Details of the Target
General Information of Target
| Target ID | LDTP02671 | |||||
|---|---|---|---|---|---|---|
| Target Name | Sodium/potassium-transporting ATPase subunit alpha-3 (ATP1A3) | |||||
| Gene Name | ATP1A3 | |||||
| Gene ID | 478 | |||||
| Synonyms |
Sodium/potassium-transporting ATPase subunit alpha-3; Na(+)/K(+) ATPase alpha-3 subunit; EC 7.2.2.13; Na(+)/K(+) ATPase alpha(III) subunit; Sodium pump subunit alpha-3 |
|||||
| 3D Structure | ||||||
| Sequence |
MGDKKDDKDSPKKNKGKERRDLDDLKKEVAMTEHKMSVEEVCRKYNTDCVQGLTHSKAQE
ILARDGPNALTPPPTTPEWVKFCRQLFGGFSILLWIGAILCFLAYGIQAGTEDDPSGDNL YLGIVLAAVVIITGCFSYYQEAKSSKIMESFKNMVPQQALVIREGEKMQVNAEEVVVGDL VEIKGGDRVPADLRIISAHGCKVDNSSLTGESEPQTRSPDCTHDNPLETRNITFFSTNCV EGTARGVVVATGDRTVMGRIATLASGLEVGKTPIAIEIEHFIQLITGVAVFLGVSFFILS LILGYTWLEAVIFLIGIIVANVPEGLLATVTVCLTLTAKRMARKNCLVKNLEAVETLGST STICSDKTGTLTQNRMTVAHMWFDNQIHEADTTEDQSGTSFDKSSHTWVALSHIAGLCNR AVFKGGQDNIPVLKRDVAGDASESALLKCIELSSGSVKLMRERNKKVAEIPFNSTNKYQL SIHETEDPNDNRYLLVMKGAPERILDRCSTILLQGKEQPLDEEMKEAFQNAYLELGGLGE RVLGFCHYYLPEEQFPKGFAFDCDDVNFTTDNLCFVGLMSMIDPPRAAVPDAVGKCRSAG IKVIMVTGDHPITAKAIAKGVGIISEGNETVEDIAARLNIPVSQVNPRDAKACVIHGTDL KDFTSEQIDEILQNHTEIVFARTSPQQKLIIVEGCQRQGAIVAVTGDGVNDSPALKKADI GVAMGIAGSDVSKQAADMILLDDNFASIVTGVEEGRLIFDNLKKSIAYTLTSNIPEITPF LLFIMANIPLPLGTITILCIDLGTDMVPAISLAYEAAESDIMKRQPRNPRTDKLVNERLI SMAYGQIGMIQALGGFFSYFVILAENGFLPGNLVGIRLNWDDRTVNDLEDSYGQQWTYEQ RKVVEFTCHTAFFVSIVVVQWADLIICKTRRNSVFQQGMKNKILIFGLFEETALAAFLSY CPGMDVALRMYPLKPSWWFCAFPYSFLIFVYDEIRKLILRRNPGGWVEKETYY |
|||||
| Target Bioclass |
Enzyme
|
|||||
| Family |
Cation transport ATPase (P-type) (TC 3.A.3) family, Type IIC subfamily
|
|||||
| Subcellular location |
Cell membrane
|
|||||
| Function |
This is the catalytic component of the active enzyme, which catalyzes the hydrolysis of ATP coupled with the exchange of sodium and potassium ions across the plasma membrane. This action creates the electrochemical gradient of sodium and potassium ions, providing the energy for active transport of various nutrients.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| CAL51 | SNV: p.Q157R | DBIA Probe Info | |||
| CCK81 | SNV: p.T272M | . | |||
| DU145 | SNV: p.K615N | DBIA Probe Info | |||
| GB1 | Insertion: p.N787QfsTer19 | . | |||
| HEC1 | SNV: p.I945V | . | |||
| HGC27 | SNV: p.A587E | . | |||
| HT | SNV: p.Y105C | . | |||
| KYSE180 | SNV: p.G699D | . | |||
| LN229 | SNV: p.A527V | . | |||
| LS123 | SNV: p.A98V | DBIA Probe Info | |||
| MCC13 | SNV: p.G938S Substitution: p.A244T |
. | |||
| MDST8 | SNV: p.G66R | DBIA Probe Info | |||
| MOLT4 | SNV: p.M168I | IA-alkyne Probe Info | |||
| NCIH1155 | SNV: p.R930Q | DBIA Probe Info | |||
| OVK18 | SNV: p.M579V | DBIA Probe Info | |||
| PC14 | SNV: p.K81M | DBIA Probe Info | |||
| PC9 | SNV: p.K81M | . | |||
| SNU1 | SNV: p.P273S | DBIA Probe Info | |||
| TE8 | SNV: p.E540K | . | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
DBIA Probe Info |
 |
C521(3.60); C62(1.05) | LDD3311 | [1] | |
|
AHL-Pu-1 Probe Info |
 |
C364(2.27) | LDD0169 | [2] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
C653(0.00); C449(0.00); C508(0.00); C546(0.00) | LDD0038 | [3] | |
|
IA-alkyne Probe Info |
 |
C449(0.00); C418(0.00); C49(0.00); C695(0.00) | LDD0036 | [3] | |
|
Lodoacetamide azide Probe Info |
 |
C449(0.00); C508(0.00); C418(0.00); C49(0.00) | LDD0037 | [3] | |
|
Compound 11 Probe Info |
 |
N.A. | LDD2213 | [4] | |
|
IPM Probe Info |
 |
C49(0.00); C653(0.00) | LDD2156 | [5] | |
|
Acrolein Probe Info |
 |
C364(0.00); C695(0.00); C418(0.00) | LDD0217 | [6] | |
|
Cinnamaldehyde Probe Info |
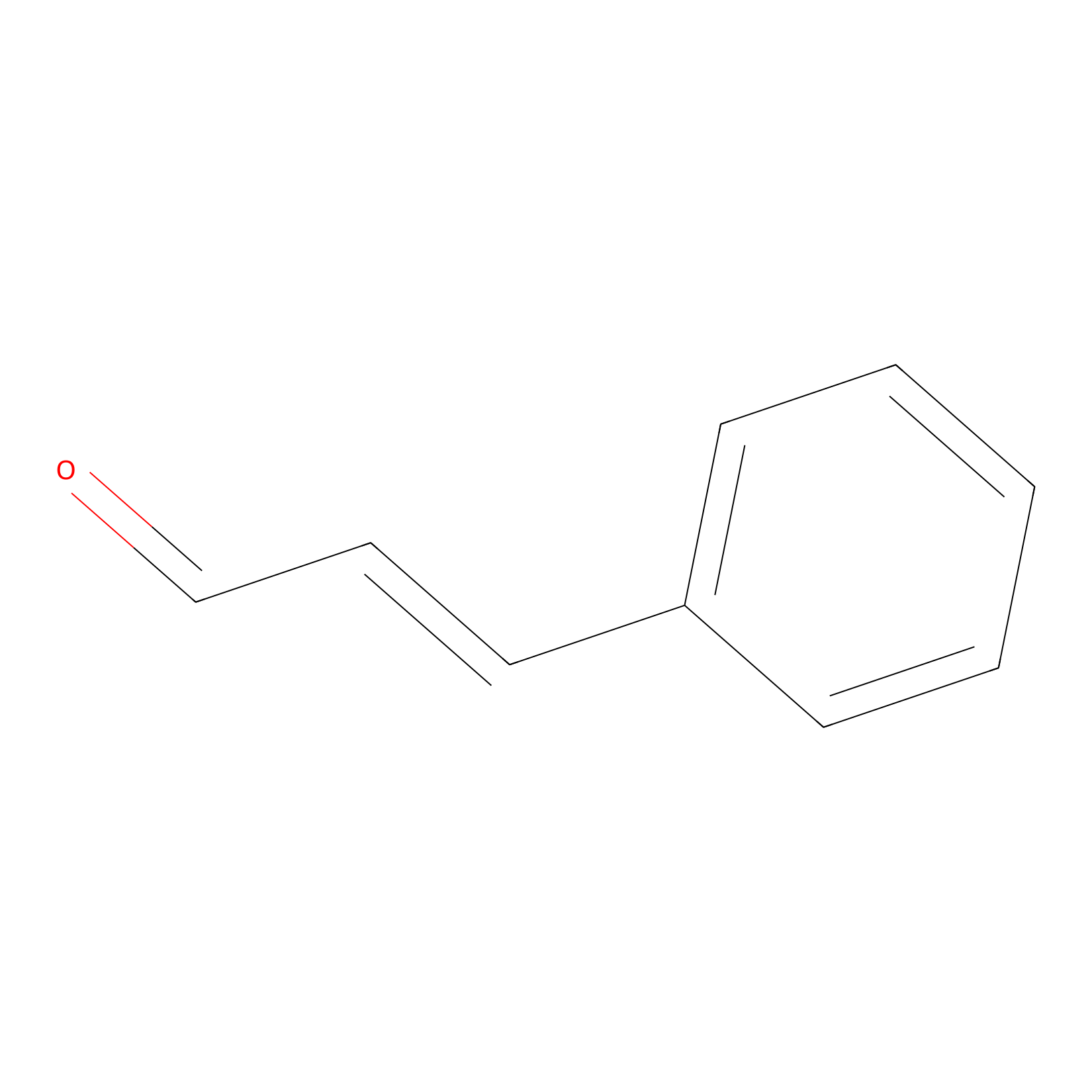 |
C418(0.00); C695(0.00) | LDD0220 | [6] | |
|
Crotonaldehyde Probe Info |
 |
C418(0.00); C364(0.00); C695(0.00) | LDD0219 | [6] | |
|
Methacrolein Probe Info |
 |
C695(0.00); C364(0.00) | LDD0218 | [6] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
FFF probe11 Probe Info |
 |
20.00 | LDD0471 | [7] | |
|
FFF probe6 Probe Info |
 |
6.77 | LDD0467 | [7] | |
|
STS-2 Probe Info |
 |
N.A. | LDD0138 | [8] | |
|
LD-F Probe Info |
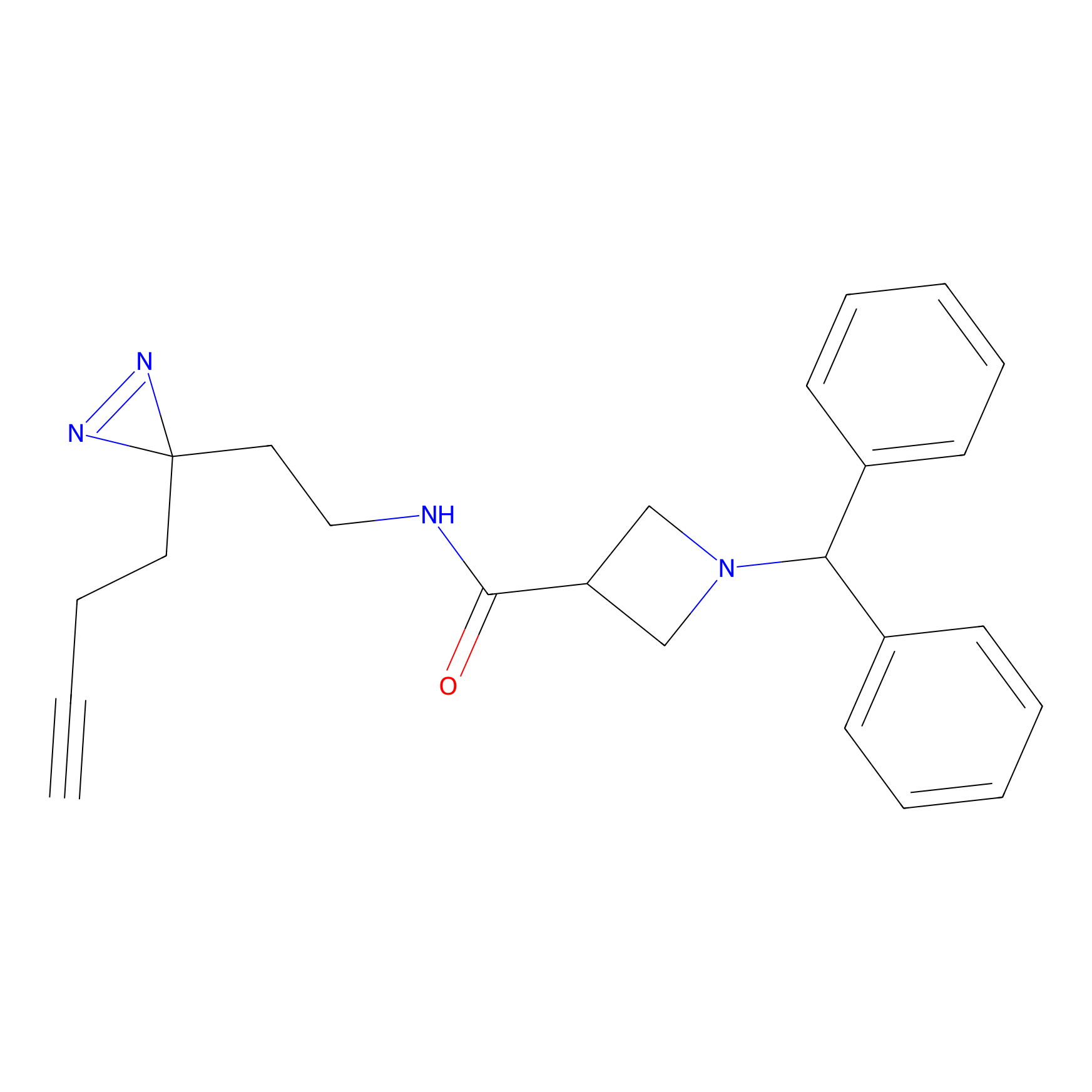 |
E352(0.00); A353(0.00); V354(0.00); E355(0.00) | LDD0015 | [9] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0026 | 4SU-RNA+native RNA | HEK-293T | C364(2.27) | LDD0169 | [2] |
| LDCM0214 | AC1 | HEK-293T | C49(1.06); C653(0.99); C221(1.06); C546(0.78) | LDD1507 | [10] |
| LDCM0215 | AC10 | HEK-293T | C49(0.99); C653(1.04); C221(0.91); C546(1.00) | LDD1508 | [10] |
| LDCM0226 | AC11 | HEK-293T | C49(0.98); C653(0.87); C221(0.99); C546(1.19) | LDD1509 | [10] |
| LDCM0237 | AC12 | HEK-293T | C49(0.94); C653(0.82); C221(1.01); C546(0.92) | LDD1510 | [10] |
| LDCM0259 | AC14 | HEK-293T | C49(1.10); C653(1.15); C546(0.96) | LDD1512 | [10] |
| LDCM0270 | AC15 | HEK-293T | C49(0.90); C653(0.92); C221(1.01); C546(0.86) | LDD1513 | [10] |
| LDCM0276 | AC17 | HEK-293T | C49(0.98); C653(1.04); C221(0.92); C546(0.88) | LDD1515 | [10] |
| LDCM0277 | AC18 | HEK-293T | C49(0.92); C653(1.00); C221(0.82); C546(0.98) | LDD1516 | [10] |
| LDCM0278 | AC19 | HEK-293T | C49(1.35); C653(1.04); C221(0.75); C546(0.99) | LDD1517 | [10] |
| LDCM0279 | AC2 | HEK-293T | C49(0.90); C653(1.04); C221(0.95); C546(1.27) | LDD1518 | [10] |
| LDCM0280 | AC20 | HEK-293T | C49(0.91); C653(0.86); C221(0.86); C546(0.79) | LDD1519 | [10] |
| LDCM0281 | AC21 | HEK-293T | C49(0.98); C653(0.95); C546(0.84) | LDD1520 | [10] |
| LDCM0282 | AC22 | HEK-293T | C49(1.06); C653(1.07); C546(1.08) | LDD1521 | [10] |
| LDCM0283 | AC23 | HEK-293T | C49(0.90); C653(1.00); C221(1.00); C546(1.10) | LDD1522 | [10] |
| LDCM0284 | AC24 | HEK-293T | C49(0.96); C653(0.98) | LDD1523 | [10] |
| LDCM0285 | AC25 | HEK-293T | C49(1.03); C653(1.04); C221(0.87); C546(0.75) | LDD1524 | [10] |
| LDCM0286 | AC26 | HEK-293T | C49(0.91); C653(1.01); C221(0.97); C546(0.96) | LDD1525 | [10] |
| LDCM0287 | AC27 | HEK-293T | C49(0.97); C653(0.95); C221(0.96); C546(1.15) | LDD1526 | [10] |
| LDCM0288 | AC28 | HEK-293T | C49(0.91); C653(1.00); C221(0.90); C546(0.91) | LDD1527 | [10] |
| LDCM0289 | AC29 | HEK-293T | C49(0.97); C653(0.99); C546(1.20) | LDD1528 | [10] |
| LDCM0290 | AC3 | HEK-293T | C49(1.03); C653(0.89); C221(0.93); C546(1.15) | LDD1529 | [10] |
| LDCM0291 | AC30 | HEK-293T | C49(1.06); C653(1.07); C546(0.92) | LDD1530 | [10] |
| LDCM0292 | AC31 | HEK-293T | C49(0.98); C653(1.00); C221(0.82); C546(0.94) | LDD1531 | [10] |
| LDCM0293 | AC32 | HEK-293T | C49(0.94); C653(1.06) | LDD1532 | [10] |
| LDCM0294 | AC33 | HEK-293T | C49(0.99); C653(1.02); C221(1.08); C546(0.81) | LDD1533 | [10] |
| LDCM0295 | AC34 | HEK-293T | C49(0.99); C653(0.99); C221(0.92); C546(1.11) | LDD1534 | [10] |
| LDCM0296 | AC35 | HEK-293T | C49(1.03); C653(0.93); C221(0.94); C546(1.21) | LDD1535 | [10] |
| LDCM0297 | AC36 | HEK-293T | C49(0.95); C653(0.89); C221(0.90); C546(0.91) | LDD1536 | [10] |
| LDCM0298 | AC37 | HEK-293T | C49(0.95); C653(0.91); C546(0.90) | LDD1537 | [10] |
| LDCM0299 | AC38 | HEK-293T | C49(1.00); C653(0.99); C546(0.95) | LDD1538 | [10] |
| LDCM0300 | AC39 | HEK-293T | C49(0.98); C653(1.02); C221(0.90); C546(0.93) | LDD1539 | [10] |
| LDCM0301 | AC4 | HEK-293T | C49(0.99); C653(0.80); C221(1.03); C546(1.18) | LDD1540 | [10] |
| LDCM0302 | AC40 | HEK-293T | C49(1.03); C653(1.01) | LDD1541 | [10] |
| LDCM0303 | AC41 | HEK-293T | C49(1.02); C653(1.05); C221(1.03); C546(0.80) | LDD1542 | [10] |
| LDCM0304 | AC42 | HEK-293T | C49(1.00); C653(0.95); C221(0.90); C546(1.22) | LDD1543 | [10] |
| LDCM0305 | AC43 | HEK-293T | C49(1.00); C653(0.86); C221(0.92); C546(1.28) | LDD1544 | [10] |
| LDCM0306 | AC44 | HEK-293T | C49(0.88); C653(0.77); C221(0.94); C546(0.96) | LDD1545 | [10] |
| LDCM0307 | AC45 | HEK-293T | C49(0.93); C653(0.99); C546(1.03) | LDD1546 | [10] |
| LDCM0308 | AC46 | HEK-293T | C49(1.03); C653(0.95); C546(1.06) | LDD1547 | [10] |
| LDCM0309 | AC47 | HEK-293T | C49(1.00); C653(0.90); C221(0.92); C546(0.90) | LDD1548 | [10] |
| LDCM0310 | AC48 | HEK-293T | C49(1.06); C653(0.89) | LDD1549 | [10] |
| LDCM0311 | AC49 | HEK-293T | C49(1.03); C653(0.93); C221(1.02); C546(0.80) | LDD1550 | [10] |
| LDCM0312 | AC5 | HEK-293T | C49(0.93); C653(0.93); C546(0.87) | LDD1551 | [10] |
| LDCM0313 | AC50 | HEK-293T | C49(0.95); C653(1.01); C221(1.02); C546(0.92) | LDD1552 | [10] |
| LDCM0314 | AC51 | HEK-293T | C49(1.00); C653(0.93); C221(1.00); C546(1.19) | LDD1553 | [10] |
| LDCM0315 | AC52 | HEK-293T | C49(0.91); C653(0.80); C221(0.99); C546(0.93) | LDD1554 | [10] |
| LDCM0316 | AC53 | HEK-293T | C49(0.97); C653(0.93); C546(0.95) | LDD1555 | [10] |
| LDCM0317 | AC54 | HEK-293T | C49(0.96); C653(0.91); C546(1.10) | LDD1556 | [10] |
| LDCM0318 | AC55 | HEK-293T | C49(1.01); C653(0.95); C221(0.93); C546(0.97) | LDD1557 | [10] |
| LDCM0319 | AC56 | HEK-293T | C49(0.99); C653(0.91) | LDD1558 | [10] |
| LDCM0320 | AC57 | HEK-293T | C49(1.01); C653(0.95); C221(1.02); C546(1.15) | LDD1559 | [10] |
| LDCM0321 | AC58 | HEK-293T | C49(1.01); C653(0.92); C221(0.96); C546(1.04) | LDD1560 | [10] |
| LDCM0322 | AC59 | HEK-293T | C49(1.00); C653(0.87); C221(0.97); C546(1.27) | LDD1561 | [10] |
| LDCM0323 | AC6 | HEK-293T | C49(0.99); C653(0.97); C546(1.12) | LDD1562 | [10] |
| LDCM0324 | AC60 | HEK-293T | C49(1.00); C653(0.88); C221(0.90); C546(0.78) | LDD1563 | [10] |
| LDCM0325 | AC61 | HEK-293T | C49(0.95); C653(0.88); C546(1.07) | LDD1564 | [10] |
| LDCM0326 | AC62 | HEK-293T | C49(1.02); C653(0.86); C546(1.03) | LDD1565 | [10] |
| LDCM0327 | AC63 | HEK-293T | C49(1.03); C653(0.95); C221(1.00); C546(0.99) | LDD1566 | [10] |
| LDCM0328 | AC64 | HEK-293T | C49(1.04); C653(0.92) | LDD1567 | [10] |
| LDCM0334 | AC7 | HEK-293T | C49(0.90); C653(0.98); C221(1.03); C546(0.96) | LDD1568 | [10] |
| LDCM0345 | AC8 | HEK-293T | C49(1.01); C653(1.01) | LDD1569 | [10] |
| LDCM0248 | AKOS034007472 | HEK-293T | C49(0.99); C653(0.90); C546(0.88) | LDD1511 | [10] |
| LDCM0356 | AKOS034007680 | HEK-293T | C49(1.01); C653(1.05); C221(1.02); C546(0.98) | LDD1570 | [10] |
| LDCM0275 | AKOS034007705 | HEK-293T | C49(1.01); C653(0.98) | LDD1514 | [10] |
| LDCM0108 | Chloroacetamide | HeLa | C364(0.00); C418(0.00); C346(0.00); C695(0.00) | LDD0222 | [6] |
| LDCM0367 | CL1 | HEK-293T | C49(0.86); C653(0.97); C221(1.16); C546(1.35) | LDD1571 | [10] |
| LDCM0368 | CL10 | HEK-293T | C49(1.32); C653(1.15); C546(0.83) | LDD1572 | [10] |
| LDCM0369 | CL100 | HEK-293T | C49(0.99); C653(0.93); C546(1.45) | LDD1573 | [10] |
| LDCM0370 | CL101 | HEK-293T | C49(0.94); C653(0.93); C221(0.87); C546(1.20) | LDD1574 | [10] |
| LDCM0371 | CL102 | HEK-293T | C49(1.04); C653(0.90); C221(1.08) | LDD1575 | [10] |
| LDCM0372 | CL103 | HEK-293T | C49(0.92); C653(1.09); C221(1.00); C546(0.86) | LDD1576 | [10] |
| LDCM0373 | CL104 | HEK-293T | C49(0.99); C653(0.92); C546(0.99) | LDD1577 | [10] |
| LDCM0374 | CL105 | HEK-293T | C49(0.99); C653(0.92); C221(1.06); C546(0.96) | LDD1578 | [10] |
| LDCM0375 | CL106 | HEK-293T | C49(0.83); C653(0.96); C221(1.02) | LDD1579 | [10] |
| LDCM0376 | CL107 | HEK-293T | C49(0.97); C653(1.02); C221(0.98); C546(0.93) | LDD1580 | [10] |
| LDCM0377 | CL108 | HEK-293T | C49(1.05); C653(0.92); C546(1.15) | LDD1581 | [10] |
| LDCM0378 | CL109 | HEK-293T | C49(0.95); C653(0.96); C221(1.00); C546(0.94) | LDD1582 | [10] |
| LDCM0379 | CL11 | HEK-293T | C49(0.96); C653(1.10); C221(1.15); C546(0.69) | LDD1583 | [10] |
| LDCM0380 | CL110 | HEK-293T | C49(0.87); C653(1.02); C221(0.94) | LDD1584 | [10] |
| LDCM0381 | CL111 | HEK-293T | C49(0.94); C653(1.06); C221(0.92); C546(1.04) | LDD1585 | [10] |
| LDCM0382 | CL112 | HEK-293T | C49(1.16); C653(0.98); C546(0.84) | LDD1586 | [10] |
| LDCM0383 | CL113 | HEK-293T | C49(0.91); C653(1.04); C221(0.94); C546(1.26) | LDD1587 | [10] |
| LDCM0384 | CL114 | HEK-293T | C49(0.95); C653(1.02); C221(0.97) | LDD1588 | [10] |
| LDCM0385 | CL115 | HEK-293T | C49(0.95); C653(1.05); C221(1.03); C546(1.26) | LDD1589 | [10] |
| LDCM0386 | CL116 | HEK-293T | C49(0.96); C653(0.97); C546(1.46) | LDD1590 | [10] |
| LDCM0387 | CL117 | HEK-293T | C49(0.95); C653(0.94); C221(1.07); C546(1.48) | LDD1591 | [10] |
| LDCM0388 | CL118 | HEK-293T | C49(0.97); C653(0.88); C221(1.04) | LDD1592 | [10] |
| LDCM0389 | CL119 | HEK-293T | C49(0.94); C653(1.01); C221(1.02); C546(1.03) | LDD1593 | [10] |
| LDCM0390 | CL12 | HEK-293T | C49(1.08); C653(1.16) | LDD1594 | [10] |
| LDCM0391 | CL120 | HEK-293T | C49(1.00); C653(0.94); C546(1.35) | LDD1595 | [10] |
| LDCM0392 | CL121 | HEK-293T | C49(0.92); C653(0.83); C221(1.04); C546(1.28) | LDD1596 | [10] |
| LDCM0393 | CL122 | HEK-293T | C49(0.99); C653(0.86); C221(0.99) | LDD1597 | [10] |
| LDCM0394 | CL123 | HEK-293T | C49(0.81); C653(0.99); C221(0.90); C546(0.76) | LDD1598 | [10] |
| LDCM0395 | CL124 | HEK-293T | C49(1.03); C653(0.83); C546(1.28) | LDD1599 | [10] |
| LDCM0396 | CL125 | HEK-293T | C49(0.88); C653(0.73); C221(1.09); C546(1.03) | LDD1600 | [10] |
| LDCM0397 | CL126 | HEK-293T | C49(1.09); C653(0.79); C221(0.98) | LDD1601 | [10] |
| LDCM0398 | CL127 | HEK-293T | C49(1.04); C653(0.87); C221(0.83); C546(1.28) | LDD1602 | [10] |
| LDCM0399 | CL128 | HEK-293T | C49(1.08); C653(0.88); C546(1.13) | LDD1603 | [10] |
| LDCM0400 | CL13 | HEK-293T | C49(0.95); C653(0.85); C221(1.04); C546(1.12) | LDD1604 | [10] |
| LDCM0401 | CL14 | HEK-293T | C49(0.95); C653(0.91); C221(1.04) | LDD1605 | [10] |
| LDCM0402 | CL15 | HEK-293T | C49(0.78); C653(1.06); C221(1.01); C546(0.84) | LDD1606 | [10] |
| LDCM0403 | CL16 | HEK-293T | C49(0.97); C653(0.99); C546(1.08) | LDD1607 | [10] |
| LDCM0404 | CL17 | HEK-293T | C49(0.88); C653(0.98); C221(1.06); C546(0.66) | LDD1608 | [10] |
| LDCM0405 | CL18 | HEK-293T | C49(0.96); C653(0.97); C221(0.99); C546(1.24) | LDD1609 | [10] |
| LDCM0406 | CL19 | HEK-293T | C49(1.07); C653(0.82); C221(1.05); C546(0.75) | LDD1610 | [10] |
| LDCM0407 | CL2 | HEK-293T | C49(1.02); C653(0.92); C221(1.14) | LDD1611 | [10] |
| LDCM0408 | CL20 | HEK-293T | C49(0.99); C653(0.82); C221(1.14); C546(0.97) | LDD1612 | [10] |
| LDCM0409 | CL21 | HEK-293T | C49(0.96); C653(0.90); C546(0.75) | LDD1613 | [10] |
| LDCM0410 | CL22 | HEK-293T | C49(1.11); C653(1.11); C546(0.87) | LDD1614 | [10] |
| LDCM0411 | CL23 | HEK-293T | C49(1.10); C653(1.06); C221(1.23); C546(0.85) | LDD1615 | [10] |
| LDCM0412 | CL24 | HEK-293T | C49(1.08); C653(1.07) | LDD1616 | [10] |
| LDCM0413 | CL25 | HEK-293T | C49(0.89); C653(0.84); C221(1.04); C546(1.18) | LDD1617 | [10] |
| LDCM0414 | CL26 | HEK-293T | C49(0.96); C653(0.95); C221(1.10) | LDD1618 | [10] |
| LDCM0415 | CL27 | HEK-293T | C49(0.97); C653(1.02); C221(0.95); C546(1.10) | LDD1619 | [10] |
| LDCM0416 | CL28 | HEK-293T | C49(0.99); C653(0.97); C546(1.13) | LDD1620 | [10] |
| LDCM0417 | CL29 | HEK-293T | C49(0.95); C653(0.99); C221(1.04); C546(0.89) | LDD1621 | [10] |
| LDCM0418 | CL3 | HEK-293T | C49(0.92); C653(1.02); C221(1.10); C546(0.92) | LDD1622 | [10] |
| LDCM0419 | CL30 | HEK-293T | C49(1.04); C653(0.99); C221(0.99); C546(1.05) | LDD1623 | [10] |
| LDCM0420 | CL31 | HEK-293T | C49(0.99); C653(0.84); C221(1.04); C546(0.87) | LDD1624 | [10] |
| LDCM0421 | CL32 | HEK-293T | C49(0.90); C653(0.81); C221(1.02); C546(0.96) | LDD1625 | [10] |
| LDCM0422 | CL33 | HEK-293T | C49(0.85); C653(0.98); C546(0.86) | LDD1626 | [10] |
| LDCM0423 | CL34 | HEK-293T | C49(1.16); C653(1.02); C546(0.84) | LDD1627 | [10] |
| LDCM0424 | CL35 | HEK-293T | C49(1.10); C653(1.02); C221(1.18); C546(0.68) | LDD1628 | [10] |
| LDCM0425 | CL36 | HEK-293T | C49(1.10); C653(1.03) | LDD1629 | [10] |
| LDCM0426 | CL37 | HEK-293T | C49(0.79); C653(0.89); C221(1.00); C546(1.31) | LDD1630 | [10] |
| LDCM0428 | CL39 | HEK-293T | C49(0.92); C653(0.91); C221(0.96); C546(1.10) | LDD1632 | [10] |
| LDCM0429 | CL4 | HEK-293T | C49(0.91); C653(0.90); C546(0.85) | LDD1633 | [10] |
| LDCM0430 | CL40 | HEK-293T | C49(0.98); C653(1.01); C546(1.20) | LDD1634 | [10] |
| LDCM0431 | CL41 | HEK-293T | C49(1.29); C653(1.04); C221(0.95); C546(0.81) | LDD1635 | [10] |
| LDCM0432 | CL42 | HEK-293T | C49(0.99); C653(0.92); C221(0.88); C546(1.33) | LDD1636 | [10] |
| LDCM0433 | CL43 | HEK-293T | C49(0.95); C653(0.88); C221(0.91); C546(1.01) | LDD1637 | [10] |
| LDCM0434 | CL44 | HEK-293T | C49(0.95); C653(0.79); C221(1.12); C546(0.71) | LDD1638 | [10] |
| LDCM0435 | CL45 | HEK-293T | C49(0.97); C653(0.94); C546(0.86) | LDD1639 | [10] |
| LDCM0436 | CL46 | HEK-293T | C49(1.14); C653(0.89); C546(0.93) | LDD1640 | [10] |
| LDCM0437 | CL47 | HEK-293T | C49(1.07); C653(1.00); C221(1.08); C546(0.67) | LDD1641 | [10] |
| LDCM0438 | CL48 | HEK-293T | C49(1.16); C653(1.03) | LDD1642 | [10] |
| LDCM0439 | CL49 | HEK-293T | C49(0.91); C653(0.81); C221(1.13); C546(1.91) | LDD1643 | [10] |
| LDCM0440 | CL5 | HEK-293T | C49(0.99); C653(0.89); C221(1.01); C546(0.90) | LDD1644 | [10] |
| LDCM0441 | CL50 | HEK-293T | C49(1.08); C653(0.92); C221(1.05) | LDD1645 | [10] |
| LDCM0443 | CL52 | HEK-293T | C49(1.01); C653(1.01); C546(1.01) | LDD1646 | [10] |
| LDCM0444 | CL53 | HEK-293T | C49(1.09); C653(1.00); C221(1.07); C546(0.62) | LDD1647 | [10] |
| LDCM0445 | CL54 | HEK-293T | C49(0.86); C653(0.98); C221(0.90); C546(0.76) | LDD1648 | [10] |
| LDCM0446 | CL55 | HEK-293T | C49(1.06); C653(0.93); C221(0.94); C546(1.06) | LDD1649 | [10] |
| LDCM0447 | CL56 | HEK-293T | C49(0.94); C653(0.76); C221(1.08); C546(0.78) | LDD1650 | [10] |
| LDCM0448 | CL57 | HEK-293T | C49(0.94); C653(0.95); C546(0.92) | LDD1651 | [10] |
| LDCM0449 | CL58 | HEK-293T | C49(1.02); C653(1.01); C546(0.75) | LDD1652 | [10] |
| LDCM0450 | CL59 | HEK-293T | C49(0.99); C653(0.94); C221(1.14); C546(0.82) | LDD1653 | [10] |
| LDCM0451 | CL6 | HEK-293T | C49(0.86); C653(0.95); C221(0.93); C546(0.89) | LDD1654 | [10] |
| LDCM0452 | CL60 | HEK-293T | C49(1.12); C653(1.02) | LDD1655 | [10] |
| LDCM0453 | CL61 | HEK-293T | C49(0.89); C653(0.91); C221(0.98); C546(1.15) | LDD1656 | [10] |
| LDCM0454 | CL62 | HEK-293T | C49(0.99); C653(0.96); C221(1.19) | LDD1657 | [10] |
| LDCM0455 | CL63 | HEK-293T | C49(1.01); C653(0.99); C221(1.02); C546(1.20) | LDD1658 | [10] |
| LDCM0456 | CL64 | HEK-293T | C49(0.90); C653(0.97); C546(1.24) | LDD1659 | [10] |
| LDCM0457 | CL65 | HEK-293T | C49(1.02); C653(1.01); C221(1.14); C546(0.99) | LDD1660 | [10] |
| LDCM0458 | CL66 | HEK-293T | C49(0.95); C653(0.88); C221(1.00); C546(1.04) | LDD1661 | [10] |
| LDCM0459 | CL67 | HEK-293T | C49(0.96); C653(0.89); C221(1.05); C546(0.89) | LDD1662 | [10] |
| LDCM0460 | CL68 | HEK-293T | C49(0.95); C653(0.76); C221(1.25); C546(0.70) | LDD1663 | [10] |
| LDCM0461 | CL69 | HEK-293T | C49(1.02); C653(0.86); C546(0.79) | LDD1664 | [10] |
| LDCM0462 | CL7 | HEK-293T | C49(1.15); C653(0.87); C221(1.05); C546(0.94) | LDD1665 | [10] |
| LDCM0463 | CL70 | HEK-293T | C49(1.20); C653(0.95); C546(0.90) | LDD1666 | [10] |
| LDCM0464 | CL71 | HEK-293T | C49(1.10); C653(0.93); C221(1.18); C546(0.89) | LDD1667 | [10] |
| LDCM0465 | CL72 | HEK-293T | C49(1.04); C653(1.02) | LDD1668 | [10] |
| LDCM0466 | CL73 | HEK-293T | C49(0.86); C653(0.85); C221(1.04); C546(1.30) | LDD1669 | [10] |
| LDCM0467 | CL74 | HEK-293T | C49(1.11); C653(0.86); C221(1.08) | LDD1670 | [10] |
| LDCM0469 | CL76 | HEK-293T | C49(1.03); C653(0.93); C546(1.08) | LDD1672 | [10] |
| LDCM0470 | CL77 | HEK-293T | C49(0.97); C653(0.91); C221(1.00); C546(0.64) | LDD1673 | [10] |
| LDCM0471 | CL78 | HEK-293T | C49(0.96); C653(0.97); C221(1.05); C546(1.23) | LDD1674 | [10] |
| LDCM0472 | CL79 | HEK-293T | C49(0.95); C653(0.88); C221(0.95); C546(1.11) | LDD1675 | [10] |
| LDCM0473 | CL8 | HEK-293T | C49(1.06); C653(0.82); C221(0.83); C546(0.83) | LDD1676 | [10] |
| LDCM0474 | CL80 | HEK-293T | C49(0.91); C653(0.76); C221(1.00); C546(0.81) | LDD1677 | [10] |
| LDCM0475 | CL81 | HEK-293T | C49(0.96); C653(0.90); C546(0.94) | LDD1678 | [10] |
| LDCM0476 | CL82 | HEK-293T | C49(1.07); C653(0.97); C546(0.90) | LDD1679 | [10] |
| LDCM0477 | CL83 | HEK-293T | C49(1.10); C653(0.92); C221(0.98); C546(0.89) | LDD1680 | [10] |
| LDCM0478 | CL84 | HEK-293T | C49(0.99); C653(0.98) | LDD1681 | [10] |
| LDCM0479 | CL85 | HEK-293T | C49(0.93); C653(0.83); C221(1.01); C546(1.78) | LDD1682 | [10] |
| LDCM0480 | CL86 | HEK-293T | C49(1.18); C653(0.80); C221(0.99) | LDD1683 | [10] |
| LDCM0481 | CL87 | HEK-293T | C49(0.89); C653(0.82); C221(0.94); C546(1.08) | LDD1684 | [10] |
| LDCM0482 | CL88 | HEK-293T | C49(1.05); C653(0.79); C546(1.13) | LDD1685 | [10] |
| LDCM0483 | CL89 | HEK-293T | C49(1.02); C653(0.84); C221(1.08); C546(0.99) | LDD1686 | [10] |
| LDCM0484 | CL9 | HEK-293T | C49(0.90); C653(0.90); C546(0.90) | LDD1687 | [10] |
| LDCM0485 | CL90 | HEK-293T | C49(0.85); C653(0.95); C221(0.71); C546(0.97) | LDD1688 | [10] |
| LDCM0486 | CL91 | HEK-293T | C49(0.94); C653(0.68); C221(0.93); C546(1.19) | LDD1689 | [10] |
| LDCM0487 | CL92 | HEK-293T | C49(0.95); C653(0.73); C221(0.97); C546(0.89) | LDD1690 | [10] |
| LDCM0488 | CL93 | HEK-293T | C49(0.91); C653(0.87); C546(0.96) | LDD1691 | [10] |
| LDCM0489 | CL94 | HEK-293T | C49(1.02); C653(0.90); C546(1.09) | LDD1692 | [10] |
| LDCM0490 | CL95 | HEK-293T | C49(0.90); C653(0.93); C221(0.97); C546(0.67) | LDD1693 | [10] |
| LDCM0491 | CL96 | HEK-293T | C49(1.08); C653(0.90) | LDD1694 | [10] |
| LDCM0492 | CL97 | HEK-293T | C49(0.98); C653(0.88); C221(1.00); C546(1.60) | LDD1695 | [10] |
| LDCM0493 | CL98 | HEK-293T | C49(1.07); C653(0.96); C221(1.12) | LDD1696 | [10] |
| LDCM0494 | CL99 | HEK-293T | C49(0.97); C653(0.95); C221(0.91); C546(0.91) | LDD1697 | [10] |
| LDCM0495 | E2913 | HEK-293T | C49(0.84); C653(1.01); C221(1.15); C546(0.89) | LDD1698 | [10] |
| LDCM0625 | F8 | Ramos | C201(1.87); C695(1.43); C49(2.43); C546(0.73) | LDD2187 | [11] |
| LDCM0572 | Fragment10 | Ramos | C201(0.66); C695(2.89); C49(1.86); C546(0.53) | LDD2189 | [11] |
| LDCM0573 | Fragment11 | Ramos | C201(0.80); C695(0.18); C49(1.18); C546(0.15) | LDD2190 | [11] |
| LDCM0574 | Fragment12 | Ramos | C201(1.09); C695(5.62); C49(0.78); C546(0.73) | LDD2191 | [11] |
| LDCM0575 | Fragment13 | Ramos | C201(1.59); C695(1.17); C49(0.70); C546(0.92) | LDD2192 | [11] |
| LDCM0576 | Fragment14 | Ramos | C201(1.17); C695(6.15); C49(1.35); C546(1.78) | LDD2193 | [11] |
| LDCM0579 | Fragment20 | Ramos | C201(1.14); C695(8.24); C49(0.72); C546(0.90) | LDD2194 | [11] |
| LDCM0580 | Fragment21 | Ramos | C201(1.37); C695(1.40); C49(0.95); C546(1.22) | LDD2195 | [11] |
| LDCM0582 | Fragment23 | Ramos | C201(0.51); C695(1.07); C49(1.14); C546(0.69) | LDD2196 | [11] |
| LDCM0578 | Fragment27 | Ramos | C201(1.47); C695(0.98); C49(0.98); C546(1.06) | LDD2197 | [11] |
| LDCM0586 | Fragment28 | Ramos | C201(0.54); C695(0.92); C49(0.79); C546(1.14) | LDD2198 | [11] |
| LDCM0588 | Fragment30 | Ramos | C201(1.98); C695(1.19); C49(0.77); C546(0.51) | LDD2199 | [11] |
| LDCM0589 | Fragment31 | Ramos | C201(1.81); C695(1.77); C49(1.00); C546(0.91) | LDD2200 | [11] |
| LDCM0590 | Fragment32 | Ramos | C201(1.09); C695(5.77); C49(1.11); C546(0.43) | LDD2201 | [11] |
| LDCM0468 | Fragment33 | HEK-293T | C49(0.94); C653(0.92); C221(1.09); C546(0.99) | LDD1671 | [10] |
| LDCM0596 | Fragment38 | Ramos | C201(1.84); C695(1.65); C49(1.28); C546(3.37) | LDD2203 | [11] |
| LDCM0566 | Fragment4 | Ramos | C201(0.82); C695(5.20); C49(1.03); C546(0.84) | LDD2184 | [11] |
| LDCM0427 | Fragment51 | HEK-293T | C49(1.00); C653(0.99); C221(1.11) | LDD1631 | [10] |
| LDCM0610 | Fragment52 | Ramos | C201(2.47); C695(1.10); C49(0.45); C546(0.52) | LDD2204 | [11] |
| LDCM0614 | Fragment56 | Ramos | C201(3.91); C695(1.07); C49(1.06); C546(1.24) | LDD2205 | [11] |
| LDCM0569 | Fragment7 | Ramos | C201(0.59); C695(4.16); C49(2.57); C546(0.83) | LDD2186 | [11] |
| LDCM0571 | Fragment9 | Ramos | C201(0.77); C695(10.79); C49(0.80); C546(0.79) | LDD2188 | [11] |
| LDCM0107 | IAA | HeLa | C364(0.00); H610(0.00); C695(0.00); C418(0.00) | LDD0221 | [6] |
| LDCM0022 | KB02 | HEK-293T | C221(1.09); C49(0.93); C653(1.03) | LDD1492 | [10] |
| LDCM0023 | KB03 | HEK-293T | C221(0.96); C49(1.00); C653(0.85) | LDD1497 | [10] |
| LDCM0024 | KB05 | G361 | C521(3.60); C62(1.05) | LDD3311 | [1] |
| LDCM0109 | NEM | HeLa | C695(0.00); H610(0.00) | LDD0223 | [6] |
| LDCM0627 | NUDT7-COV-1 | HEK-293T | C346(1.05) | LDD2206 | [12] |
| LDCM0628 | OTUB2-COV-1 | HEK-293T | C364(0.85); C695(0.78); C346(0.61) | LDD2207 | [12] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Eyes absent homolog 3 (EYA3) | EYA family | Q99504 | |||
| Calcium/calmodulin-dependent protein kinase type II subunit alpha (CAMK2A) | CAMK Ser/Thr protein kinase family | Q9UQM7 | |||
Transporter and channel
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Bcl-2-related ovarian killer protein (BOK) | Bcl-2 family | Q9UMX3 | |||
Transcription factor
GPCR
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Neuropeptide FF receptor 1 (NPFFR1) | G-protein coupled receptor 1 family | Q9GZQ6 | |||
Other
References
