Details of the Target
General Information of Target
| Target ID | LDTP14045 | |||||
|---|---|---|---|---|---|---|
| Target Name | Disheveled-associated activator of morphogenesis 1 (DAAM1) | |||||
| Gene Name | DAAM1 | |||||
| Gene ID | 23002 | |||||
| Synonyms |
KIAA0666; Disheveled-associated activator of morphogenesis 1 |
|||||
| 3D Structure | ||||||
| Sequence |
MSRLIVKNLPNGMKEERFRQLFAAFGTLTDCSLKFTKDGKFRKFGFIGFKSEEEAQKAQK
HFNKSFIDTSRITVEFCKSFGDPAKPRAWSKHAQKPSQPKQPPKDSTTPEIKKDEKKKKV AGQLEKLKEDTEFQEFLSVHQRRAQAATWANDGLDAEPSKGKSKPASDYLNFDSDSGQES EEEGAGEDLEEEASLEPKAAVQKELSDMDYLKSKMVKAGSSSSSEEEESEDEAVHCDEGS EAEEEDSSATPVLQERDSKGAGQEQGMPAGKKRPPEARAETEKPANQKEPTTCHTVKLRG APFNVTEKNVMEFLAPLKPVAIRIVRNAHGNKTGYIFVDFSNEEEVKQALKCNREYMGGR YIEVFREKNVPTTKGAPKNTTKSWQGRILGENEEEEDLAESGRLFVRNLPYTSTEEDLEK LFSKYGPLSELHYPIDSLTKKPKGFAFITFMFPEHAVKAYSEVDGQVFQGRMLHVLPSTI KKEASEDASALGSSSYKKKKEAQDKANSASSHNWNTLFMGPNAVADAIAQKYNATKSQVF DHETKGSVAVRVALGETQLVQEVRRFLIDNGVSLDSFSQAAAERSKTVILVKNLPAGTLA AQLQETFGHFGSLGRVLLPEGGITAIVEFLEPLEARKAFRHLAYSKFHHVPLYLEWAPVG VFSSTAPQKKKLQDTPSEPMEKDPAEPETVPDGETPEDENPTEEGADNSSAKMEEEEEEE EEEEESLPGCTLFIKNLNFDTTEEKLKEVFSKVGTVKSCSISKKKNKAGVLLSMGFGFVE YRKPEQAQKALKQLQGHVVDGHKLEVRISERATKPAVTLARKKQVPRKQTTSKILVRNIP FQAHSREIRELFSTFGELKTVRLPKKMTGTGTHRGFGFVDFLTKQDAKRAFNALCHSTHL YGRRLVLEWADSEVTLQALRRKTAAHFHEPPKKKRSVVLDEILEQLEGSDSDSEEQTLQL |
|||||
| Target Bioclass |
Other
|
|||||
| Family |
Formin homology family
|
|||||
| Subcellular location |
Cytoplasm
|
|||||
| Function |
Binds to disheveled (Dvl) and Rho, and mediates Wnt-induced Dvl-Rho complex formation. May play a role as a scaffolding protein to recruit Rho-GDP and Rho-GEF, thereby enhancing Rho-GTP formation. Can direct nucleation and elongation of new actin filaments. Involved in building functional cilia. Involved in the organization of the subapical actin network in multiciliated epithelial cells. Together with DAAM2, required for myocardial maturation and sarcomere assembly.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
P1 Probe Info |
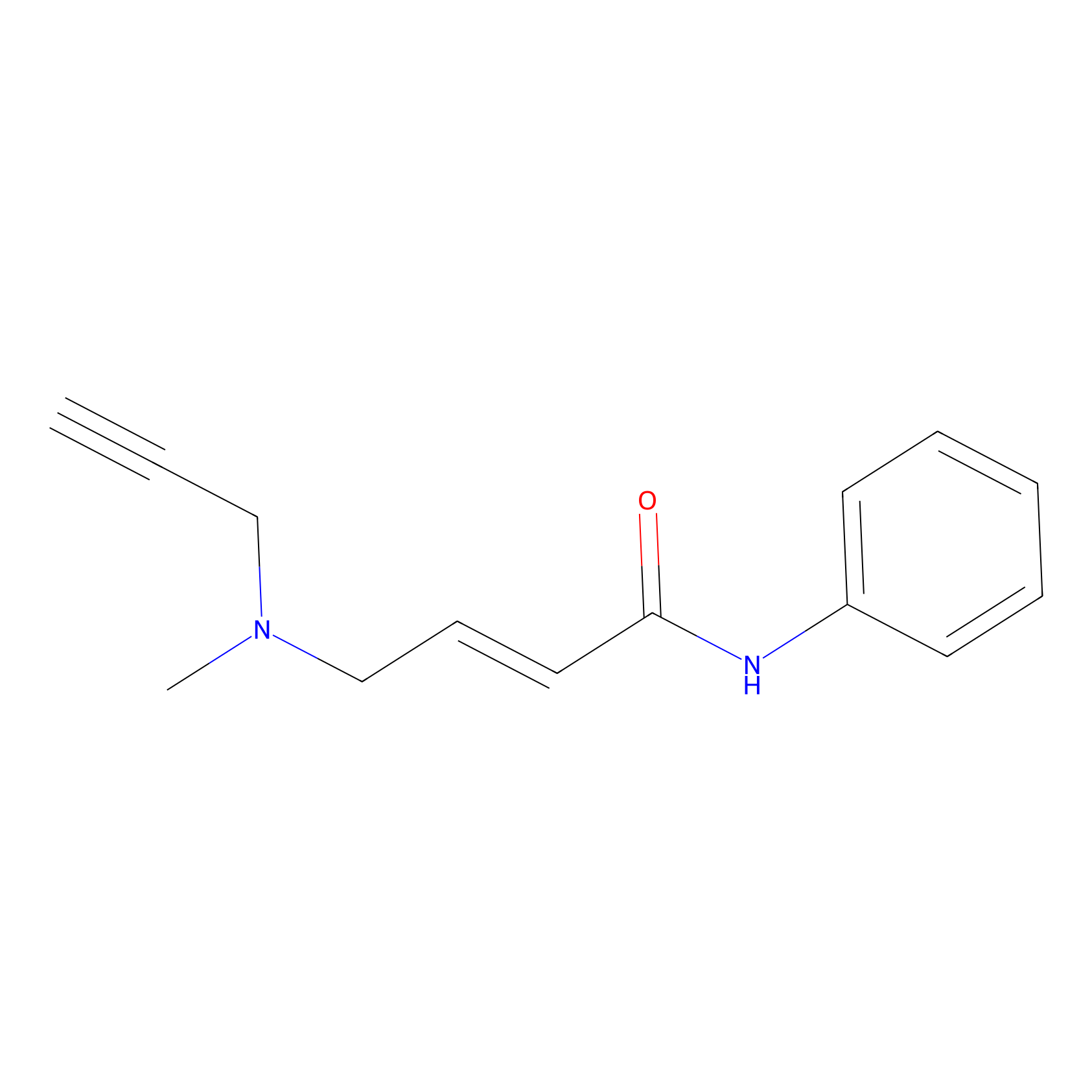 |
3.89 | LDD0452 | [1] | |
|
P2 Probe Info |
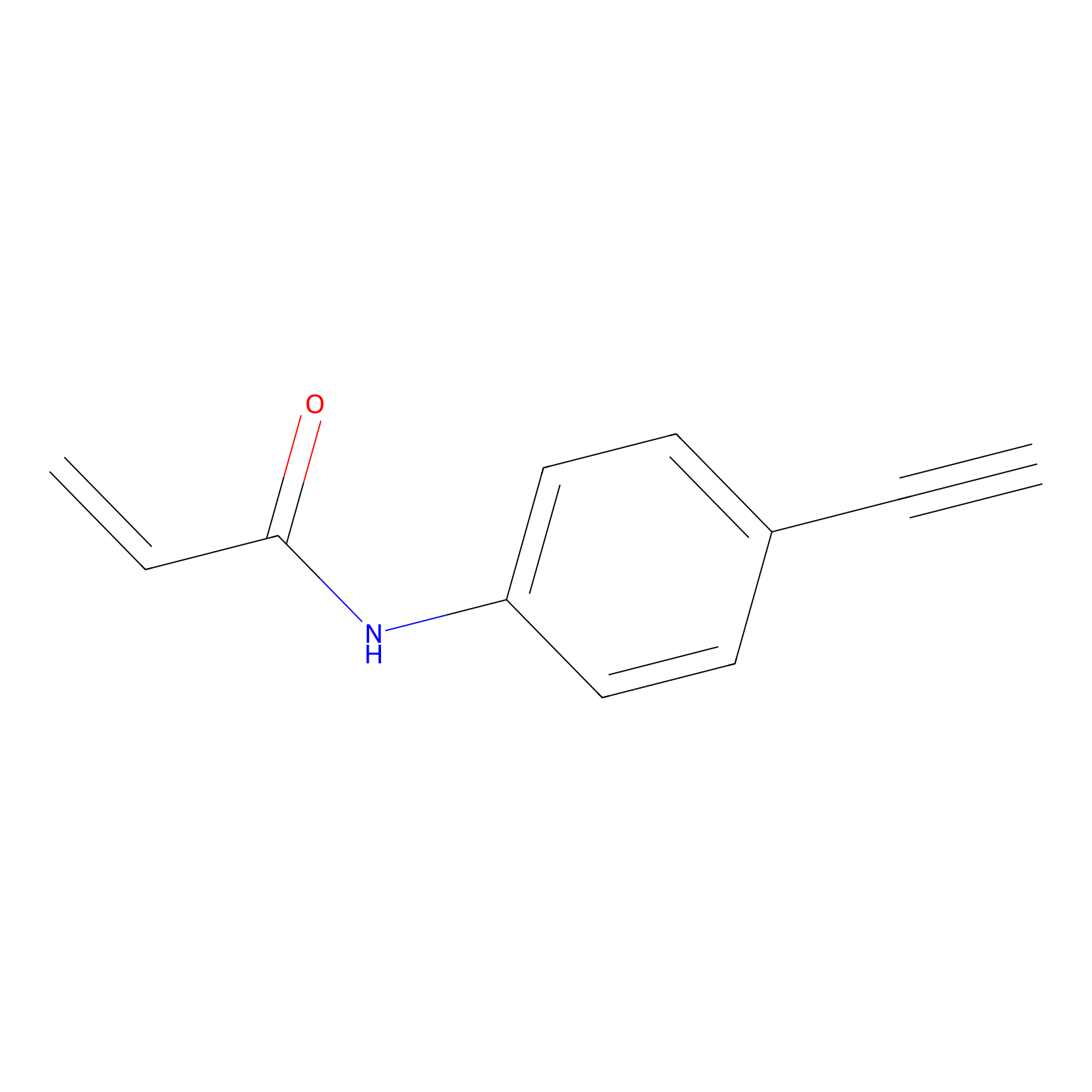 |
2.29 | LDD0453 | [1] | |
|
P3 Probe Info |
 |
1.68 | LDD0454 | [1] | |
|
P8 Probe Info |
 |
9.46 | LDD0455 | [1] | |
|
STPyne Probe Info |
 |
K138(10.00); K780(1.14) | LDD0277 | [2] | |
|
BTD Probe Info |
 |
C696(0.66) | LDD2122 | [3] | |
|
DBIA Probe Info |
 |
C87(0.87) | LDD1514 | [4] | |
|
N1 Probe Info |
 |
E365(0.00); K369(0.00); M363(0.00); T367(0.00) | LDD0245 | [5] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
N.A. | LDD0038 | [6] | |
|
Lodoacetamide azide Probe Info |
 |
N.A. | LDD0037 | [6] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0284 | AC24 | HEK-293T | C87(0.97) | LDD1523 | [4] |
| LDCM0293 | AC32 | HEK-293T | C87(0.95) | LDD1532 | [4] |
| LDCM0302 | AC40 | HEK-293T | C87(1.11) | LDD1541 | [4] |
| LDCM0310 | AC48 | HEK-293T | C87(0.95) | LDD1549 | [4] |
| LDCM0319 | AC56 | HEK-293T | C87(0.98) | LDD1558 | [4] |
| LDCM0328 | AC64 | HEK-293T | C87(0.89) | LDD1567 | [4] |
| LDCM0345 | AC8 | HEK-293T | C87(0.93) | LDD1569 | [4] |
| LDCM0275 | AKOS034007705 | HEK-293T | C87(0.87) | LDD1514 | [4] |
| LDCM0369 | CL100 | HEK-293T | C87(0.99) | LDD1573 | [4] |
| LDCM0373 | CL104 | HEK-293T | C87(1.17) | LDD1577 | [4] |
| LDCM0377 | CL108 | HEK-293T | C87(0.98) | LDD1581 | [4] |
| LDCM0382 | CL112 | HEK-293T | C87(0.87) | LDD1586 | [4] |
| LDCM0386 | CL116 | HEK-293T | C87(1.11) | LDD1590 | [4] |
| LDCM0390 | CL12 | HEK-293T | C87(1.03) | LDD1594 | [4] |
| LDCM0391 | CL120 | HEK-293T | C87(1.12) | LDD1595 | [4] |
| LDCM0395 | CL124 | HEK-293T | C87(1.49) | LDD1599 | [4] |
| LDCM0399 | CL128 | HEK-293T | C87(0.98) | LDD1603 | [4] |
| LDCM0403 | CL16 | HEK-293T | C87(1.24) | LDD1607 | [4] |
| LDCM0412 | CL24 | HEK-293T | C87(1.18) | LDD1616 | [4] |
| LDCM0416 | CL28 | HEK-293T | C87(1.12) | LDD1620 | [4] |
| LDCM0425 | CL36 | HEK-293T | C87(1.32) | LDD1629 | [4] |
| LDCM0429 | CL4 | HEK-293T | C87(1.27) | LDD1633 | [4] |
| LDCM0430 | CL40 | HEK-293T | C87(1.04) | LDD1634 | [4] |
| LDCM0438 | CL48 | HEK-293T | C87(1.04) | LDD1642 | [4] |
| LDCM0443 | CL52 | HEK-293T | C87(1.06) | LDD1646 | [4] |
| LDCM0452 | CL60 | HEK-293T | C87(1.16) | LDD1655 | [4] |
| LDCM0456 | CL64 | HEK-293T | C87(1.10) | LDD1659 | [4] |
| LDCM0465 | CL72 | HEK-293T | C87(1.05) | LDD1668 | [4] |
| LDCM0469 | CL76 | HEK-293T | C87(1.24) | LDD1672 | [4] |
| LDCM0478 | CL84 | HEK-293T | C87(1.01) | LDD1681 | [4] |
| LDCM0482 | CL88 | HEK-293T | C87(0.96) | LDD1685 | [4] |
| LDCM0491 | CL96 | HEK-293T | C87(0.98) | LDD1694 | [4] |
| LDCM0529 | Nucleophilic fragment 27b | MDA-MB-231 | C696(0.66) | LDD2122 | [3] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Transforming protein RhoA (RHOA) | Rho family | P61586 | |||
Other
References
