Details of the Target
General Information of Target
| Target ID | LDTP01012 | |||||
|---|---|---|---|---|---|---|
| Target Name | Selenoprotein F (SELENOF) | |||||
| Gene Name | SELENOF | |||||
| Gene ID | 9403 | |||||
| Synonyms |
15-Sep; Selenoprotein F; 15 kDa selenoprotein |
|||||
| 3D Structure | ||||||
| Sequence |
MVAMAAGPSGCLVPAFGLRLLLATVLQAVSAFGAEFSSEACRELGFSSNLLCSSCDLLGQ
FNLLQLDPDCRGCCQEEAQFETKKLYAGAILEVCGUKLGRFPQVQAFVRSDKPKLFRGLQ IKYVRGSDPVLKLLDDNGNIAEELSILKWNTDSVEEFLSEKLERI |
|||||
| Target Bioclass |
Other
|
|||||
| Family |
Selenoprotein M/F family
|
|||||
| Subcellular location |
Endoplasmic reticulum lumen
|
|||||
| Function |
May be involved in redox reactions associated with the formation of disulfide bonds. May contribute to the quality control of protein folding in the endoplasmic reticulum. May regulate protein folding by enhancing the catalytic activity of UGGT1/UGCGL1 and UGGT2/UGCGL2.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Target Site Mutations in Different Cell Lines
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
YN-1 Probe Info |
 |
100.00 | LDD0444 | [1] | |
|
Johansson_61 Probe Info |
 |
_(7.83) | LDD1485 | [2] | |
|
YY4-yne Probe Info |
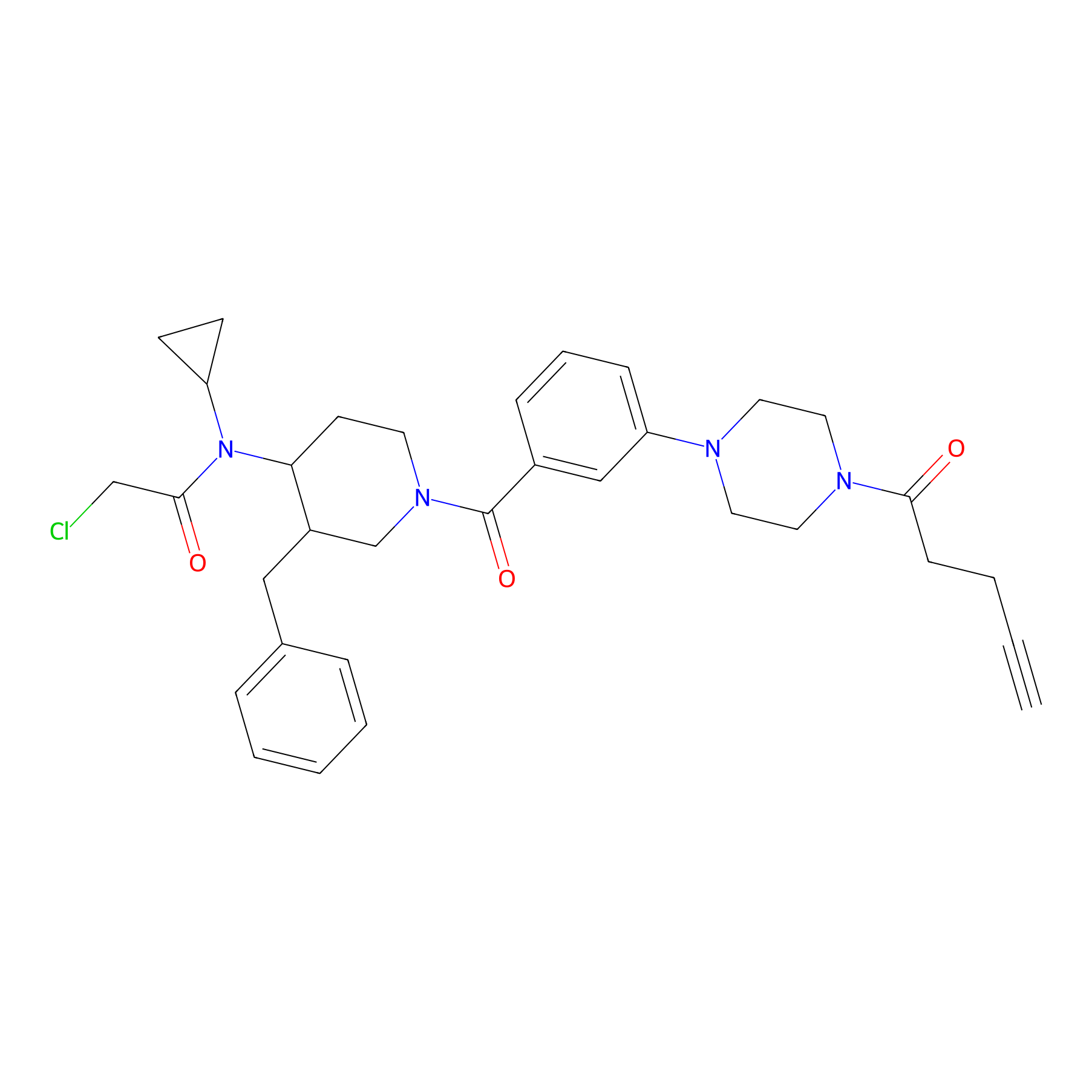 |
3.27 | LDD0400 | [3] | |
|
DBIA Probe Info |
 |
C70(4.70); C71(4.70) | LDD0080 | [4] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0217 | [5] | |
|
Crotonaldehyde Probe Info |
 |
N.A. | LDD0219 | [5] | |
|
Methacrolein Probe Info |
 |
C41(0.00); C74(0.00) | LDD0218 | [5] | |
|
AOyne Probe Info |
 |
15.00 | LDD0443 | [6] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0237 | AC12 | HEK-293T | C73(1.14) | LDD1510 | [7] |
| LDCM0259 | AC14 | HEK-293T | C74(1.03) | LDD1512 | [7] |
| LDCM0270 | AC15 | HEK-293T | C73(1.16) | LDD1513 | [7] |
| LDCM0280 | AC20 | HEK-293T | C73(1.08) | LDD1519 | [7] |
| LDCM0282 | AC22 | HEK-293T | C74(1.01) | LDD1521 | [7] |
| LDCM0283 | AC23 | HEK-293T | C73(0.92) | LDD1522 | [7] |
| LDCM0284 | AC24 | HEK-293T | C74(1.02) | LDD1523 | [7] |
| LDCM0288 | AC28 | HEK-293T | C73(1.10) | LDD1527 | [7] |
| LDCM0291 | AC30 | HEK-293T | C74(0.90) | LDD1530 | [7] |
| LDCM0292 | AC31 | HEK-293T | C73(0.87) | LDD1531 | [7] |
| LDCM0293 | AC32 | HEK-293T | C74(0.92) | LDD1532 | [7] |
| LDCM0297 | AC36 | HEK-293T | C73(1.05) | LDD1536 | [7] |
| LDCM0299 | AC38 | HEK-293T | C74(1.04) | LDD1538 | [7] |
| LDCM0300 | AC39 | HEK-293T | C73(0.88) | LDD1539 | [7] |
| LDCM0301 | AC4 | HEK-293T | C73(0.98) | LDD1540 | [7] |
| LDCM0302 | AC40 | HEK-293T | C74(1.17) | LDD1541 | [7] |
| LDCM0306 | AC44 | HEK-293T | C73(1.15) | LDD1545 | [7] |
| LDCM0308 | AC46 | HEK-293T | C74(1.05) | LDD1547 | [7] |
| LDCM0309 | AC47 | HEK-293T | C73(0.90) | LDD1548 | [7] |
| LDCM0310 | AC48 | HEK-293T | C74(1.05) | LDD1549 | [7] |
| LDCM0315 | AC52 | HEK-293T | C73(1.04) | LDD1554 | [7] |
| LDCM0317 | AC54 | HEK-293T | C74(1.06) | LDD1556 | [7] |
| LDCM0318 | AC55 | HEK-293T | C73(0.88) | LDD1557 | [7] |
| LDCM0319 | AC56 | HEK-293T | C74(1.05) | LDD1558 | [7] |
| LDCM0323 | AC6 | HEK-293T | C74(1.04) | LDD1562 | [7] |
| LDCM0324 | AC60 | HEK-293T | C73(1.06) | LDD1563 | [7] |
| LDCM0326 | AC62 | HEK-293T | C74(1.07) | LDD1565 | [7] |
| LDCM0327 | AC63 | HEK-293T | C73(1.25) | LDD1566 | [7] |
| LDCM0328 | AC64 | HEK-293T | C74(1.13) | LDD1567 | [7] |
| LDCM0334 | AC7 | HEK-293T | C73(0.94) | LDD1568 | [7] |
| LDCM0345 | AC8 | HEK-293T | C74(1.08) | LDD1569 | [7] |
| LDCM0275 | AKOS034007705 | HEK-293T | C74(1.06) | LDD1514 | [7] |
| LDCM0108 | Chloroacetamide | HeLa | N.A. | LDD0222 | [5] |
| LDCM0368 | CL10 | HEK-293T | C74(1.46) | LDD1572 | [7] |
| LDCM0372 | CL103 | HEK-293T | C74(1.19) | LDD1576 | [7] |
| LDCM0376 | CL107 | HEK-293T | C74(1.21) | LDD1580 | [7] |
| LDCM0379 | CL11 | HEK-293T | C73(0.91) | LDD1583 | [7] |
| LDCM0381 | CL111 | HEK-293T | C74(1.05) | LDD1585 | [7] |
| LDCM0385 | CL115 | HEK-293T | C74(1.12) | LDD1589 | [7] |
| LDCM0389 | CL119 | HEK-293T | C74(1.13) | LDD1593 | [7] |
| LDCM0390 | CL12 | HEK-293T | C74(1.16) | LDD1594 | [7] |
| LDCM0394 | CL123 | HEK-293T | C74(1.06) | LDD1598 | [7] |
| LDCM0398 | CL127 | HEK-293T | C74(1.04) | LDD1602 | [7] |
| LDCM0402 | CL15 | HEK-293T | C74(1.27) | LDD1606 | [7] |
| LDCM0408 | CL20 | HEK-293T | C73(1.53) | LDD1612 | [7] |
| LDCM0410 | CL22 | HEK-293T | C74(1.48) | LDD1614 | [7] |
| LDCM0411 | CL23 | HEK-293T | C73(0.99) | LDD1615 | [7] |
| LDCM0412 | CL24 | HEK-293T | C74(1.36) | LDD1616 | [7] |
| LDCM0415 | CL27 | HEK-293T | C74(1.21) | LDD1619 | [7] |
| LDCM0418 | CL3 | HEK-293T | C74(1.26) | LDD1622 | [7] |
| LDCM0421 | CL32 | HEK-293T | C73(1.12) | LDD1625 | [7] |
| LDCM0423 | CL34 | HEK-293T | C74(1.30) | LDD1627 | [7] |
| LDCM0424 | CL35 | HEK-293T | C73(0.84) | LDD1628 | [7] |
| LDCM0425 | CL36 | HEK-293T | C74(1.30) | LDD1629 | [7] |
| LDCM0428 | CL39 | HEK-293T | C74(1.08) | LDD1632 | [7] |
| LDCM0434 | CL44 | HEK-293T | C73(1.13) | LDD1638 | [7] |
| LDCM0436 | CL46 | HEK-293T | C74(1.17) | LDD1640 | [7] |
| LDCM0437 | CL47 | HEK-293T | C73(1.04) | LDD1641 | [7] |
| LDCM0438 | CL48 | HEK-293T | C74(1.21) | LDD1642 | [7] |
| LDCM0447 | CL56 | HEK-293T | C73(1.47) | LDD1650 | [7] |
| LDCM0449 | CL58 | HEK-293T | C74(1.27) | LDD1652 | [7] |
| LDCM0450 | CL59 | HEK-293T | C73(0.93) | LDD1653 | [7] |
| LDCM0452 | CL60 | HEK-293T | C74(1.32) | LDD1655 | [7] |
| LDCM0455 | CL63 | HEK-293T | C74(1.14) | LDD1658 | [7] |
| LDCM0460 | CL68 | HEK-293T | C73(1.17) | LDD1663 | [7] |
| LDCM0463 | CL70 | HEK-293T | C74(1.22) | LDD1666 | [7] |
| LDCM0464 | CL71 | HEK-293T | C73(1.25) | LDD1667 | [7] |
| LDCM0465 | CL72 | HEK-293T | C74(1.29) | LDD1668 | [7] |
| LDCM0473 | CL8 | HEK-293T | C73(1.13) | LDD1676 | [7] |
| LDCM0474 | CL80 | HEK-293T | C73(1.11) | LDD1677 | [7] |
| LDCM0476 | CL82 | HEK-293T | C74(1.22) | LDD1679 | [7] |
| LDCM0477 | CL83 | HEK-293T | C73(0.83) | LDD1680 | [7] |
| LDCM0478 | CL84 | HEK-293T | C74(1.23) | LDD1681 | [7] |
| LDCM0481 | CL87 | HEK-293T | C74(1.16) | LDD1684 | [7] |
| LDCM0487 | CL92 | HEK-293T | C73(1.07) | LDD1690 | [7] |
| LDCM0489 | CL94 | HEK-293T | C74(1.38) | LDD1692 | [7] |
| LDCM0490 | CL95 | HEK-293T | C73(1.04) | LDD1693 | [7] |
| LDCM0491 | CL96 | HEK-293T | C74(1.12) | LDD1694 | [7] |
| LDCM0494 | CL99 | HEK-293T | C74(1.17) | LDD1697 | [7] |
| LDCM0495 | E2913 | HEK-293T | C74(1.26) | LDD1698 | [7] |
| LDCM0468 | Fragment33 | HEK-293T | C74(1.12) | LDD1671 | [7] |
| LDCM0615 | Fragment63-R | Jurkat | _(9.54) | LDD1487 | [2] |
| LDCM0617 | Fragment63-S | Jurkat | _(16.42) | LDD1490 | [2] |
| LDCM0569 | Fragment7 | Jurkat | _(7.83) | LDD1485 | [2] |
| LDCM0022 | KB02 | HCT 116 | C70(4.70); C71(4.70) | LDD0080 | [4] |
| LDCM0023 | KB03 | HCT 116 | C70(4.80); C71(4.80) | LDD0081 | [4] |
| LDCM0024 | KB05 | HCT 116 | C70(3.30); C71(3.30) | LDD0082 | [4] |
| LDCM0154 | YY4 | T cell | 3.27 | LDD0400 | [3] |
The Interaction Atlas With This Target
References
