Details of the Target
General Information of Target
| Target ID | LDTP09976 | |||||
|---|---|---|---|---|---|---|
| Target Name | Sarcoplasmic/endoplasmic reticulum calcium ATPase 3 (ATP2A3) | |||||
| Gene Name | ATP2A3 | |||||
| Gene ID | 489 | |||||
| Synonyms |
Sarcoplasmic/endoplasmic reticulum calcium ATPase 3; SERCA3; SR Ca(2+)-ATPase 3; EC 7.2.2.10; Calcium pump 3 |
|||||
| 3D Structure | ||||||
| Sequence |
MPDPAKSAPAPKKGSKKAVTKAQKKDGKKRKRSRKESYSVYVYKVLKQVHPDTGISSKAM
GIMNSFVNDIFERIAGEASRLAHYNKRSTITSREIQTAVRLLLPGELAKHAVSEGTKAVT KYTSSK |
|||||
| Target Bioclass |
Enzyme
|
|||||
| Family |
Cation transport ATPase (P-type) (TC 3.A.3) family, Type IIA subfamily
|
|||||
| Subcellular location |
Nucleus membrane
|
|||||
| Function |
This magnesium-dependent enzyme catalyzes the hydrolysis of ATP coupled with the transport of calcium. Transports calcium ions from the cytosol into the sarcoplasmic/endoplasmic reticulum lumen. Contributes to calcium sequestration involved in muscular excitation/contraction.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
DBIA Probe Info |
 |
C636(4.54); C364(1.70) | LDD3310 | [1] | |
|
SAHA-CA-4PAP Probe Info |
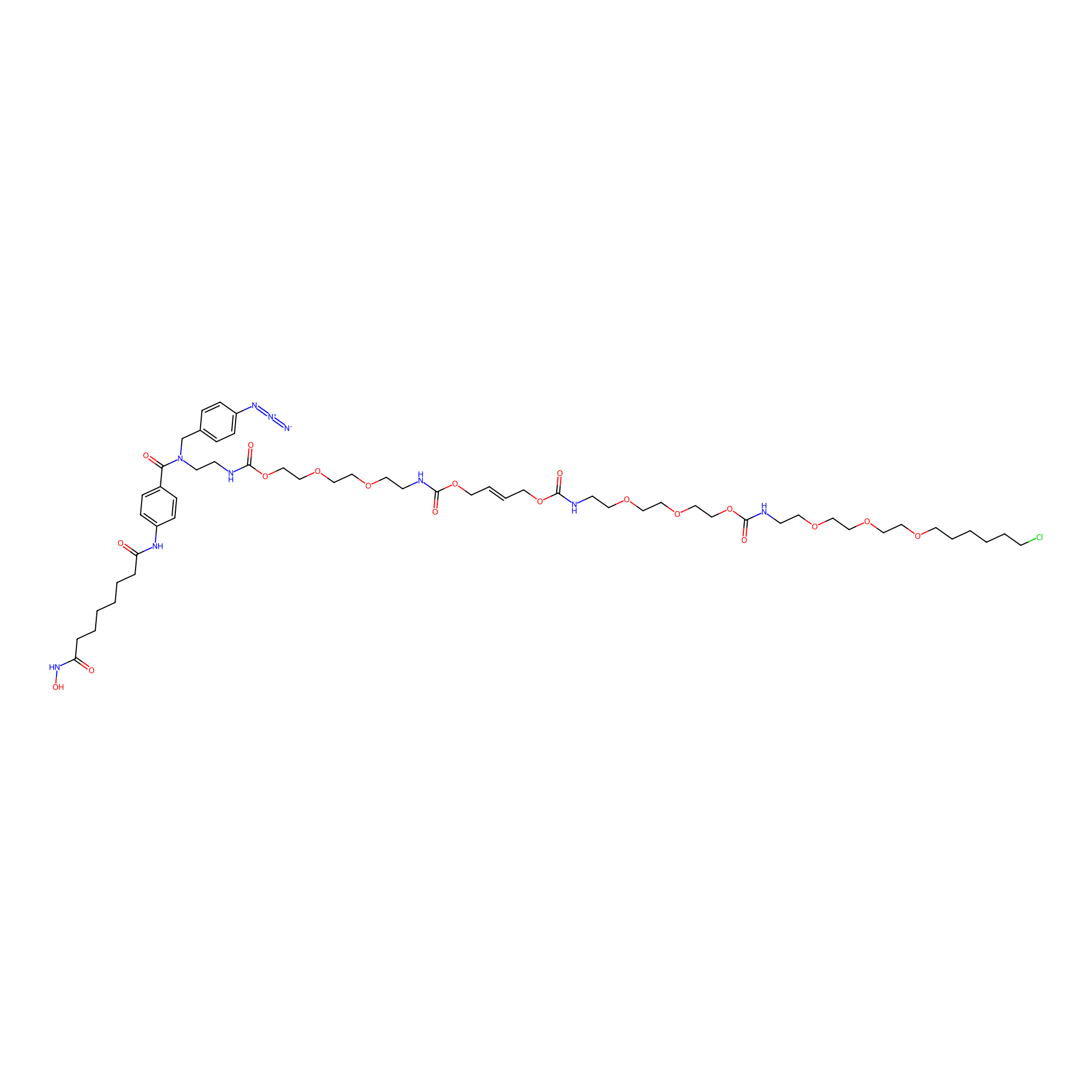 |
4.20 | LDD0269 | [2] | |
|
Johansson_61 Probe Info |
 |
_(5.64) | LDD1485 | [3] | |
|
AHL-Pu-1 Probe Info |
 |
C447(2.03) | LDD0168 | [4] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
C561(0.00); C581(0.00); C614(0.00); C498(0.00) | LDD0038 | [5] | |
|
IA-alkyne Probe Info |
 |
C471(0.00); C614(0.00); C561(0.00); C581(0.00) | LDD0036 | [5] | |
|
IPIAA_H Probe Info |
 |
N.A. | LDD0030 | [6] | |
|
IPIAA_L Probe Info |
 |
N.A. | LDD0031 | [6] | |
|
Lodoacetamide azide Probe Info |
 |
C561(0.00); C614(0.00); C498(0.00); C581(0.00) | LDD0037 | [5] | |
|
Compound 10 Probe Info |
 |
N.A. | LDD2216 | [7] | |
|
Acrolein Probe Info |
 |
C447(0.00); C364(0.00); C636(0.00) | LDD0217 | [8] | |
|
Cinnamaldehyde Probe Info |
 |
C364(0.00); C447(0.00) | LDD0220 | [8] | |
|
Crotonaldehyde Probe Info |
 |
C364(0.00); C447(0.00) | LDD0219 | [8] | |
|
Methacrolein Probe Info |
 |
C447(0.00); C364(0.00) | LDD0218 | [8] | |
|
NAIA_5 Probe Info |
 |
C614(0.00); C498(0.00); C636(0.00) | LDD2223 | [9] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
FFF probe11 Probe Info |
 |
20.00 | LDD0472 | [10] | |
|
FFF probe13 Probe Info |
 |
10.34 | LDD0476 | [10] | |
|
FFF probe3 Probe Info |
 |
20.00 | LDD0465 | [10] | |
|
STS-1 Probe Info |
 |
N.A. | LDD0136 | [11] | |
|
STS-2 Probe Info |
 |
N.A. | LDD0138 | [11] | |
|
VE-P Probe Info |
 |
N.A. | LDD0396 | [12] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0025 | 4SU-RNA | HEK-293T | C447(2.03) | LDD0168 | [4] |
| LDCM0108 | Chloroacetamide | HeLa | C447(0.00); C364(0.00); C636(0.00) | LDD0222 | [8] |
| LDCM0632 | CL-Sc | Hep-G2 | C636(2.11); C447(0.97) | LDD2227 | [9] |
| LDCM0634 | CY-0357 | Hep-G2 | C581(0.98) | LDD2228 | [9] |
| LDCM0625 | F8 | Ramos | C636(0.53); C364(1.65); C447(0.77); C471(0.98) | LDD2187 | [13] |
| LDCM0572 | Fragment10 | Ramos | C636(3.22); C364(0.39); C447(0.97); C561(0.31) | LDD2189 | [13] |
| LDCM0573 | Fragment11 | Ramos | C636(0.36); C364(0.48); C447(0.15); C561(0.26) | LDD2190 | [13] |
| LDCM0574 | Fragment12 | Ramos | C636(7.00); C364(0.66); C447(1.48); C561(0.63) | LDD2191 | [13] |
| LDCM0575 | Fragment13 | Ramos | C636(1.46); C364(0.76); C447(0.96); C561(1.16) | LDD2192 | [13] |
| LDCM0576 | Fragment14 | Ramos | C636(2.29); C364(1.89); C447(2.52); C561(0.70) | LDD2193 | [13] |
| LDCM0579 | Fragment20 | Ramos | C636(7.69); C364(0.75); C447(3.12); C561(0.80) | LDD2194 | [13] |
| LDCM0580 | Fragment21 | Ramos | C636(1.79); C364(0.67); C447(1.37); C471(0.97) | LDD2195 | [13] |
| LDCM0582 | Fragment23 | Ramos | C636(2.20); C364(0.54); C447(1.43); C561(0.29) | LDD2196 | [13] |
| LDCM0578 | Fragment27 | Ramos | C636(1.26); C364(0.65); C447(0.96); C471(1.00) | LDD2197 | [13] |
| LDCM0586 | Fragment28 | Ramos | C636(0.93); C364(0.56); C447(1.39); C471(0.38) | LDD2198 | [13] |
| LDCM0588 | Fragment30 | Ramos | C636(2.25); C364(0.70); C447(0.78); C561(2.17) | LDD2199 | [13] |
| LDCM0589 | Fragment31 | Ramos | C636(1.91); C364(0.78); C447(1.29); C561(1.61) | LDD2200 | [13] |
| LDCM0590 | Fragment32 | Ramos | C636(2.79); C364(0.41); C447(0.89); C561(0.46) | LDD2201 | [13] |
| LDCM0468 | Fragment33 | Ramos | C636(1.18); C364(0.50); C447(0.67); C561(2.96) | LDD2202 | [13] |
| LDCM0596 | Fragment38 | Ramos | C636(2.14); C364(0.78); C447(0.72); C471(1.17) | LDD2203 | [13] |
| LDCM0566 | Fragment4 | Ramos | C636(1.87); C364(0.91); C447(1.67); C561(1.43) | LDD2184 | [13] |
| LDCM0610 | Fragment52 | Ramos | C636(1.66); C364(0.83); C447(0.65); C561(0.36) | LDD2204 | [13] |
| LDCM0614 | Fragment56 | Ramos | C636(1.47); C364(0.76); C447(0.75); C561(2.29) | LDD2205 | [13] |
| LDCM0617 | Fragment63-S | Jurkat | _(20.00) | LDD1490 | [3] |
| LDCM0569 | Fragment7 | Jurkat | _(5.64) | LDD1485 | [3] |
| LDCM0571 | Fragment9 | Ramos | C636(9.80); C364(0.65); C447(3.05); C561(0.55) | LDD2188 | [13] |
| LDCM0107 | IAA | HeLa | C364(0.00); C636(0.00); C447(0.00) | LDD0221 | [8] |
| LDCM0022 | KB02 | Ramos | C636(1.37); C364(0.79); C447(1.05); C561(0.77) | LDD2182 | [13] |
| LDCM0023 | KB03 | Ramos | C636(4.34); C364(1.24); C447(1.74); C561(1.15) | LDD2183 | [13] |
| LDCM0024 | KB05 | COLO792 | C636(4.54); C364(1.70) | LDD3310 | [1] |
| LDCM0109 | NEM | HeLa | N.A. | LDD0232 | [8] |
| LDCM0131 | RA190 | MM1.R | C581(1.55) | LDD0304 | [14] |
| LDCM0096 | SAHA | K562 | 4.20 | LDD0269 | [2] |
The Interaction Atlas With This Target
References
