Details of the Target
General Information of Target
| Target ID | LDTP08448 | |||||
|---|---|---|---|---|---|---|
| Target Name | ELKS/Rab6-interacting/CAST family member 1 (ERC1) | |||||
| Gene Name | ERC1 | |||||
| Gene ID | 23085 | |||||
| Synonyms |
ELKS; KIAA1081; RAB6IP2; ELKS/Rab6-interacting/CAST family member 1; ERC-1; Rab6-interacting protein 2 |
|||||
| 3D Structure | ||||||
| Sequence |
MYGSARSVGKVEPSSQSPGRSPRLPRSPRLGHRRTNSTGGSSGSSVGGGSGKTLSMENIQ
SLNAAYATSGPMYLSDHENVGSETPKSTMTLGRSGGRLPYGVRMTAMGSSPNIASSGVAS DTIAFGEHHLPPVSMASTVPHSLRQARDNTIMDLQTQLKEVLRENDLLRKDVEVKESKLS SSMNSIKTFWSPELKKERALRKDEASKITIWKEQYRVVQEENQHMQMTIQALQDELRIQR DLNQLFQQDSSSRTGEPCVAELTEENFQRLHAEHERQAKELFLLRKTLEEMELRIETQKQ TLNARDESIKKLLEMLQSKGLSAKATEEDHERTRRLAEAEMHVHHLESLLEQKEKENSML REEMHRRFENAPDSAKTKALQTVIEMKDSKISSMERGLRDLEEEIQMLKSNGALSTEERE EEMKQMEVYRSHSKFMKNKVEQLKEELSSKEAQWEELKKKAAGLQAEIGQVKQELSRKDT ELLALQTKLETLTNQFSDSKQHIEVLKESLTAKEQRAAILQTEVDALRLRLEEKETMLNK KTKQIQDMAEEKGTQAGEIHDLKDMLDVKERKVNVLQKKIENLQEQLRDKEKQMSSLKER VKSLQADTTNTDTALTTLEEALAEKERTIERLKEQRDRDEREKQEEIDNYKKDLKDLKEK VSLLQGDLSEKEASLLDLKEHASSLASSGLKKDSRLKTLEIALEQKKEECLKMESQLKKA HEAALEARASPEMSDRIQHLEREITRYKDESSKAQAEVDRLLEILKEVENEKNDKDKKIA ELERQVKDQNKKVANLKHKEQVEKKKSAQMLEEARRREDNLNDSSQQLQDSLRKKDDRIE ELEEALRESVQITAEREMVLAQEESARTNAEKQVEELLMAMEKVKQELESMKAKLSSTQQ SLAEKETHLTNLRAERRKHLEEVLEMKQEALLAAISEKDANIALLELSSSKKKTQEEVAA LKREKDRLVQQLKQQTQNRMKLMADNYEDDHFKSSHSNQTNHKPSPDQIIQPLLELDQNR SKLKLYIGHLTTLCHDRDPLILRGLTPPASYNLDDDQAAWENELQKMTRGQLQDELEKGE RDNAELQEFANAILQQIADHCPDILEQVVNALEESS |
|||||
| Target Bioclass |
Other
|
|||||
| Subcellular location |
Cytoplasm, cytoskeleton, microtubule organizing center, centrosome
|
|||||
| Function |
Regulatory subunit of the IKK complex. Probably recruits IkappaBalpha/NFKBIA to the complex. May be involved in the organization of the cytomatrix at the nerve terminals active zone (CAZ) which regulates neurotransmitter release. May be involved in vesicle trafficking at the CAZ. May be involved in Rab-6 regulated endosomes to Golgi transport.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| 769P | SNV: p.N941I | . | |||
| AN3CA | Deletion: p.K953del | . | |||
| C4II | SNV: p.E1061K | . | |||
| CCK81 | SNV: p.R396C | . | |||
| COLO792 | SNV: p.R979Ter | . | |||
| HCT15 | SNV: p.K927E | . | |||
| HT115 | SNV: p.D656Y | . | |||
| IGR37 | SNV: p.H721Q | . | |||
| IGR39 | SNV: p.H721Q | . | |||
| KYSE30 | SNV: p.S669L | DBIA Probe Info | |||
| MCC13 | SNV: p.S948L | . | |||
| MFE319 | SNV: p.L401P; p.M565I | . | |||
| PF382 | SNV: p.D171G | . | |||
| RKO | Deletion: p.G96DfsTer9 | . | |||
| RL952 | SNV: p.E256K | . | |||
| SKOV3 | SNV: p.Y747Ter | . | |||
| TE4 | SNV: p.E83K | . | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
14.21 | LDD0402 | [2] | |
|
STPyne Probe Info |
 |
K178(10.00); K212(10.00); K311(1.23); K319(10.00) | LDD0277 | [3] | |
|
DBIA Probe Info |
 |
C258(2.25) | LDD3320 | [4] | |
|
AHL-Pu-1 Probe Info |
 |
C258(2.29) | LDD0168 | [5] | |
|
5E-2FA Probe Info |
 |
N.A. | LDD2235 | [6] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0165 | [7] | |
|
Lodoacetamide azide Probe Info |
 |
N.A. | LDD0037 | [8] | |
|
IPM Probe Info |
 |
N.A. | LDD0025 | [9] | |
|
NAIA_4 Probe Info |
 |
N.A. | LDD2226 | [10] | |
|
TFBX Probe Info |
 |
N.A. | LDD0027 | [9] | |
|
NHS Probe Info |
 |
N.A. | LDD0010 | [11] | |
|
VSF Probe Info |
 |
N.A. | LDD0007 | [11] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0217 | [12] | |
|
Methacrolein Probe Info |
 |
N.A. | LDD0218 | [12] | |
|
W1 Probe Info |
 |
N.A. | LDD0236 | [13] | |
|
AOyne Probe Info |
 |
11.30 | LDD0443 | [14] | |
|
NAIA_5 Probe Info |
 |
N.A. | LDD2223 | [10] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C052 Probe Info |
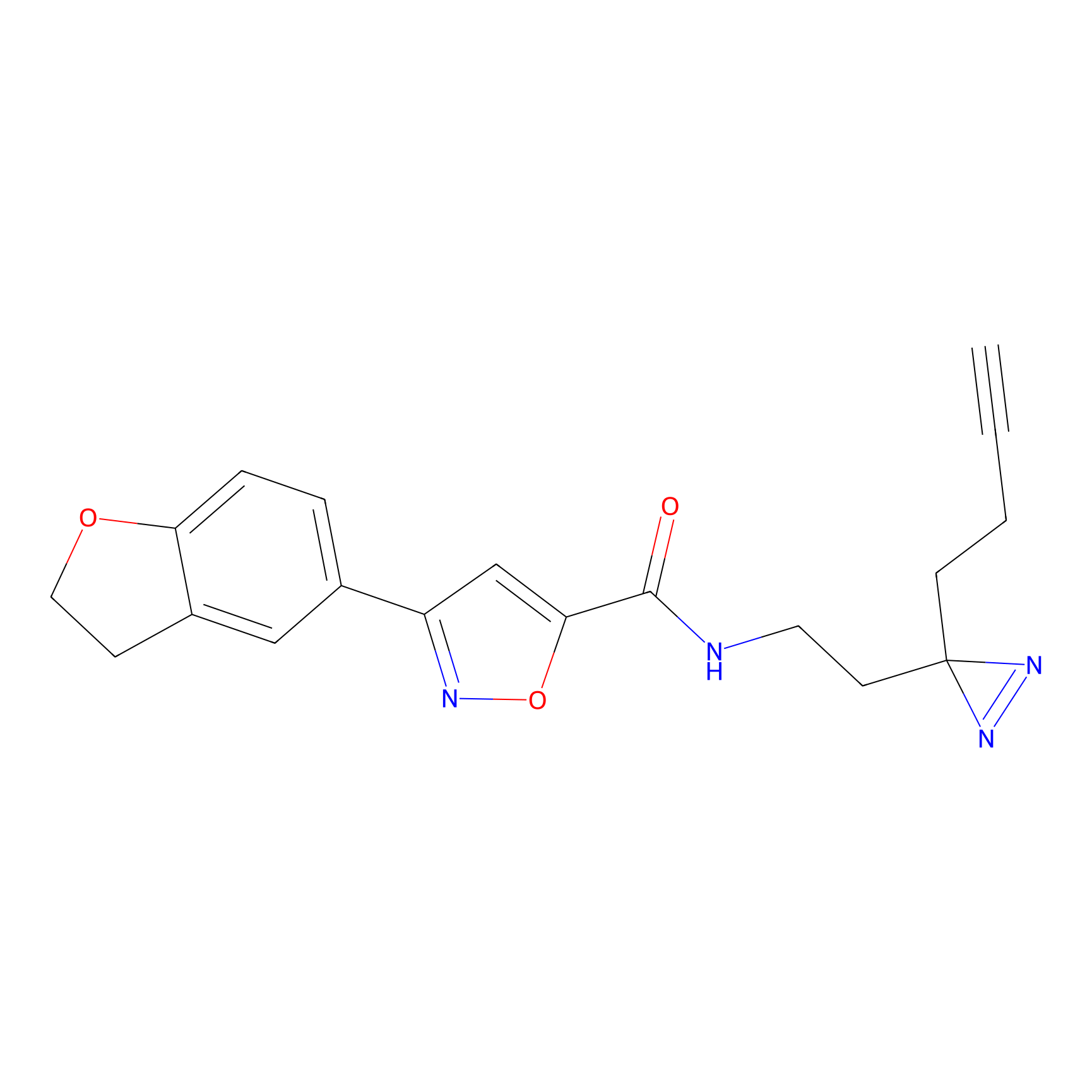 |
5.66 | LDD1750 | [15] | |
|
C391 Probe Info |
 |
13.36 | LDD2050 | [15] | |
|
STS-2 Probe Info |
 |
N.A. | LDD0139 | [16] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0025 | 4SU-RNA | HEK-293T | C258(2.29) | LDD0168 | [5] |
| LDCM0214 | AC1 | HEK-293T | C258(0.95) | LDD1507 | [17] |
| LDCM0215 | AC10 | HEK-293T | C258(1.05) | LDD1508 | [17] |
| LDCM0226 | AC11 | HEK-293T | C258(1.06) | LDD1509 | [17] |
| LDCM0237 | AC12 | HEK-293T | C258(1.02) | LDD1510 | [17] |
| LDCM0259 | AC14 | HEK-293T | C258(1.02) | LDD1512 | [17] |
| LDCM0270 | AC15 | HEK-293T | C258(1.08) | LDD1513 | [17] |
| LDCM0276 | AC17 | HEK-293T | C258(0.95) | LDD1515 | [17] |
| LDCM0277 | AC18 | HEK-293T | C258(0.93) | LDD1516 | [17] |
| LDCM0278 | AC19 | HEK-293T | C258(1.14) | LDD1517 | [17] |
| LDCM0279 | AC2 | HEK-293T | C258(1.21) | LDD1518 | [17] |
| LDCM0280 | AC20 | HEK-293T | C258(0.96) | LDD1519 | [17] |
| LDCM0281 | AC21 | HEK-293T | C258(1.10) | LDD1520 | [17] |
| LDCM0282 | AC22 | HEK-293T | C258(1.02) | LDD1521 | [17] |
| LDCM0283 | AC23 | HEK-293T | C258(1.10) | LDD1522 | [17] |
| LDCM0284 | AC24 | HEK-293T | C258(0.98) | LDD1523 | [17] |
| LDCM0285 | AC25 | HEK-293T | C258(0.93) | LDD1524 | [17] |
| LDCM0286 | AC26 | HEK-293T | C258(0.95) | LDD1525 | [17] |
| LDCM0287 | AC27 | HEK-293T | C258(1.05) | LDD1526 | [17] |
| LDCM0288 | AC28 | HEK-293T | C258(1.04) | LDD1527 | [17] |
| LDCM0289 | AC29 | HEK-293T | C258(1.02) | LDD1528 | [17] |
| LDCM0290 | AC3 | HEK-293T | C258(0.97) | LDD1529 | [17] |
| LDCM0291 | AC30 | HEK-293T | C258(1.01) | LDD1530 | [17] |
| LDCM0292 | AC31 | HEK-293T | C258(1.19) | LDD1531 | [17] |
| LDCM0293 | AC32 | HEK-293T | C258(0.95) | LDD1532 | [17] |
| LDCM0294 | AC33 | HEK-293T | C258(1.09) | LDD1533 | [17] |
| LDCM0295 | AC34 | HEK-293T | C258(1.08) | LDD1534 | [17] |
| LDCM0296 | AC35 | HEK-293T | C258(1.10) | LDD1535 | [17] |
| LDCM0297 | AC36 | HEK-293T | C258(1.04) | LDD1536 | [17] |
| LDCM0298 | AC37 | HEK-293T | C258(1.05) | LDD1537 | [17] |
| LDCM0299 | AC38 | HEK-293T | C258(0.96) | LDD1538 | [17] |
| LDCM0300 | AC39 | HEK-293T | C258(1.01) | LDD1539 | [17] |
| LDCM0301 | AC4 | HEK-293T | C258(0.98) | LDD1540 | [17] |
| LDCM0302 | AC40 | HEK-293T | C258(0.93) | LDD1541 | [17] |
| LDCM0303 | AC41 | HEK-293T | C258(1.13) | LDD1542 | [17] |
| LDCM0304 | AC42 | HEK-293T | C258(1.29) | LDD1543 | [17] |
| LDCM0305 | AC43 | HEK-293T | C258(1.13) | LDD1544 | [17] |
| LDCM0306 | AC44 | HEK-293T | C258(1.02) | LDD1545 | [17] |
| LDCM0307 | AC45 | HEK-293T | C258(1.12) | LDD1546 | [17] |
| LDCM0308 | AC46 | HEK-293T | C258(0.96) | LDD1547 | [17] |
| LDCM0309 | AC47 | HEK-293T | C258(1.07) | LDD1548 | [17] |
| LDCM0310 | AC48 | HEK-293T | C258(0.98) | LDD1549 | [17] |
| LDCM0311 | AC49 | HEK-293T | C258(1.08) | LDD1550 | [17] |
| LDCM0312 | AC5 | HEK-293T | C258(1.02) | LDD1551 | [17] |
| LDCM0313 | AC50 | HEK-293T | C258(1.31) | LDD1552 | [17] |
| LDCM0314 | AC51 | HEK-293T | C258(1.06) | LDD1553 | [17] |
| LDCM0315 | AC52 | HEK-293T | C258(0.99) | LDD1554 | [17] |
| LDCM0316 | AC53 | HEK-293T | C258(1.12) | LDD1555 | [17] |
| LDCM0317 | AC54 | HEK-293T | C258(1.00) | LDD1556 | [17] |
| LDCM0318 | AC55 | HEK-293T | C258(1.18) | LDD1557 | [17] |
| LDCM0319 | AC56 | HEK-293T | C258(0.94) | LDD1558 | [17] |
| LDCM0320 | AC57 | HEK-293T | C258(1.00) | LDD1559 | [17] |
| LDCM0321 | AC58 | HEK-293T | C258(1.05) | LDD1560 | [17] |
| LDCM0322 | AC59 | HEK-293T | C258(1.04) | LDD1561 | [17] |
| LDCM0323 | AC6 | HEK-293T | C258(0.88) | LDD1562 | [17] |
| LDCM0324 | AC60 | HEK-293T | C258(1.07) | LDD1563 | [17] |
| LDCM0325 | AC61 | HEK-293T | C258(1.00) | LDD1564 | [17] |
| LDCM0326 | AC62 | HEK-293T | C258(0.98) | LDD1565 | [17] |
| LDCM0327 | AC63 | HEK-293T | C258(1.00) | LDD1566 | [17] |
| LDCM0328 | AC64 | HEK-293T | C258(1.05) | LDD1567 | [17] |
| LDCM0334 | AC7 | HEK-293T | C258(1.09) | LDD1568 | [17] |
| LDCM0345 | AC8 | HEK-293T | C258(0.97) | LDD1569 | [17] |
| LDCM0248 | AKOS034007472 | HEK-293T | C258(1.14) | LDD1511 | [17] |
| LDCM0356 | AKOS034007680 | HEK-293T | C258(1.07) | LDD1570 | [17] |
| LDCM0275 | AKOS034007705 | HEK-293T | C258(1.02) | LDD1514 | [17] |
| LDCM0156 | Aniline | NCI-H1299 | N.A. | LDD0404 | [2] |
| LDCM0108 | Chloroacetamide | HeLa | N.A. | LDD0222 | [12] |
| LDCM0367 | CL1 | HEK-293T | C258(1.04) | LDD1571 | [17] |
| LDCM0368 | CL10 | HEK-293T | C258(0.87) | LDD1572 | [17] |
| LDCM0369 | CL100 | HEK-293T | C258(1.02) | LDD1573 | [17] |
| LDCM0370 | CL101 | HEK-293T | C258(1.11) | LDD1574 | [17] |
| LDCM0371 | CL102 | HEK-293T | C258(0.92) | LDD1575 | [17] |
| LDCM0372 | CL103 | HEK-293T | C258(1.12) | LDD1576 | [17] |
| LDCM0373 | CL104 | HEK-293T | C258(1.00) | LDD1577 | [17] |
| LDCM0374 | CL105 | HEK-293T | C258(1.05) | LDD1578 | [17] |
| LDCM0375 | CL106 | HEK-293T | C258(1.03) | LDD1579 | [17] |
| LDCM0376 | CL107 | HEK-293T | C258(1.15) | LDD1580 | [17] |
| LDCM0377 | CL108 | HEK-293T | C258(1.06) | LDD1581 | [17] |
| LDCM0378 | CL109 | HEK-293T | C258(1.00) | LDD1582 | [17] |
| LDCM0379 | CL11 | HEK-293T | C258(1.08) | LDD1583 | [17] |
| LDCM0380 | CL110 | HEK-293T | C258(0.96) | LDD1584 | [17] |
| LDCM0381 | CL111 | HEK-293T | C258(1.07) | LDD1585 | [17] |
| LDCM0382 | CL112 | HEK-293T | C258(1.09) | LDD1586 | [17] |
| LDCM0383 | CL113 | HEK-293T | C258(1.03) | LDD1587 | [17] |
| LDCM0384 | CL114 | HEK-293T | C258(0.97) | LDD1588 | [17] |
| LDCM0385 | CL115 | HEK-293T | C258(1.20) | LDD1589 | [17] |
| LDCM0386 | CL116 | HEK-293T | C258(1.09) | LDD1590 | [17] |
| LDCM0387 | CL117 | HEK-293T | C258(0.98) | LDD1591 | [17] |
| LDCM0388 | CL118 | HEK-293T | C258(1.22) | LDD1592 | [17] |
| LDCM0389 | CL119 | HEK-293T | C258(1.06) | LDD1593 | [17] |
| LDCM0390 | CL12 | HEK-293T | C258(1.02) | LDD1594 | [17] |
| LDCM0391 | CL120 | HEK-293T | C258(1.08) | LDD1595 | [17] |
| LDCM0392 | CL121 | HEK-293T | C258(1.10) | LDD1596 | [17] |
| LDCM0393 | CL122 | HEK-293T | C258(1.09) | LDD1597 | [17] |
| LDCM0394 | CL123 | HEK-293T | C258(1.21) | LDD1598 | [17] |
| LDCM0395 | CL124 | HEK-293T | C258(1.15) | LDD1599 | [17] |
| LDCM0396 | CL125 | HEK-293T | C258(1.05) | LDD1600 | [17] |
| LDCM0397 | CL126 | HEK-293T | C258(0.99) | LDD1601 | [17] |
| LDCM0398 | CL127 | HEK-293T | C258(1.15) | LDD1602 | [17] |
| LDCM0399 | CL128 | HEK-293T | C258(1.13) | LDD1603 | [17] |
| LDCM0400 | CL13 | HEK-293T | C258(1.11) | LDD1604 | [17] |
| LDCM0401 | CL14 | HEK-293T | C258(1.00) | LDD1605 | [17] |
| LDCM0402 | CL15 | HEK-293T | C258(1.07) | LDD1606 | [17] |
| LDCM0403 | CL16 | HEK-293T | C258(1.02) | LDD1607 | [17] |
| LDCM0404 | CL17 | HEK-293T | C258(0.92) | LDD1608 | [17] |
| LDCM0405 | CL18 | HEK-293T | C258(1.22) | LDD1609 | [17] |
| LDCM0406 | CL19 | HEK-293T | C258(1.07) | LDD1610 | [17] |
| LDCM0407 | CL2 | HEK-293T | C258(1.05) | LDD1611 | [17] |
| LDCM0408 | CL20 | HEK-293T | C258(0.92) | LDD1612 | [17] |
| LDCM0409 | CL21 | HEK-293T | C258(1.08) | LDD1613 | [17] |
| LDCM0410 | CL22 | HEK-293T | C258(0.97) | LDD1614 | [17] |
| LDCM0411 | CL23 | HEK-293T | C258(1.23) | LDD1615 | [17] |
| LDCM0412 | CL24 | HEK-293T | C258(0.93) | LDD1616 | [17] |
| LDCM0413 | CL25 | HEK-293T | C258(1.06) | LDD1617 | [17] |
| LDCM0414 | CL26 | HEK-293T | C258(0.99) | LDD1618 | [17] |
| LDCM0415 | CL27 | HEK-293T | C258(1.11) | LDD1619 | [17] |
| LDCM0416 | CL28 | HEK-293T | C258(1.06) | LDD1620 | [17] |
| LDCM0417 | CL29 | HEK-293T | C258(1.04) | LDD1621 | [17] |
| LDCM0418 | CL3 | HEK-293T | C258(1.06) | LDD1622 | [17] |
| LDCM0419 | CL30 | HEK-293T | C258(1.40) | LDD1623 | [17] |
| LDCM0420 | CL31 | HEK-293T | C258(1.13) | LDD1624 | [17] |
| LDCM0421 | CL32 | HEK-293T | C258(1.01) | LDD1625 | [17] |
| LDCM0422 | CL33 | HEK-293T | C258(0.99) | LDD1626 | [17] |
| LDCM0423 | CL34 | HEK-293T | C258(1.01) | LDD1627 | [17] |
| LDCM0424 | CL35 | HEK-293T | C258(1.08) | LDD1628 | [17] |
| LDCM0425 | CL36 | HEK-293T | C258(1.05) | LDD1629 | [17] |
| LDCM0426 | CL37 | HEK-293T | C258(1.02) | LDD1630 | [17] |
| LDCM0428 | CL39 | HEK-293T | C258(1.14) | LDD1632 | [17] |
| LDCM0429 | CL4 | HEK-293T | C258(1.08) | LDD1633 | [17] |
| LDCM0430 | CL40 | HEK-293T | C258(1.07) | LDD1634 | [17] |
| LDCM0431 | CL41 | HEK-293T | C258(0.95) | LDD1635 | [17] |
| LDCM0432 | CL42 | HEK-293T | C258(1.22) | LDD1636 | [17] |
| LDCM0433 | CL43 | HEK-293T | C258(1.06) | LDD1637 | [17] |
| LDCM0434 | CL44 | HEK-293T | C258(1.06) | LDD1638 | [17] |
| LDCM0435 | CL45 | HEK-293T | C258(1.11) | LDD1639 | [17] |
| LDCM0436 | CL46 | HEK-293T | C258(1.02) | LDD1640 | [17] |
| LDCM0437 | CL47 | HEK-293T | C258(1.09) | LDD1641 | [17] |
| LDCM0438 | CL48 | HEK-293T | C258(0.90) | LDD1642 | [17] |
| LDCM0439 | CL49 | HEK-293T | C258(1.21) | LDD1643 | [17] |
| LDCM0440 | CL5 | HEK-293T | C258(0.98) | LDD1644 | [17] |
| LDCM0441 | CL50 | HEK-293T | C258(1.04) | LDD1645 | [17] |
| LDCM0443 | CL52 | HEK-293T | C258(1.11) | LDD1646 | [17] |
| LDCM0444 | CL53 | HEK-293T | C258(1.13) | LDD1647 | [17] |
| LDCM0445 | CL54 | HEK-293T | C258(1.09) | LDD1648 | [17] |
| LDCM0446 | CL55 | HEK-293T | C258(1.09) | LDD1649 | [17] |
| LDCM0447 | CL56 | HEK-293T | C258(1.01) | LDD1650 | [17] |
| LDCM0448 | CL57 | HEK-293T | C258(0.99) | LDD1651 | [17] |
| LDCM0449 | CL58 | HEK-293T | C258(0.96) | LDD1652 | [17] |
| LDCM0450 | CL59 | HEK-293T | C258(1.14) | LDD1653 | [17] |
| LDCM0451 | CL6 | HEK-293T | C258(1.02) | LDD1654 | [17] |
| LDCM0452 | CL60 | HEK-293T | C258(1.02) | LDD1655 | [17] |
| LDCM0453 | CL61 | HEK-293T | C258(1.01) | LDD1656 | [17] |
| LDCM0454 | CL62 | HEK-293T | C258(1.15) | LDD1657 | [17] |
| LDCM0455 | CL63 | HEK-293T | C258(1.13) | LDD1658 | [17] |
| LDCM0456 | CL64 | HEK-293T | C258(1.07) | LDD1659 | [17] |
| LDCM0457 | CL65 | HEK-293T | C258(1.03) | LDD1660 | [17] |
| LDCM0458 | CL66 | HEK-293T | C258(1.06) | LDD1661 | [17] |
| LDCM0459 | CL67 | HEK-293T | C258(1.13) | LDD1662 | [17] |
| LDCM0460 | CL68 | HEK-293T | C258(1.07) | LDD1663 | [17] |
| LDCM0461 | CL69 | HEK-293T | C258(1.09) | LDD1664 | [17] |
| LDCM0462 | CL7 | HEK-293T | C258(1.08) | LDD1665 | [17] |
| LDCM0463 | CL70 | HEK-293T | C258(0.96) | LDD1666 | [17] |
| LDCM0464 | CL71 | HEK-293T | C258(1.12) | LDD1667 | [17] |
| LDCM0465 | CL72 | HEK-293T | C258(1.04) | LDD1668 | [17] |
| LDCM0466 | CL73 | HEK-293T | C258(1.06) | LDD1669 | [17] |
| LDCM0467 | CL74 | HEK-293T | C258(1.06) | LDD1670 | [17] |
| LDCM0469 | CL76 | HEK-293T | C258(1.13) | LDD1672 | [17] |
| LDCM0470 | CL77 | HEK-293T | C258(1.07) | LDD1673 | [17] |
| LDCM0471 | CL78 | HEK-293T | C258(1.16) | LDD1674 | [17] |
| LDCM0472 | CL79 | HEK-293T | C258(1.05) | LDD1675 | [17] |
| LDCM0473 | CL8 | HEK-293T | C258(0.83) | LDD1676 | [17] |
| LDCM0474 | CL80 | HEK-293T | C258(1.05) | LDD1677 | [17] |
| LDCM0475 | CL81 | HEK-293T | C258(1.08) | LDD1678 | [17] |
| LDCM0476 | CL82 | HEK-293T | C258(0.92) | LDD1679 | [17] |
| LDCM0477 | CL83 | HEK-293T | C258(1.05) | LDD1680 | [17] |
| LDCM0478 | CL84 | HEK-293T | C258(1.03) | LDD1681 | [17] |
| LDCM0479 | CL85 | HEK-293T | C258(0.92) | LDD1682 | [17] |
| LDCM0480 | CL86 | HEK-293T | C258(1.02) | LDD1683 | [17] |
| LDCM0481 | CL87 | HEK-293T | C258(1.27) | LDD1684 | [17] |
| LDCM0482 | CL88 | HEK-293T | C258(1.13) | LDD1685 | [17] |
| LDCM0483 | CL89 | HEK-293T | C258(0.87) | LDD1686 | [17] |
| LDCM0484 | CL9 | HEK-293T | C258(1.08) | LDD1687 | [17] |
| LDCM0485 | CL90 | HEK-293T | C258(1.01) | LDD1688 | [17] |
| LDCM0486 | CL91 | HEK-293T | C258(0.96) | LDD1689 | [17] |
| LDCM0487 | CL92 | HEK-293T | C258(0.98) | LDD1690 | [17] |
| LDCM0488 | CL93 | HEK-293T | C258(1.15) | LDD1691 | [17] |
| LDCM0489 | CL94 | HEK-293T | C258(0.99) | LDD1692 | [17] |
| LDCM0490 | CL95 | HEK-293T | C258(1.10) | LDD1693 | [17] |
| LDCM0491 | CL96 | HEK-293T | C258(0.93) | LDD1694 | [17] |
| LDCM0492 | CL97 | HEK-293T | C258(1.04) | LDD1695 | [17] |
| LDCM0493 | CL98 | HEK-293T | C258(1.02) | LDD1696 | [17] |
| LDCM0494 | CL99 | HEK-293T | C258(1.18) | LDD1697 | [17] |
| LDCM0495 | E2913 | HEK-293T | C258(1.22) | LDD1698 | [17] |
| LDCM0625 | F8 | Ramos | C258(1.14) | LDD2187 | [18] |
| LDCM0572 | Fragment10 | Ramos | C258(1.40) | LDD2189 | [18] |
| LDCM0573 | Fragment11 | Ramos | C258(1.24) | LDD2190 | [18] |
| LDCM0574 | Fragment12 | Ramos | C258(0.97) | LDD2191 | [18] |
| LDCM0575 | Fragment13 | Ramos | C258(0.91) | LDD2192 | [18] |
| LDCM0576 | Fragment14 | Ramos | C258(0.99) | LDD2193 | [18] |
| LDCM0579 | Fragment20 | Ramos | C258(1.09) | LDD2194 | [18] |
| LDCM0580 | Fragment21 | Ramos | C258(1.03) | LDD2195 | [18] |
| LDCM0582 | Fragment23 | Ramos | C258(1.02) | LDD2196 | [18] |
| LDCM0578 | Fragment27 | Ramos | C258(0.93) | LDD2197 | [18] |
| LDCM0586 | Fragment28 | Ramos | C258(0.65) | LDD2198 | [18] |
| LDCM0588 | Fragment30 | Ramos | C258(1.14) | LDD2199 | [18] |
| LDCM0589 | Fragment31 | Ramos | C258(0.90) | LDD2200 | [18] |
| LDCM0590 | Fragment32 | Ramos | C258(1.40) | LDD2201 | [18] |
| LDCM0468 | Fragment33 | HEK-293T | C258(1.19) | LDD1671 | [17] |
| LDCM0596 | Fragment38 | Ramos | C258(0.95) | LDD2203 | [18] |
| LDCM0566 | Fragment4 | Ramos | C258(0.79) | LDD2184 | [18] |
| LDCM0427 | Fragment51 | HEK-293T | C258(0.96) | LDD1631 | [17] |
| LDCM0610 | Fragment52 | Ramos | C258(0.88) | LDD2204 | [18] |
| LDCM0614 | Fragment56 | Ramos | C258(0.93) | LDD2205 | [18] |
| LDCM0569 | Fragment7 | Ramos | C258(1.16) | LDD2186 | [18] |
| LDCM0571 | Fragment9 | Ramos | C258(1.17) | LDD2188 | [18] |
| LDCM0107 | IAA | HeLa | H721(0.00); H274(0.00) | LDD0221 | [12] |
| LDCM0022 | KB02 | HEK-293T | C258(0.99) | LDD1492 | [17] |
| LDCM0023 | KB03 | HEK-293T | C258(1.02) | LDD1497 | [17] |
| LDCM0024 | KB05 | RVH421 | C258(2.25) | LDD3320 | [4] |
| LDCM0109 | NEM | HeLa | H908(0.00); H721(0.00) | LDD0223 | [12] |
| LDCM0627 | NUDT7-COV-1 | HEK-293T | C258(0.88) | LDD2206 | [19] |
| LDCM0628 | OTUB2-COV-1 | HEK-293T | C258(0.43) | LDD2207 | [19] |
References
