Details of the Target
General Information of Target
| Target ID | LDTP09225 | |||||
|---|---|---|---|---|---|---|
| Target Name | sn-1-specific diacylglycerol lipase ABHD11 (ABHD11) | |||||
| Gene Name | ABHD11 | |||||
| Gene ID | 83451 | |||||
| Synonyms |
WBSCR21; sn-1-specific diacylglycerol lipase ABHD11; EC 3.1.1.116; Alpha/beta hydrolase domain-containing protein 11; Abhydrolase domain-containing protein 11; Williams-Beuren syndrome chromosomal region 21 protein
|
|||||
| 3D Structure | ||||||
| Sequence |
MLRWTRAWRLPREGLGPHGPSFARVPVAPSSSSGGRGGAEPRPLPLSYRLLDGEAALPAV
VFLHGLFGSKTNFNSIAKILAQQTGRRVLTVDARNHGDSPHSPDMSYEIMSQDLQDLLPQ LGLVPCVVVGHSMGGKTAMLLALQRPELVERLIAVDISPVESTGVSHFATYVAAMRAINI ADELPRSRARKLADEQLSSVIQDMAVRQHLLTNLVEVDGRFVWRVNLDALTQHLDKILAF PQRQESYLGPTLFLLGGNSQFVHPSHHPEIMRLFPRAQMQTVPNAGHWIHADRPQDFIAA IRGFLV |
|||||
| Target Bioclass |
Enzyme
|
|||||
| Family |
AB hydrolase superfamily
|
|||||
| Function |
Catalyzes the hydrolysis of diacylglycerol in vitro and may function as a key regulator in lipid metabolism, namely by regulating the intracellular levels of diacylglycerol. 1,2-diacyl-sn-glycerols are the preferred substrate over 1,3-diacyl-sn-glycerols. The enzyme hydrolyzes stearate in preference to palmitate from the sn-1 position of 1,2-diacyl-sn-glycerols. Maintains the functional lipoylation of the 2-oxoglutarate dehydrogenase complex (OGDHc) through its interaction with the OGDHc by preventing the formation of lipoyl adducts. In addition, is also required for the expansion and differentiation of embryonic stem cells (ESCs).
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C-Sul Probe Info |
 |
3.04 | LDD0066 | [1] | |
|
1oxF11yne Probe Info |
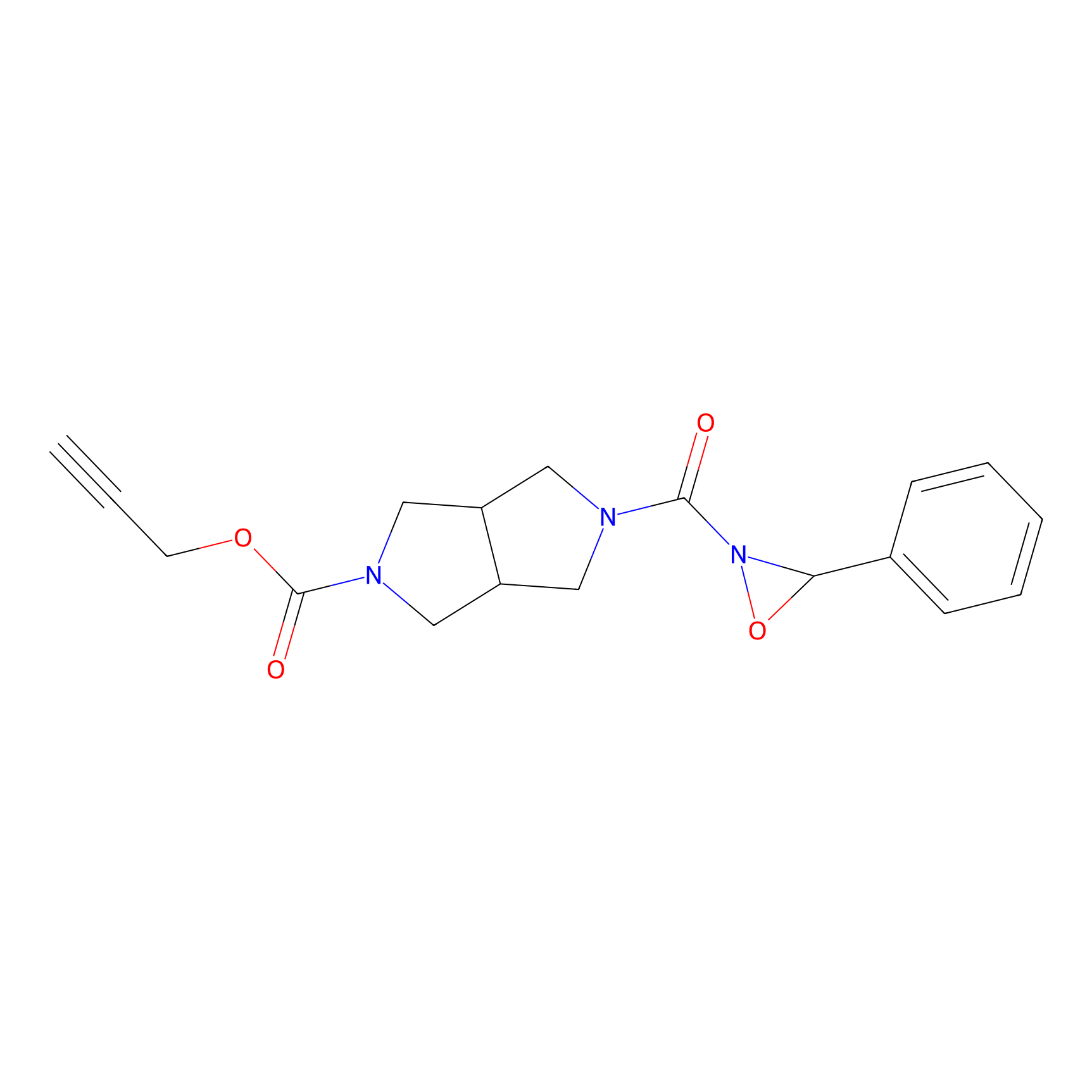 |
N.A. | LDD0193 | [2] | |
|
STPyne Probe Info |
 |
K87(1.85) | LDD0277 | [3] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0227 | [4] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD2241 | [5] | |
|
OSF Probe Info |
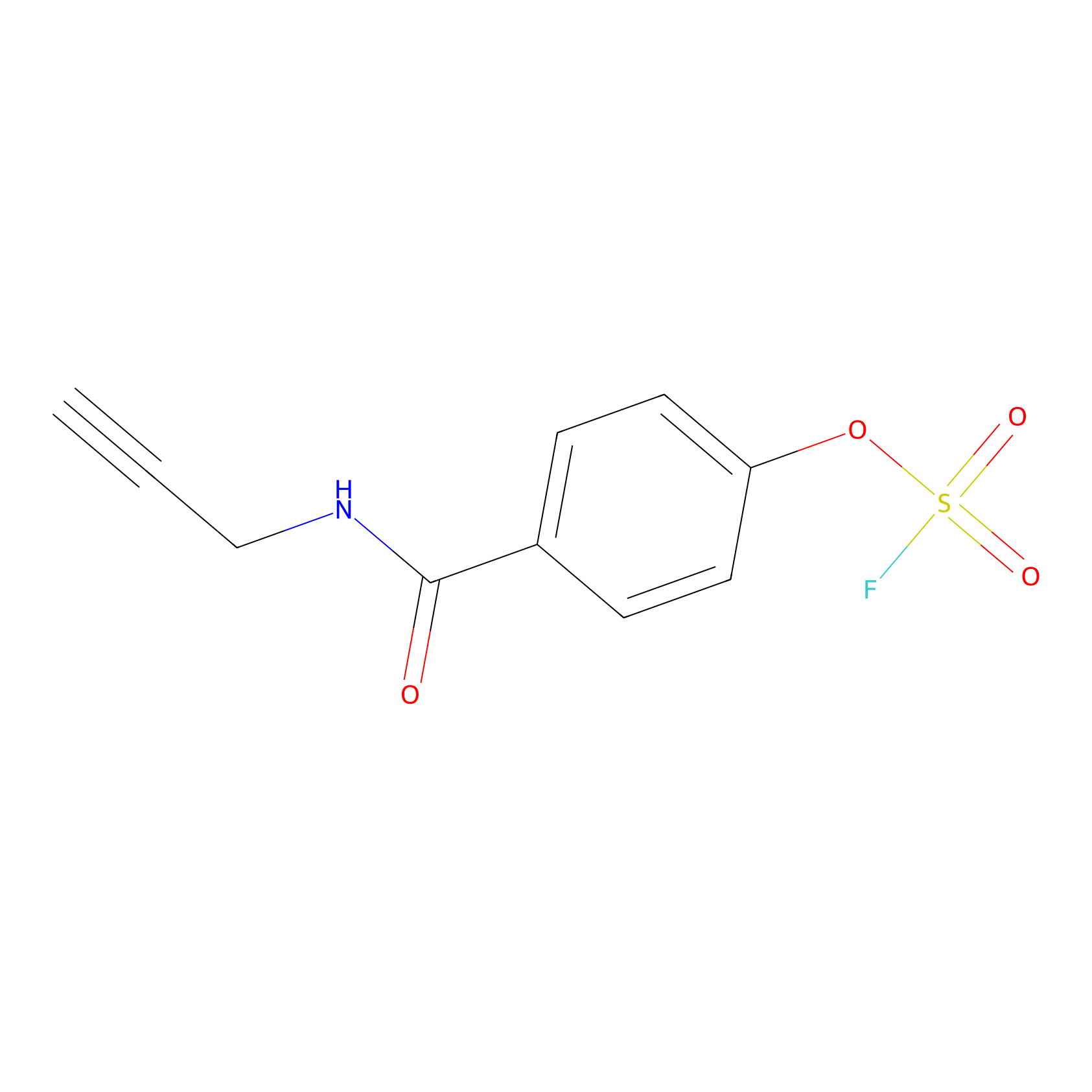 |
N.A. | LDD0029 | [6] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C158 Probe Info |
 |
26.72 | LDD1838 | [7] | |
|
C269 Probe Info |
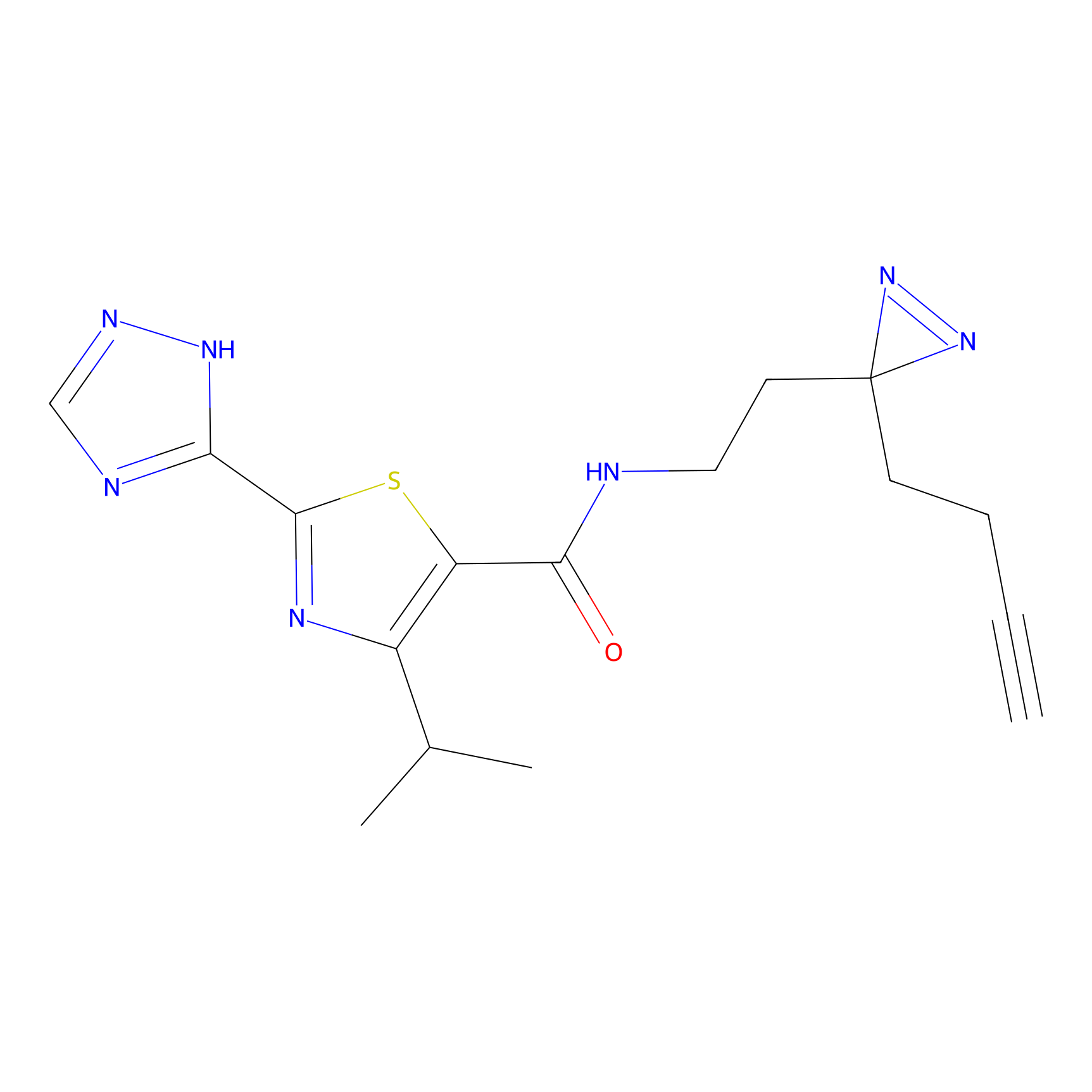 |
46.21 | LDD1939 | [7] | |
|
C363 Probe Info |
 |
19.70 | LDD2024 | [7] | |
|
C383 Probe Info |
 |
16.68 | LDD2042 | [7] | |
Competitor(s) Related to This Target
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transporter and channel
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| MyoD family inhibitor (MDFI) | MDFI family | Q99750 | |||
Transcription factor
Other
References
