBIA Probe Info | |||
| SW480 | SNV: p.M1053I | . | |||
| TCCSUP | SNV: p.G1650A | DBIA Probe Info | |||
| TE11 | SNV: p.D352V | DBIA Probe Info | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
15.00 | LDD0402 | [1] | |
|
P3 Probe Info |
 |
1.71 | LDD0450 | [2] | |
|
P8 Probe Info |
 |
2.37 | LDD0451 | [2] | |
|
TH211 Probe Info |
 |
Y942(15.28); Y935(5.68) | LDD0260 | [3] | |
|
TH216 Probe Info |
 |
Y942(20.00); Y935(19.46) | LDD0259 | [3] | |
|
STPyne Probe Info |
 |
K16(6.42); K1802(10.00); K2055(4.94); K2108(1.64) | LDD0277 | [4] | |
|
BTD Probe Info |
 |
C2070(1.05) | LDD1700 | [5] | |
|
AHL-Pu-1 Probe Info |
 |
C2380(2.66); C1551(4.62) | LDD0169 | [6] | |
|
DBIA Probe Info |
 |
C2380(44.16) | LDD0209 | [7] | |
|
ATP probe Probe Info |
 |
N.A. | LDD0199 | [8] | |
|
1d-yne Probe Info |
 |
N.A. | LDD0358 | [9] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
C2070(0.00); C1551(0.00); C2380(0.00); C92(0.00) | LDD0038 | [10] | |
|
IA-alkyne Probe Info |
 |
C92(0.00); C2070(0.00) | LDD0036 | [10] | |
|
IPIAA_L Probe Info |
 |
N.A. | LDD0031 | [11] | |
|
Lodoacetamide azide Probe Info |
 |
C2070(0.00); C92(0.00); C1551(0.00); C2380(0.00) | LDD0037 | [10] | |
|
JW-RF-010 Probe Info |
 |
N.A. | LDD0026 | [12] | |
|
TFBX Probe Info |
 |
C92(0.00); C2070(0.00) | LDD0027 | [12] | |
|
WYneN Probe Info |
 |
C92(0.00); C2070(0.00) | LDD0021 | [13] | |
|
WYneO Probe Info |
 |
N.A. | LDD0022 | [13] | |
|
Compound 10 Probe Info |
 |
C2070(0.00); C92(0.00) | LDD2216 | [14] | |
|
Compound 11 Probe Info |
 |
C2070(0.00); C92(0.00) | LDD2213 | [14] | |
|
ENE Probe Info |
 |
N.A. | LDD0006 | [13] | |
|
IPM Probe Info |
 |
C2070(0.00); C2380(0.00); C1551(0.00); C92(0.00) | LDD0005 | [13] | |
|
NPM Probe Info |
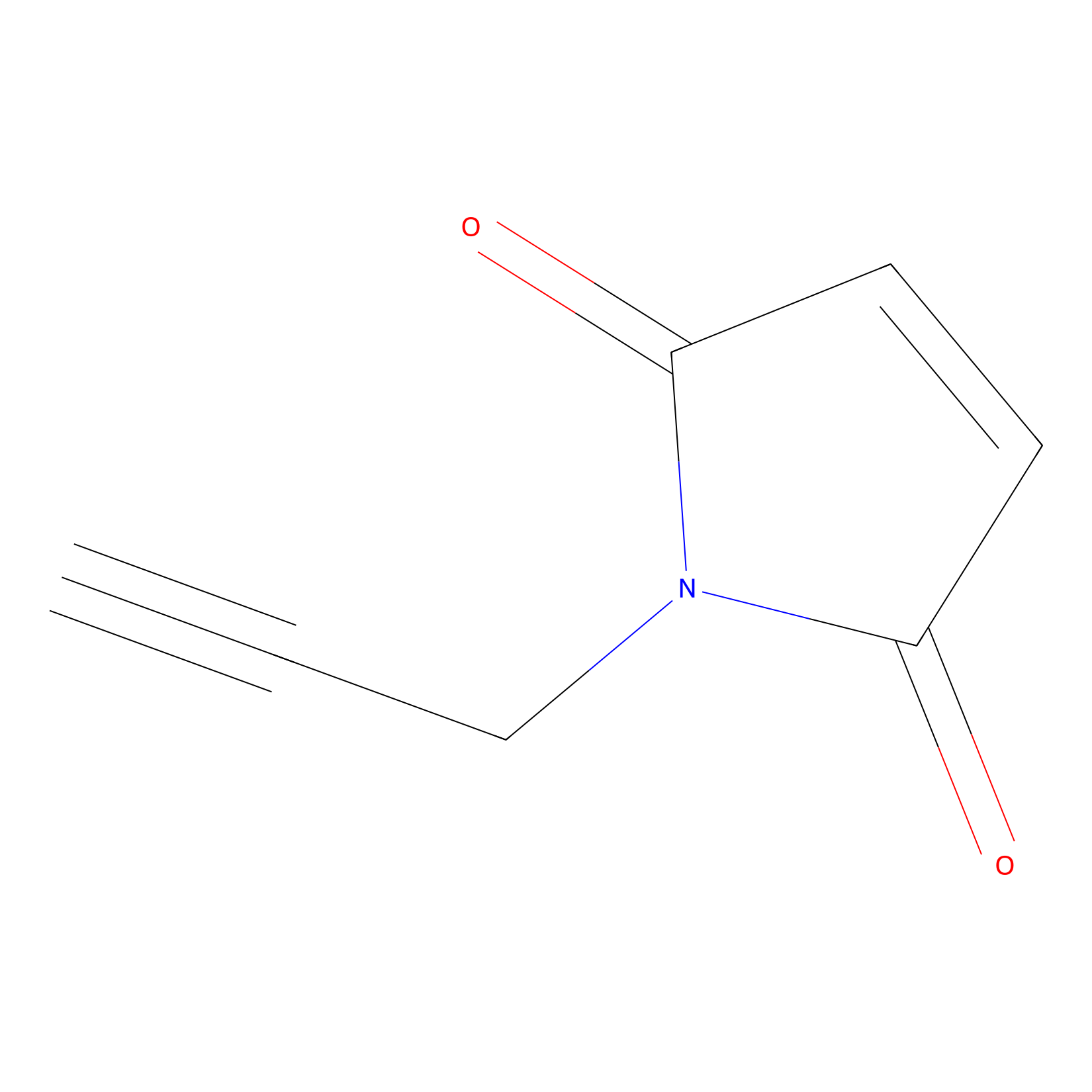 |
N.A. | LDD0016 | [13] | |
|
SF Probe Info |
 |
Y949(0.00); Y928, Y921(0.00) | LDD0028 | [15] | |
|
VSF Probe Info |
 |
C1551(0.00); C92(0.00) | LDD0007 | [13] | |
|
Phosphinate-6 Probe Info |
 |
C92(0.00); C2070(0.00); C1551(0.00) | LDD0018 | [16] | |
|
1c-yne Probe Info |
 |
N.A. | LDD0228 | [9] | |
|
Acrolein Probe Info |
 |
C1551(0.00); C2070(0.00) | LDD0217 | [17] | |
|
Crotonaldehyde Probe Info |
 |
N.A. | LDD0219 | [17] | |
|
Methacrolein Probe Info |
 |
C1551(0.00); C2070(0.00) | LDD0218 | [17] | |
|
W1 Probe Info |
 |
C2070(0.00); C2380(0.00) | LDD0236 | [18] | |
|
NAIA_5 Probe Info |
 |
N.A. | LDD2223 | [19] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
FFF probe11 Probe Info |
 |
5.78 | LDD0472 | [20] | |
|
FFF probe13 Probe Info |
 |
20.00 | LDD0475 | [20] | |
|
FFF probe3 Probe Info |
 |
6.11 | LDD0465 | [20] | |
|
STS-1 Probe Info |
 |
N.A. | LDD0137 | [21] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0519 | 1-(6-methoxy-3,4-dihydroquinolin-1(2H)-yl)-2-nitroethan-1-one | MDA-MB-231 | C2109(1.07) | LDD2112 | [5] |
| LDCM0502 | 1-(Cyanoacetyl)piperidine | MDA-MB-231 | C92(0.71) | LDD2095 | [5] |
| LDCM0537 | 2-Cyano-N,N-dimethylacetamide | MDA-MB-231 | C2070(0.95) | LDD2130 | [5] |
| LDCM0524 | 2-Cyano-N-(2-morpholin-4-yl-ethyl)-acetamide | MDA-MB-231 | C2070(0.86); C2380(0.98) | LDD2117 | [5] |
| LDCM0558 | 2-Cyano-N-phenylacetamide | MDA-MB-231 | C2070(1.13) | LDD2152 | [5] |
| LDCM0510 | 3-(4-(Hydroxydiphenylmethyl)piperidin-1-yl)-3-oxopropanenitrile | MDA-MB-231 | C2109(1.09) | LDD2103 | [5] |
| LDCM0539 | 3-(4-Isopropylpiperazin-1-yl)-3-oxopropanenitrile | MDA-MB-231 | C2070(0.73) | LDD2132 | [5] |
| LDCM0026 | 4SU-RNA+native RNA | HEK-293T | C2380(2.66); C1551(4.62) | LDD0169 | [6] |
| LDCM0520 | AKOS000195272 | MDA-MB-231 | C2070(1.16) | LDD2113 | [5] |
| LDCM0156 | Aniline | NCI-H1299 | 12.96 | LDD0403 | [1] |
| LDCM0498 | BS-3668 | MDA-MB-231 | C2070(0.42) | LDD2091 | [5] |
| LDCM0108 | Chloroacetamide | HeLa | C1551(0.00); C2070(0.00); H953(0.00) | LDD0222 | [17] |
| LDCM0632 | CL-Sc | Hep-G2 | C2070(1.53); C92(0.74); C92(0.69); C2109(0.69) | LDD2227 | [19] |
| LDCM0634 | CY-0357 | Hep-G2 | C92(3.34) | LDD2228 | [19] |
| LDCM0198 | Dimethyl Fumarate(DMF) | T cell | C92(11.68) | LDD0526 | [22] |
| LDCM0213 | Electrophilic fragment 2 | MDA-MB-231 | C92(1.87); C1551(1.15); C2070(1.15) | LDD1702 | [5] |
| LDCM0625 | F8 | Ramos | C92(0.94); C2070(1.03) | LDD2187 | [23] |
| LDCM0572 | Fragment10 | Ramos | C92(2.96); C2070(1.19) | LDD2189 | [23] |
| LDCM0573 | Fragment11 | Ramos | C92(3.55) | LDD2190 | [23] |
| LDCM0574 | Fragment12 | Ramos | C92(1.27); C2070(1.65) | LDD2191 | [23] |
| LDCM0575 | Fragment13 | Ramos | C92(1.02); C2070(0.77) | LDD2192 | [23] |
| LDCM0576 | Fragment14 | Ramos | C92(1.31); C2070(0.77) | LDD2193 | [23] |
| LDCM0579 | Fragment20 | Ramos | C92(1.13); C2070(0.78) | LDD2194 | [23] |
| LDCM0580 | Fragment21 | Ramos | C92(0.79); C2070(0.81) | LDD2195 | [23] |
| LDCM0582 | Fragment23 | Ramos | C92(0.95); C2070(1.55) | LDD2196 | [23] |
| LDCM0578 | Fragment27 | Ramos | C92(0.87); C2070(0.83) | LDD2197 | [23] |
| LDCM0586 | Fragment28 | Ramos | C92(0.73); C2070(1.39) | LDD2198 | [23] |
| LDCM0588 | Fragment30 | Ramos | C92(1.36); C2070(1.06) | LDD2199 | [23] |
| LDCM0589 | Fragment31 | Ramos | C92(0.85); C2070(1.26) | LDD2200 | [23] |
| LDCM0590 | Fragment32 | Ramos | C92(1.85); C2070(2.98) | LDD2201 | [23] |
| LDCM0468 | Fragment33 | Ramos | C92(1.21); C2070(1.09) | LDD2202 | [23] |
| LDCM0596 | Fragment38 | Ramos | C92(0.85); C2070(1.00) | LDD2203 | [23] |
| LDCM0566 | Fragment4 | Ramos | C92(0.58); C2070(0.73) | LDD2184 | [23] |
| LDCM0610 | Fragment52 | Ramos | C92(1.46); C2070(1.01) | LDD2204 | [23] |
| LDCM0614 | Fragment56 | Ramos | C92(1.25); C2070(3.05) | LDD2205 | [23] |
| LDCM0569 | Fragment7 | Ramos | C92(1.31); C2070(1.17) | LDD2186 | [23] |
| LDCM0571 | Fragment9 | Ramos | C92(2.42); C2070(0.88) | LDD2188 | [23] |
| LDCM0107 | IAA | HeLa | N.A. | LDD0221 | [17] |
| LDCM0022 | KB02 | Ramos | C92(1.44); C2070(1.14) | LDD2182 | [23] |
| LDCM0023 | KB03 | Jurkat | C2380(44.16) | LDD0209 | [7] |
| LDCM0024 | KB05 | COLO792 | C2420(2.79) | LDD3310 | [24] |
| LDCM0509 | N-(4-bromo-3,5-dimethylphenyl)-2-nitroacetamide | MDA-MB-231 | C2109(1.01) | LDD2102 | [5] |
| LDCM0528 | N-(4-bromophenyl)-2-cyano-N-phenylacetamide | MDA-MB-231 | C2070(0.59) | LDD2121 | [5] |
| LDCM0109 | NEM | HeLa | N.A. | LDD0223 | [17] |
| LDCM0496 | Nucleophilic fragment 11a | MDA-MB-231 | C2070(0.55) | LDD2089 | [5] |
| LDCM0500 | Nucleophilic fragment 13a | MDA-MB-231 | C2070(0.98) | LDD2093 | [5] |
| LDCM0503 | Nucleophilic fragment 14b | MDA-MB-231 | C2070(0.81); C2109(0.98) | LDD2096 | [5] |
| LDCM0505 | Nucleophilic fragment 15b | MDA-MB-231 | C2109(1.20) | LDD2098 | [5] |
| LDCM0506 | Nucleophilic fragment 16a | MDA-MB-231 | C2070(0.79) | LDD2099 | [5] |
| LDCM0512 | Nucleophilic fragment 19a | MDA-MB-231 | C2109(1.18) | LDD2105 | [5] |
| LDCM0513 | Nucleophilic fragment 19b | MDA-MB-231 | C2109(0.91) | LDD2106 | [5] |
| LDCM0514 | Nucleophilic fragment 20a | MDA-MB-231 | C2070(0.99) | LDD2107 | [5] |
| LDCM0515 | Nucleophilic fragment 20b | MDA-MB-231 | C2109(0.83) | LDD2108 | [5] |
| LDCM0516 | Nucleophilic fragment 21a | MDA-MB-231 | C2070(0.82) | LDD2109 | [5] |
| LDCM0518 | Nucleophilic fragment 22a | MDA-MB-231 | C2070(0.74); C2109(0.93) | LDD2111 | [5] |
| LDCM0523 | Nucleophilic fragment 24b | MDA-MB-231 | C2070(0.93) | LDD2116 | [5] |
| LDCM0525 | Nucleophilic fragment 25b | MDA-MB-231 | C2070(0.70) | LDD2118 | [5] |
| LDCM0526 | Nucleophilic fragment 26a | MDA-MB-231 | C2070(2.11); C2109(2.10) | LDD2119 | [5] |
| LDCM0529 | Nucleophilic fragment 27b | MDA-MB-231 | C92(0.73); C2070(0.62) | LDD2122 | [5] |
| LDCM0530 | Nucleophilic fragment 28a | MDA-MB-231 | C2070(0.79) | LDD2123 | [5] |
| LDCM0531 | Nucleophilic fragment 28b | MDA-MB-231 | C2070(0.92) | LDD2124 | [5] |
| LDCM0532 | Nucleophilic fragment 29a | MDA-MB-231 | C2070(0.87); C2109(0.75) | LDD2125 | [5] |
| LDCM0533 | Nucleophilic fragment 29b | MDA-MB-231 | C2070(0.70) | LDD2126 | [5] |
| LDCM0534 | Nucleophilic fragment 30a | MDA-MB-231 | C2070(0.70) | LDD2127 | [5] |
| LDCM0535 | Nucleophilic fragment 30b | MDA-MB-231 | C2070(1.24) | LDD2128 | [5] |
| LDCM0536 | Nucleophilic fragment 31 | MDA-MB-231 | C2070(0.95) | LDD2129 | [5] |
| LDCM0540 | Nucleophilic fragment 35 | MDA-MB-231 | C2070(0.86) | LDD2133 | [5] |
| LDCM0541 | Nucleophilic fragment 36 | MDA-MB-231 | C2070(1.19) | LDD2134 | [5] |
| LDCM0542 | Nucleophilic fragment 37 | MDA-MB-231 | C2070(0.93); C2109(1.00) | LDD2135 | [5] |
| LDCM0543 | Nucleophilic fragment 38 | MDA-MB-231 | C92(0.85); C2070(0.98) | LDD2136 | [5] |
| LDCM0544 | Nucleophilic fragment 39 | MDA-MB-231 | C2070(0.88) | LDD2137 | [5] |
| LDCM0211 | Nucleophilic fragment 3b | MDA-MB-231 | C2070(1.05) | LDD1700 | [5] |
| LDCM0546 | Nucleophilic fragment 40 | MDA-MB-231 | C2070(0.67); C2109(0.62) | LDD2140 | [5] |
| LDCM0547 | Nucleophilic fragment 41 | MDA-MB-231 | C92(0.74); C2109(1.04) | LDD2141 | [5] |
| LDCM0549 | Nucleophilic fragment 43 | MDA-MB-231 | C2070(1.14) | LDD2143 | [5] |
| LDCM0550 | Nucleophilic fragment 5a | MDA-MB-231 | C2070(1.73) | LDD2144 | [5] |
| LDCM0552 | Nucleophilic fragment 6a | MDA-MB-231 | C2070(0.86); C2109(1.13) | LDD2146 | [5] |
| LDCM0554 | Nucleophilic fragment 7a | MDA-MB-231 | C2070(0.75) | LDD2148 | [5] |
| LDCM0555 | Nucleophilic fragment 7b | MDA-MB-231 | C92(0.57); C2070(0.64) | LDD2149 | [5] |
| LDCM0627 | NUDT7-COV-1 | HEK-293T | C2070(0.88) | LDD2206 | [25] |
| LDCM0628 | OTUB2-COV-1 | HEK-293T | C92(1.32); C2070(0.71) | LDD2207 | [25] |
| LDCM0131 | RA190 | MM1.R | C92(1.20) | LDD0304 | [26] |
The Interaction Atlas With This Target
References
