Details of the Target
General Information of Target
| Target ID | LDTP04793 | |||||
|---|---|---|---|---|---|---|
| Target Name | Poly(rC)-binding protein 3 (PCBP3) | |||||
| Gene Name | PCBP3 | |||||
| Gene ID | 54039 | |||||
| Synonyms |
PCBP3-OT1; PCBP3OT; Poly(rC)-binding protein 3; Alpha-CP3; PCBP3-overlapping transcript; PCBP3-overlapping transcript 1 |
|||||
| 3D Structure | ||||||
| Sequence |
MGEGDAFWAPSVLPHSTLSTLSHHPQPQFGRRMESKVSEGGLNVTLTIRLLMHGKEVGSI
IGKKGETVKKMREESGARINISEGNCPERIVTITGPTDAIFKAFAMIAYKFEEDIINSMS NSPATSKPPVTLRLVVPASQCGSLIGKGGSKIKEIRESTGAQVQVAGDMLPNSTERAVTI SGTPDAIIQCVKQICVVMLESPPKGATIPYRPKPASTPVIFAGGQAYTIQGQYAIPHPDQ LTKLHQLAMQQTPFPPLGQTNPAFPGEKLPLHSSEEAQNLMGQSSGLDASPPASTHELTI PNDLIGCIIGRQGTKINEIRQMSGAQIKIANATEGSSERQITITGTPANISLAQYLINAR LTSEVTGMGTL |
|||||
| Target Bioclass |
Other
|
|||||
| Subcellular location |
Cytoplasm
|
|||||
| Function | Single-stranded nucleic acid binding protein that binds preferentially to oligo dC. | |||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Target Site Mutations in Different Cell Lines
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
15.00 | LDD0402 | [1] | |
|
BTD Probe Info |
 |
C86(9.01) | LDD1699 | [2] | |
|
AZ-9 Probe Info |
 |
E88(1.00) | LDD2208 | [3] | |
|
IPM Probe Info |
 |
C86(0.00); C141(0.00); C195(0.00) | LDD0241 | [4] | |
|
EA-probe Probe Info |
 |
N.A. | LDD0440 | [5] | |
|
DBIA Probe Info |
 |
C141(31.62); C86(14.72) | LDD0209 | [6] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
N.A. | LDD0038 | [7] | |
|
IA-alkyne Probe Info |
 |
C86(0.00); C141(0.00); C195(0.00) | LDD0032 | [8] | |
|
IPIAA_H Probe Info |
 |
C86(0.00); C195(0.00) | LDD0030 | [9] | |
|
IPIAA_L Probe Info |
 |
C195(0.00); C141(0.00); C86(0.00) | LDD0031 | [9] | |
|
Lodoacetamide azide Probe Info |
 |
N.A. | LDD0037 | [7] | |
|
Dyn-2 Probe Info |
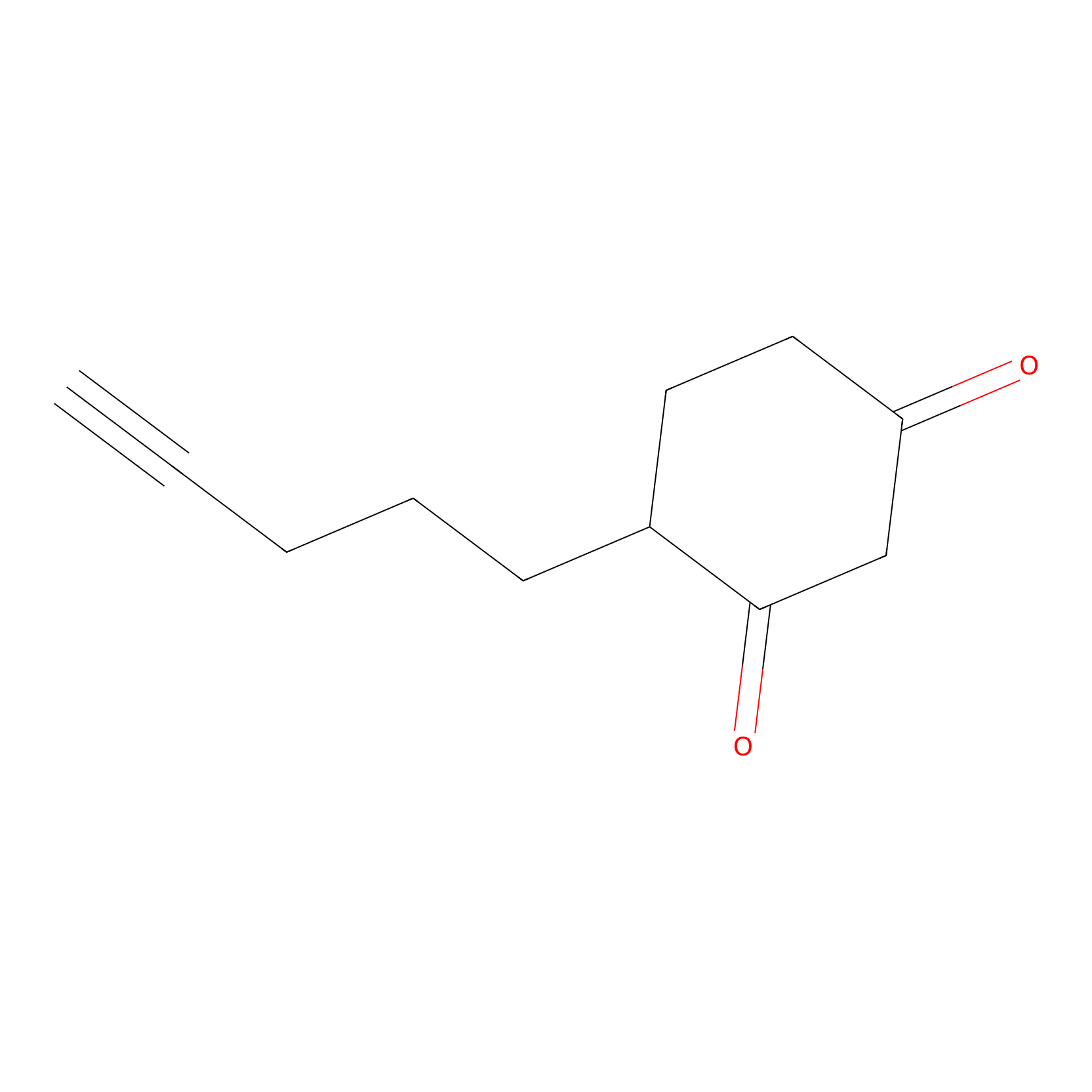 |
N.A. | LDD0013 | [10] | |
|
JW-RF-010 Probe Info |
 |
C195(0.00); C86(0.00); C141(0.00) | LDD0026 | [11] | |
|
TFBX Probe Info |
 |
C195(0.00); C141(0.00); C86(0.00) | LDD0027 | [11] | |
|
WYneO Probe Info |
 |
C86(0.00); C141(0.00) | LDD0022 | [10] | |
|
ENE Probe Info |
 |
C141(0.00); C86(0.00) | LDD0006 | [10] | |
|
PPMS Probe Info |
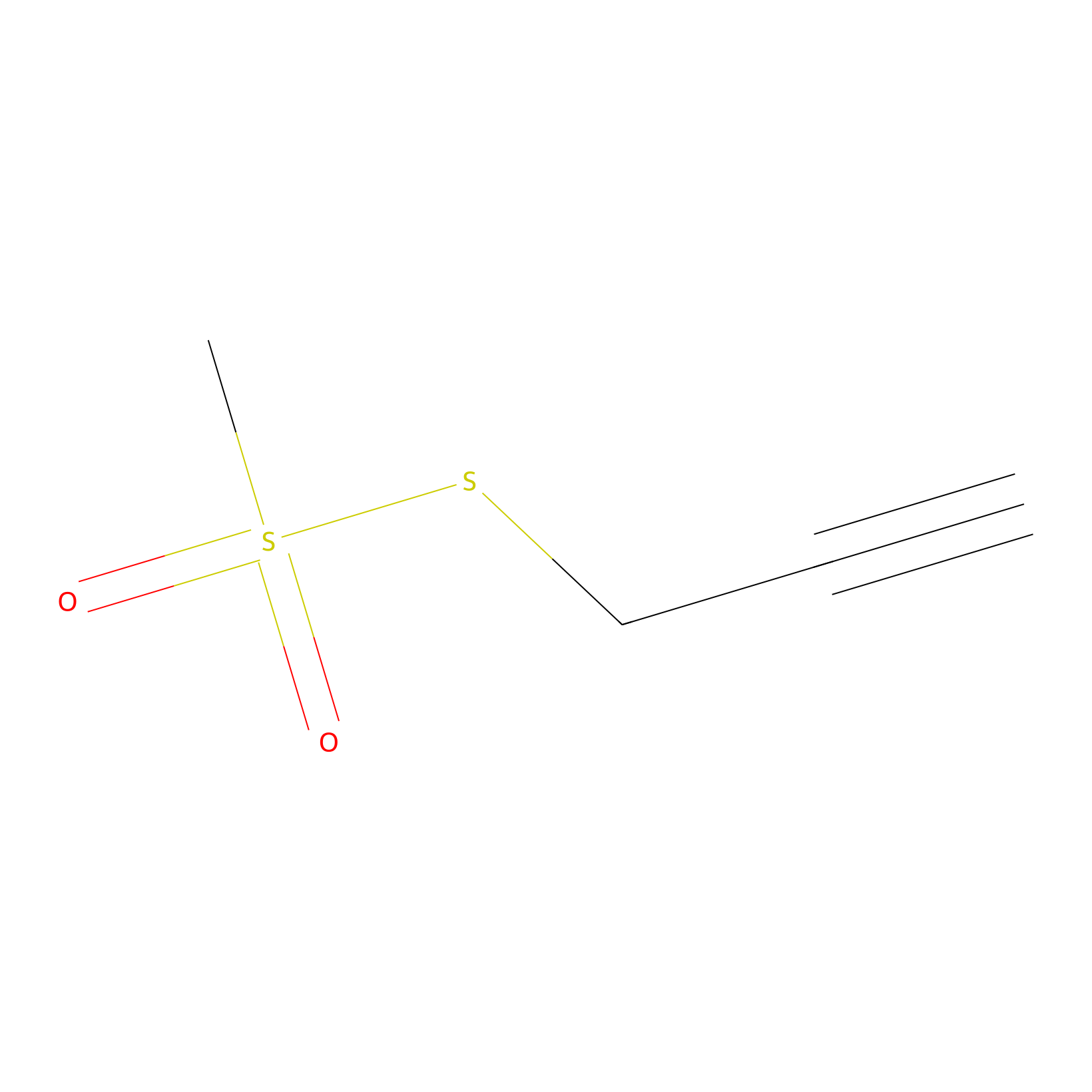 |
C86(0.00); C141(0.00) | LDD0008 | [10] | |
|
Acrolein Probe Info |
 |
C141(0.00); C195(0.00); C86(0.00); H53(0.00) | LDD0217 | [12] | |
|
Cinnamaldehyde Probe Info |
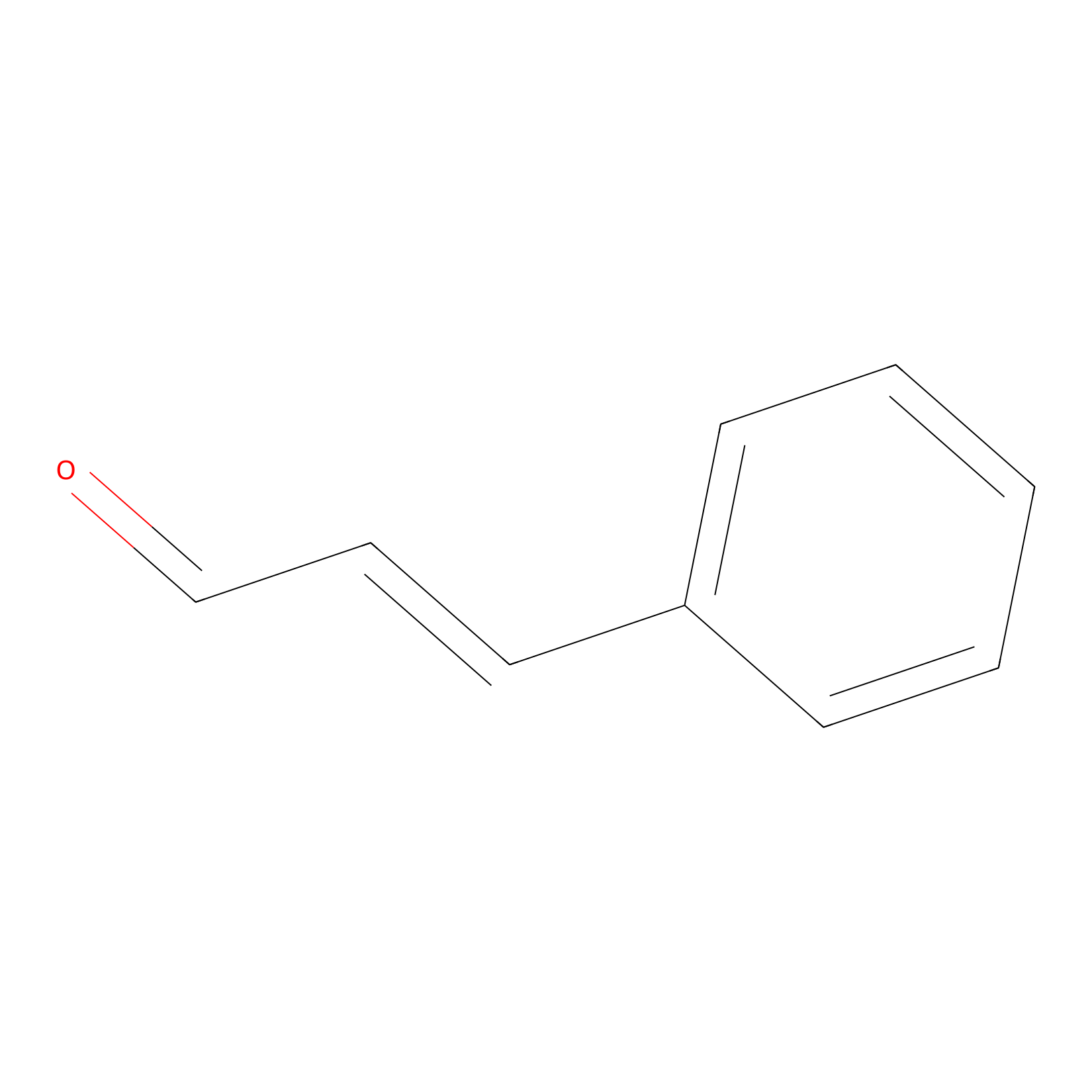 |
C86(0.00); C141(0.00) | LDD0220 | [12] | |
|
Crotonaldehyde Probe Info |
 |
C86(0.00); C141(0.00); C195(0.00) | LDD0219 | [12] | |
|
Methacrolein Probe Info |
 |
C86(0.00); C141(0.00); C195(0.00) | LDD0218 | [12] | |
|
W1 Probe Info |
 |
C141(0.00); C86(0.00); C195(0.00) | LDD0236 | [4] | |
|
NAIA_5 Probe Info |
 |
C86(0.00); C141(0.00); C195(0.00) | LDD2223 | [13] | |
|
HHS-465 Probe Info |
 |
K63(0.00); K55(0.00) | LDD2240 | [14] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
FFF probe13 Probe Info |
 |
9.94 | LDD0475 | [15] | |
|
FFF probe2 Probe Info |
 |
7.02 | LDD0463 | [15] | |
|
JN0003 Probe Info |
 |
16.16 | LDD0469 | [15] | |
|
STS-2 Probe Info |
 |
N.A. | LDD0138 | [16] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0548 | 1-(4-(Benzo[d][1,3]dioxol-5-ylmethyl)piperazin-1-yl)-2-nitroethan-1-one | MDA-MB-231 | C195(0.60); C141(0.44) | LDD2142 | [2] |
| LDCM0519 | 1-(6-methoxy-3,4-dihydroquinolin-1(2H)-yl)-2-nitroethan-1-one | MDA-MB-231 | C195(0.70); C141(0.57) | LDD2112 | [2] |
| LDCM0537 | 2-Cyano-N,N-dimethylacetamide | MDA-MB-231 | C141(0.85) | LDD2130 | [2] |
| LDCM0524 | 2-Cyano-N-(2-morpholin-4-yl-ethyl)-acetamide | MDA-MB-231 | C141(0.70) | LDD2117 | [2] |
| LDCM0558 | 2-Cyano-N-phenylacetamide | MDA-MB-231 | C195(1.45) | LDD2152 | [2] |
| LDCM0510 | 3-(4-(Hydroxydiphenylmethyl)piperidin-1-yl)-3-oxopropanenitrile | MDA-MB-231 | C141(1.31) | LDD2103 | [2] |
| LDCM0539 | 3-(4-Isopropylpiperazin-1-yl)-3-oxopropanenitrile | MDA-MB-231 | C141(0.48) | LDD2132 | [2] |
| LDCM0538 | 4-(Cyanoacetyl)morpholine | MDA-MB-231 | C141(0.66) | LDD2131 | [2] |
| LDCM0545 | Acetamide | MDA-MB-231 | C141(0.41) | LDD2138 | [2] |
| LDCM0520 | AKOS000195272 | MDA-MB-231 | C195(0.90) | LDD2113 | [2] |
| LDCM0156 | Aniline | NCI-H1299 | 11.79 | LDD0403 | [1] |
| LDCM0020 | ARS-1620 | HCC44 | C141(1.05); C86(0.93) | LDD2171 | [17] |
| LDCM0498 | BS-3668 | MDA-MB-231 | C195(1.15); C141(0.46) | LDD2091 | [2] |
| LDCM0108 | Chloroacetamide | HeLa | C195(0.00); C141(0.00); C86(0.00); H53(0.00) | LDD0222 | [12] |
| LDCM0632 | CL-Sc | Hep-G2 | C195(1.06) | LDD2227 | [13] |
| LDCM0175 | Ethacrynic acid | HeLa | N.A. | LDD0440 | [5] |
| LDCM0573 | Fragment11 | Ramos | C195(6.79) | LDD2190 | [18] |
| LDCM0576 | Fragment14 | Ramos | C195(0.30) | LDD2193 | [18] |
| LDCM0586 | Fragment28 | Ramos | C195(1.14) | LDD2198 | [18] |
| LDCM0569 | Fragment7 | Ramos | C195(0.38) | LDD2186 | [18] |
| LDCM0107 | IAA | HeLa | C195(0.00); C141(0.00); C86(0.00); H53(0.00) | LDD0221 | [12] |
| LDCM0022 | KB02 | Ramos | C195(0.28) | LDD2182 | [18] |
| LDCM0023 | KB03 | Jurkat | C141(31.62); C86(14.72) | LDD0209 | [6] |
| LDCM0024 | KB05 | COLO792 | C141(2.68); C195(1.38) | LDD3310 | [19] |
| LDCM0509 | N-(4-bromo-3,5-dimethylphenyl)-2-nitroacetamide | MDA-MB-231 | C141(1.13) | LDD2102 | [2] |
| LDCM0528 | N-(4-bromophenyl)-2-cyano-N-phenylacetamide | MDA-MB-231 | C195(1.26); C141(0.61) | LDD2121 | [2] |
| LDCM0109 | NEM | HeLa | C141(0.00); H53(0.00); C86(0.00) | LDD0223 | [12] |
| LDCM0496 | Nucleophilic fragment 11a | MDA-MB-231 | C141(0.55) | LDD2089 | [2] |
| LDCM0497 | Nucleophilic fragment 11b | MDA-MB-231 | C141(1.34) | LDD2090 | [2] |
| LDCM0500 | Nucleophilic fragment 13a | MDA-MB-231 | C141(0.98) | LDD2093 | [2] |
| LDCM0501 | Nucleophilic fragment 13b | MDA-MB-231 | C141(1.26) | LDD2094 | [2] |
| LDCM0504 | Nucleophilic fragment 15a | MDA-MB-231 | C195(1.19); C141(0.63) | LDD2097 | [2] |
| LDCM0505 | Nucleophilic fragment 15b | MDA-MB-231 | C195(0.75); C141(0.72) | LDD2098 | [2] |
| LDCM0506 | Nucleophilic fragment 16a | MDA-MB-231 | C195(1.94); C141(0.80) | LDD2099 | [2] |
| LDCM0507 | Nucleophilic fragment 16b | MDA-MB-231 | C141(0.48) | LDD2100 | [2] |
| LDCM0508 | Nucleophilic fragment 17a | MDA-MB-231 | C195(1.01) | LDD2101 | [2] |
| LDCM0511 | Nucleophilic fragment 18b | MDA-MB-231 | C141(0.50) | LDD2104 | [2] |
| LDCM0512 | Nucleophilic fragment 19a | MDA-MB-231 | C141(1.81) | LDD2105 | [2] |
| LDCM0513 | Nucleophilic fragment 19b | MDA-MB-231 | C141(0.54) | LDD2106 | [2] |
| LDCM0514 | Nucleophilic fragment 20a | MDA-MB-231 | C195(1.11); C141(0.71) | LDD2107 | [2] |
| LDCM0515 | Nucleophilic fragment 20b | MDA-MB-231 | C141(0.52) | LDD2108 | [2] |
| LDCM0516 | Nucleophilic fragment 21a | MDA-MB-231 | C195(0.77); C141(0.52) | LDD2109 | [2] |
| LDCM0518 | Nucleophilic fragment 22a | MDA-MB-231 | C195(1.56) | LDD2111 | [2] |
| LDCM0521 | Nucleophilic fragment 23b | MDA-MB-231 | C141(0.83) | LDD2114 | [2] |
| LDCM0525 | Nucleophilic fragment 25b | MDA-MB-231 | C141(0.52) | LDD2118 | [2] |
| LDCM0526 | Nucleophilic fragment 26a | MDA-MB-231 | C141(2.73) | LDD2119 | [2] |
| LDCM0527 | Nucleophilic fragment 26b | MDA-MB-231 | C195(0.52); C141(0.67) | LDD2120 | [2] |
| LDCM0529 | Nucleophilic fragment 27b | MDA-MB-231 | C141(0.40) | LDD2122 | [2] |
| LDCM0530 | Nucleophilic fragment 28a | MDA-MB-231 | C195(1.20); C141(0.88) | LDD2123 | [2] |
| LDCM0532 | Nucleophilic fragment 29a | MDA-MB-231 | C195(1.13); C141(0.67) | LDD2125 | [2] |
| LDCM0533 | Nucleophilic fragment 29b | MDA-MB-231 | C141(0.50) | LDD2126 | [2] |
| LDCM0534 | Nucleophilic fragment 30a | MDA-MB-231 | C195(1.15); C141(0.79) | LDD2127 | [2] |
| LDCM0535 | Nucleophilic fragment 30b | MDA-MB-231 | C195(0.65); C141(0.75); C86(1.05) | LDD2128 | [2] |
| LDCM0536 | Nucleophilic fragment 31 | MDA-MB-231 | C195(1.71); C141(0.98) | LDD2129 | [2] |
| LDCM0540 | Nucleophilic fragment 35 | MDA-MB-231 | C195(0.74); C141(0.61) | LDD2133 | [2] |
| LDCM0541 | Nucleophilic fragment 36 | MDA-MB-231 | C195(0.57) | LDD2134 | [2] |
| LDCM0542 | Nucleophilic fragment 37 | MDA-MB-231 | C195(1.58); C141(1.14) | LDD2135 | [2] |
| LDCM0543 | Nucleophilic fragment 38 | MDA-MB-231 | C195(1.73); C141(0.94) | LDD2136 | [2] |
| LDCM0544 | Nucleophilic fragment 39 | MDA-MB-231 | C195(1.15); C141(0.74) | LDD2137 | [2] |
| LDCM0546 | Nucleophilic fragment 40 | MDA-MB-231 | C195(1.11); C141(0.55) | LDD2140 | [2] |
| LDCM0549 | Nucleophilic fragment 43 | MDA-MB-231 | C141(0.83) | LDD2143 | [2] |
| LDCM0550 | Nucleophilic fragment 5a | MDA-MB-231 | C195(1.89); C141(3.14) | LDD2144 | [2] |
| LDCM0552 | Nucleophilic fragment 6a | MDA-MB-231 | C195(0.93); C141(0.75) | LDD2146 | [2] |
| LDCM0553 | Nucleophilic fragment 6b | MDA-MB-231 | C195(1.00); C141(1.89) | LDD2147 | [2] |
| LDCM0554 | Nucleophilic fragment 7a | MDA-MB-231 | C195(0.44) | LDD2148 | [2] |
| LDCM0556 | Nucleophilic fragment 8a | MDA-MB-231 | C195(0.82); C141(0.40) | LDD2150 | [2] |
| LDCM0559 | Nucleophilic fragment 9b | MDA-MB-231 | C195(1.61) | LDD2153 | [2] |
| LDCM0021 | THZ1 | HCT 116 | C141(1.05); C86(0.93) | LDD2173 | [17] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| E3 ubiquitin-protein ligase RNF4 (RNF4) | . | P78317 | |||
Other
References
