Details of the Target
General Information of Target
| Target ID | LDTP02108 | |||||
|---|---|---|---|---|---|---|
| Target Name | Large ribosomal subunit protein P2 (RPLP2) | |||||
| Gene Name | RPLP2 | |||||
| Gene ID | 6181 | |||||
| Synonyms |
D11S2243E; RPP2; Large ribosomal subunit protein P2; 60S acidic ribosomal protein P2; Renal carcinoma antigen NY-REN-44 |
|||||
| 3D Structure | ||||||
| Sequence |
MRYVASYLLAALGGNSSPSAKDIKKILDSVGIEADDDRLNKVISELNGKNIEDVIAQGIG
KLASVPAGGAVAVSAAPGSAAPAAGSAPAAAEEKKDEKKEESEESDDDMGFGLFD |
|||||
| Target Bioclass |
Other
|
|||||
| Family |
Eukaryotic ribosomal protein P1/P2 family
|
|||||
| Function | Plays an important role in the elongation step of protein synthesis. | |||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
P2 Probe Info |
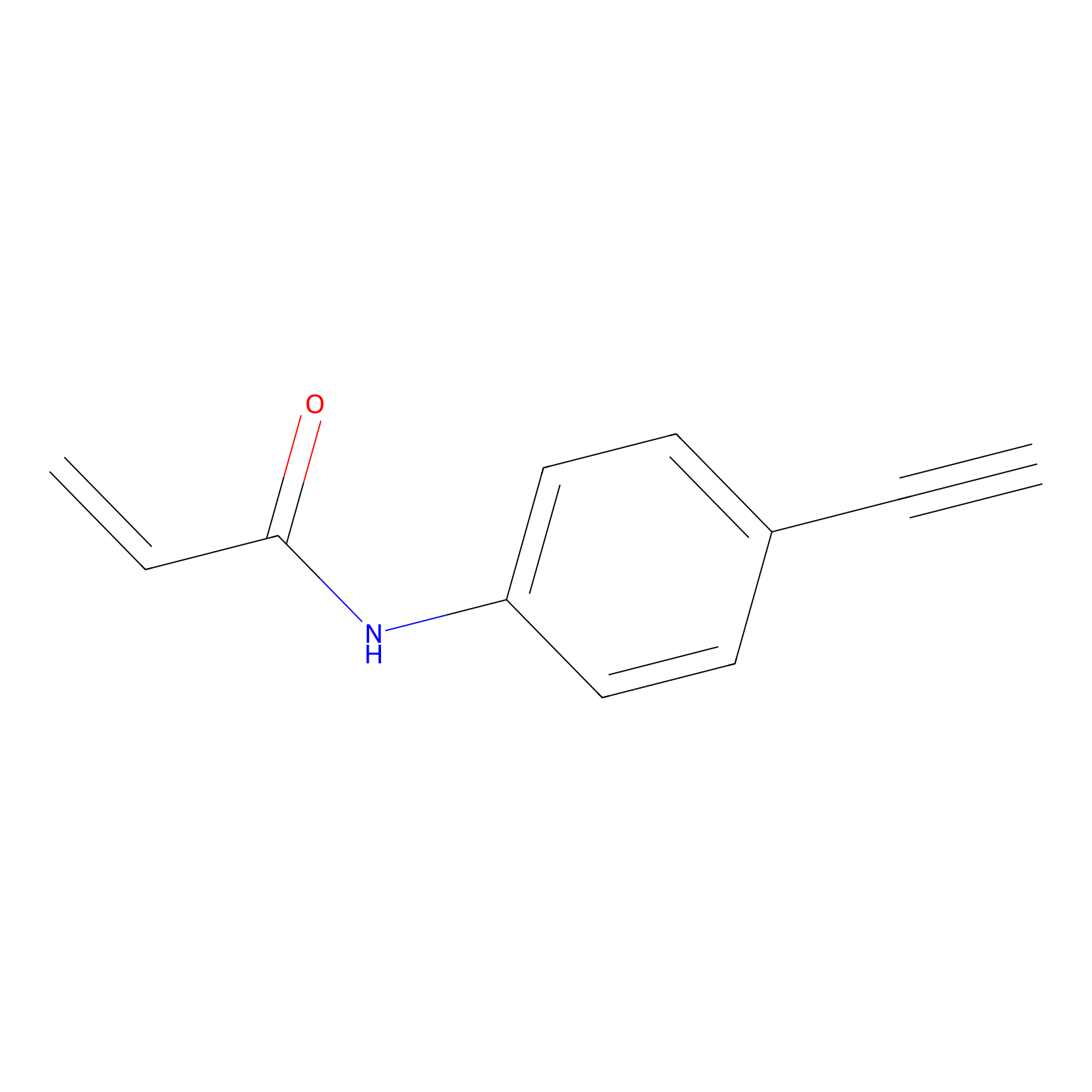 |
10.00 | LDD0449 | [1] | |
|
P8 Probe Info |
 |
10.00 | LDD0451 | [1] | |
|
AZ-9 Probe Info |
 |
3.93 | LDD0393 | [2] | |
|
FBPP2 Probe Info |
 |
2.42 | LDD0318 | [3] | |
|
C-Sul Probe Info |
 |
5.81 | LDD0066 | [4] | |
|
TH211 Probe Info |
 |
Y3(12.24) | LDD0257 | [5] | |
|
ONAyne Probe Info |
 |
K25(0.00); K41(0.00) | LDD0273 | [6] | |
|
OPA-S-S-alkyne Probe Info |
 |
K25(2.10) | LDD3494 | [7] | |
|
D5yne Probe Info |
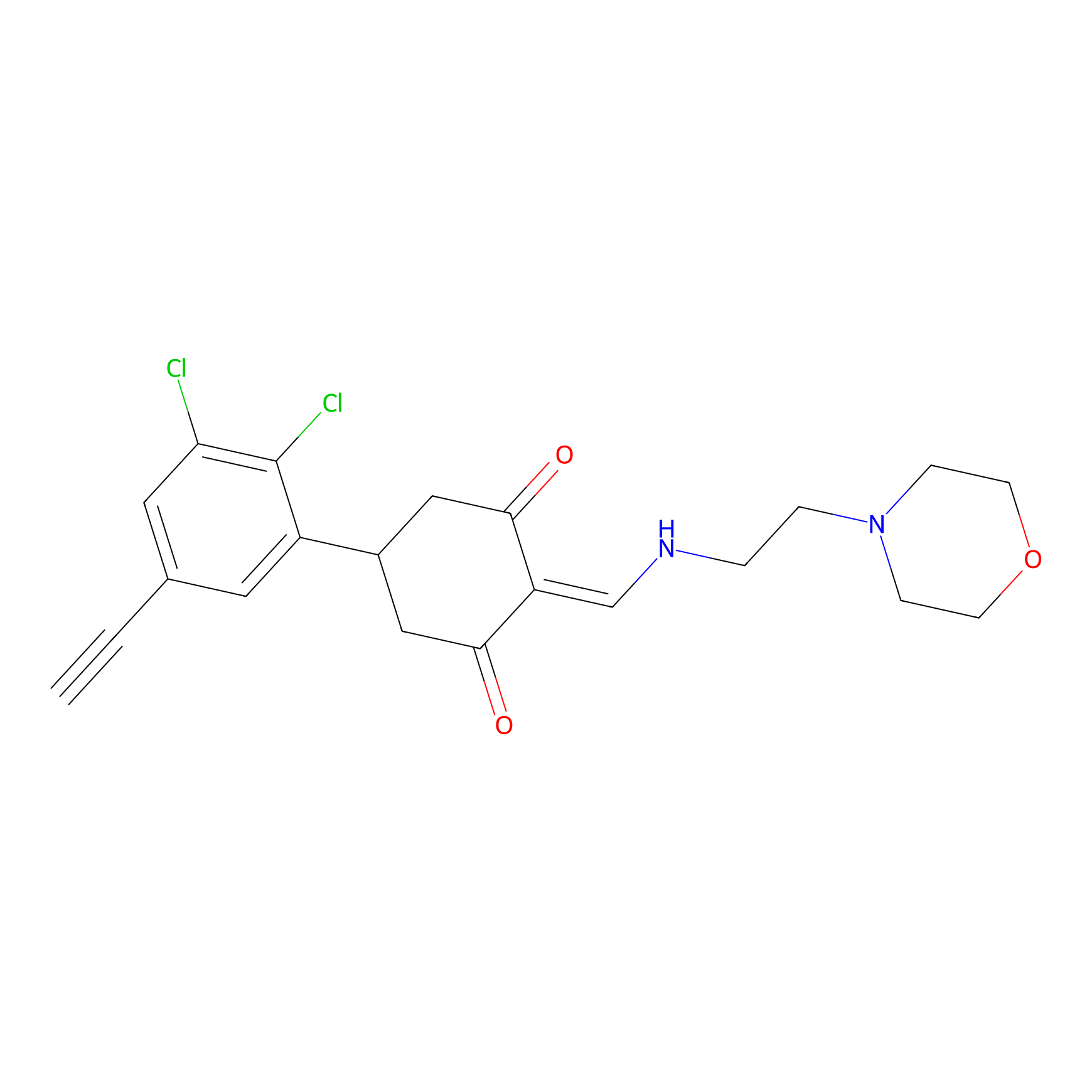 |
1.66 | LDD0055 | [8] | |
|
ATP probe Probe Info |
 |
K25(0.00); K94(0.00); K95(0.00); K98(0.00) | LDD0199 | [9] | |
|
ATP probe Probe Info |
 |
N.A. | LDD0035 | [10] | |
|
1d-yne Probe Info |
 |
N.A. | LDD0356 | [11] | |
|
NHS Probe Info |
 |
K95(0.00); K49(0.00); K94(0.00); K25(0.00) | LDD0010 | [12] | |
|
SF Probe Info |
 |
Y7(0.00); K21(0.00); K25(0.00) | LDD0028 | [13] | |
|
STPyne Probe Info |
 |
N.A. | LDD0009 | [12] | |
|
1c-yne Probe Info |
 |
K94(0.00); K25(0.00); K49(0.00); K41(0.00) | LDD0228 | [11] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
FFF probe14 Probe Info |
 |
5.50 | LDD0477 | [14] | |
|
Diazir Probe Info |
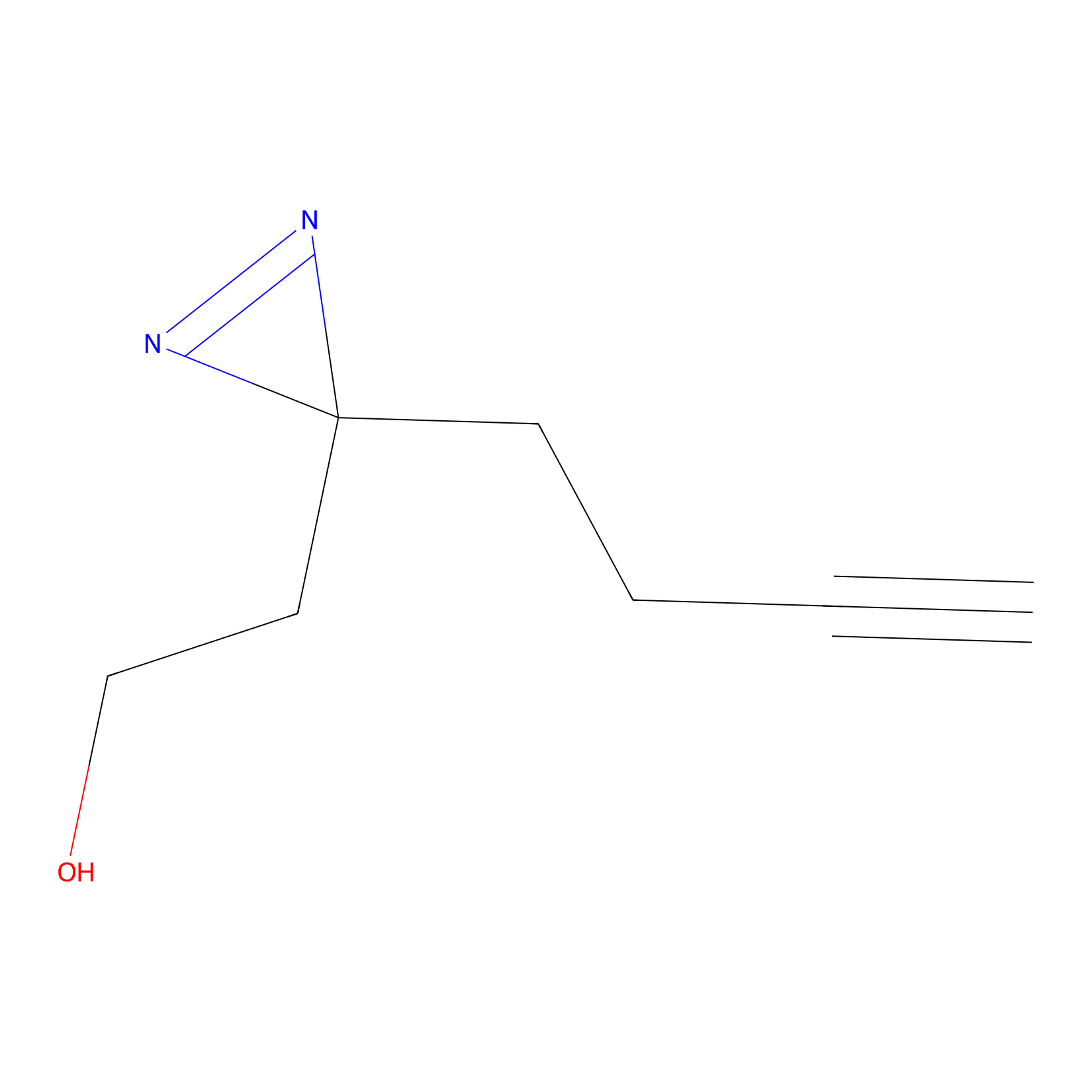 |
E52(0.00); E93(0.00) | LDD0011 | [12] | |
Competitor(s) Related to This Target
The Interaction Atlas With This Target
References
