Details of the Target
General Information of Target
| Target ID | LDTP08024 | |||||
|---|---|---|---|---|---|---|
| Target Name | Probable helicase senataxin (SETX) | |||||
| Gene Name | SETX | |||||
| Gene ID | 23064 | |||||
| Synonyms |
ALS4; KIAA0625; SCAR1; Probable helicase senataxin; EC 3.6.4.-; Amyotrophic lateral sclerosis 4 protein; SEN1 homolog; Senataxin |
|||||
| 3D Structure | ||||||
| Sequence |
MSTCCWCTPGGASTIDFLKRYASNTPSGEFQTADEDLCYCLECVAEYHKARDELPFLHEV
LWELETLRLINHFEKSMKAEIGDDDELYIVDNNGEMPLFDITGQDFENKLRVPLLEILKY PYLLLHERVNELCVEALCRMEQANCSFQVFDKHPGIYLFLVHPNEMVRRWAILTARNLGK VDRDDYYDLQEVLLCLFKVIELGLLESPDIYTSSVLEKGKLILLPSHMYDTTNYKSYWLG ICMLLTILEEQAMDSLLLGSDKQNDFMQSILHTMEREADDDSVDPFWPALHCFMVILDRL GSKVWGQLMDPIVAFQTIINNASYNREIRHIRNSSVRTKLEPESYLDDMVTCSQIVYNYN PEKTKKDSGWRTAICPDYCPNMYEEMETLASVLQSDIGQDMRVHNSTFLWFIPFVQSLMD LKDLGVAYIAQVVNHLYSEVKEVLNQTDAVCDKVTEFFLLILVSVIELHRNKKCLHLLWV SSQQWVEAVVKCAKLPTTAFTRSSEKSSGNCSKGTAMISSLSLHSMPSNSVQLAYVQLIR SLLKEGYQLGQQSLCKRFWDKLNLFLRGNLSLGWQLTSQETHELQSCLKQIIRNIKFKAP PCNTFVDLTSACKISPASYNKEESEQMGKTSRKDMHCLEASSPTFSKEPMKVQDSVLIKA DNTIEGDNNEQNYIKDVKLEDHLLAGSCLKQSSKNIFTERAEDQIKISTRKQKSVKEISS YTPKDCTSRNGPERGCDRGIIVSTRLLTDSSTDALEKVSTSNEDFSLKDDALAKTSKRKT KVQKDEICAKLSHVIKKQHRKSTLVDNTINLDENLTVSNIESFYSRKDTGVQKGDGFIHN LSLDPSGVLDDKNGEQKSQNNVLPKEKQLKNEELVIFSFHENNCKIQEFHVDGKELIPFT EMTNASEKKSSPFKDLMTVPESRDEEMSNSTSVIYSNLTREQAPDISPKSDTLTDSQIDR DLHKLSLLAQASVITFPSDSPQNSSQLQRKVKEDKRCFTANQNNVGDTSRGQVIIISDSD DDDDERILSLEKLTKQDKICLEREHPEQHVSTVNSKEEKNPVKEEKTETLFQFEESDSQC FEFESSSEVFSVWQDHPDDNNSVQDGEKKCLAPIANTTNGQGCTDYVSEVVKKGAEGIEE HTRPRSISVEEFCEIEVKKPKRKRSEKPMAEDPVRPSSSVRNEGQSDTNKRDLVGNDFKS IDRRTSTPNSRIQRATTVSQKKSSKLCTCTEPIRKVPVSKTPKKTHSDAKKGQNRSSNYL SCRTTPAIVPPKKFRQCPEPTSTAEKLGLKKGPRKAYELSQRSLDYVAQLRDHGKTVGVV DTRKKTKLISPQNLSVRNNKKLLTSQELQMQRQIRPKSQKNRRRLSDCESTDVKRAGSHT AQNSDIFVPESDRSDYNCTGGTEVLANSNRKQLIKCMPSEPETIKAKHGSPATDDACPLN QCDSVVLNGTVPTNEVIVSTSEDPLGGGDPTARHIEMAALKEGEPDSSSDAEEDNLFLTQ NDPEDMDLCSQMENDNYKLIELIHGKDTVEVEEDSVSRPQLESLSGTKCKYKDCLETTKN QGEYCPKHSEVKAADEDVFRKPGLPPPASKPLRPTTKIFSSKSTSRIAGLSKSLETSSAL SPSLKNKSKGIQSILKVPQPVPLIAQKPVGEMKNSCNVLHPQSPNNSNRQGCKVPFGESK YFPSSSPVNILLSSQSVSDTFVKEVLKWKYEMFLNFGQCGPPASLCQSISRPVPVRFHNY GDYFNVFFPLMVLNTFETVAQEWLNSPNRENFYQLQVRKFPADYIKYWEFAVYLEECELA KQLYPKENDLVFLAPERINEEKKDTERNDIQDLHEYHSGYVHKFRRTSVMRNGKTECYLS IQTQENFPANLNELVNCIVISSLVTTQRKLKAMSLLGSRNQLARAVLNPNPMDFCTKDLL TTTSERIIAYLRDFNEDQKKAIETAYAMVKHSPSVAKICLIHGPPGTGKSKTIVGLLYRL LTENQRKGHSDENSNAKIKQNRVLVCAPSNAAVDELMKKIILEFKEKCKDKKNPLGNCGD INLVRLGPEKSINSEVLKFSLDSQVNHRMKKELPSHVQAMHKRKEFLDYQLDELSRQRAL CRGGREIQRQELDENISKVSKERQELASKIKEVQGRPQKTQSIIILESHIICCTLSTSGG LLLESAFRGQGGVPFSCVIVDEAGQSCEIETLTPLIHRCNKLILVGDPKQLPPTVISMKA QEYGYDQSMMARFCRLLEENVEHNMISRLPILQLTVQYRMHPDICLFPSNYVYNRNLKTN RQTEAIRCSSDWPFQPYLVFDVGDGSERRDNDSYINVQEIKLVMEIIKLIKDKRKDVSFR NIGIITHYKAQKTMIQKDLDKEFDRKGPAEVDTVDAFQGRQKDCVIVTCVRANSIQGSIG FLASLQRLNVTITRAKYSLFILGHLRTLMENQHWNQLIQDAQKRGAIIKTCDKNYRHDAV KILKLKPVLQRSLTHPPTIAPEGSRPQGGLPSSKLDSGFAKTSVAASLYHTPSDSKEITL TVTSKDPERPPVHDQLQDPRLLKRMGIEVKGGIFLWDPQPSSPQHPGATPPTGEPGFPVV HQDLSHIQQPAAVVAALSSHKPPVRGEPPAASPEASTCQSKCDDPEEELCHRREARAFSE GEQEKCGSETHHTRRNSRWDKRTLEQEDSSSKKRKLL |
|||||
| Target Bioclass |
Enzyme
|
|||||
| Family |
DNA2/NAM7 helicase family
|
|||||
| Subcellular location |
Nucleus
|
|||||
| Function |
Probable RNA/DNA helicase involved in diverse aspects of RNA metabolism and genomic integrity. Plays a role in transcription regulation by its ability to modulate RNA Polymerase II (Pol II) binding to chromatin and through its interaction with proteins involved in transcription. Contributes to the mRNA splicing efficiency and splice site selection. Required for the resolution of R-loop RNA-DNA hybrid formation at G-rich pause sites located downstream of the poly(A) site, allowing XRN2 recruitment and XRN2-mediated degradation of the downstream cleaved RNA and hence efficient RNA polymerase II (RNAp II) transcription termination. Required for the 3' transcriptional termination of PER1 and CRY2, thus playing an important role in the circadian rhythm regulation. Involved in DNA double-strand breaks damage response generated by oxidative stress. In association with RRP45, targets the RNA exosome complex to sites of transcription-induced DNA damage. Plays a role in the development and maturation of germ cells: essential for male meiosis, acting at the interface of transcription and meiotic recombination, and in the process of gene silencing during meiotic sex chromosome inactivation (MSCI). May be involved in telomeric stability through the regulation of telomere repeat-containing RNA (TERRA) transcription. Plays a role in neurite outgrowth in hippocampal cells through FGF8-activated signaling pathways. Inhibits retinoic acid-induced apoptosis.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| AGS | SNV: p.S571F | DBIA Probe Info | |||
| CAKI1 | SNV: p.Y378D | . | |||
| CAL33 | SNV: p.W238Ter | . | |||
| CCK81 | Deletion: p.K473NfsTer8 | DBIA Probe Info | |||
| HEC1B | SNV: p.N2001S | . | |||
| HT115 | SNV: p.K75Q; p.D681N; p.V1388I; p.K2032N | . | |||
| IPC298 | Deletion: p.E2644del | DBIA Probe Info | |||
| JHH6 | SNV: p.S714C | DBIA Probe Info | |||
| JURKAT | Insertion: p.R989KfsTer7 | . | |||
| KNS42 | Insertion: p.I2020NfsTer4 | DBIA Probe Info | |||
| MCC26 | SNV: p.V465M | DBIA Probe Info | |||
| MDST8 | Deletion: p.I614SfsTer37 | DBIA Probe Info | |||
| NCIH1155 | SNV: p.S2398L | DBIA Probe Info | |||
| NUGC3 | Deletion: p.F1748SfsTer3 | DBIA Probe Info | |||
| OV56 | SNV: p.L2463V | DBIA Probe Info | |||
| RL952 | SNV: p.M140V | DBIA Probe Info | |||
| SNU1 | Deletion: p.S214del | DBIA Probe Info | |||
| TE4 | SNV: p.T3I | DBIA Probe Info | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
15.00 | LDD0402 | [1] | |
|
STPyne Probe Info |
 |
K2019(10.00) | LDD0277 | [2] | |
|
Probe 1 Probe Info |
 |
Y673(45.65) | LDD3495 | [3] | |
|
BTD Probe Info |
 |
C511(0.67) | LDD2099 | [4] | |
|
MCL-14 Probe Info |
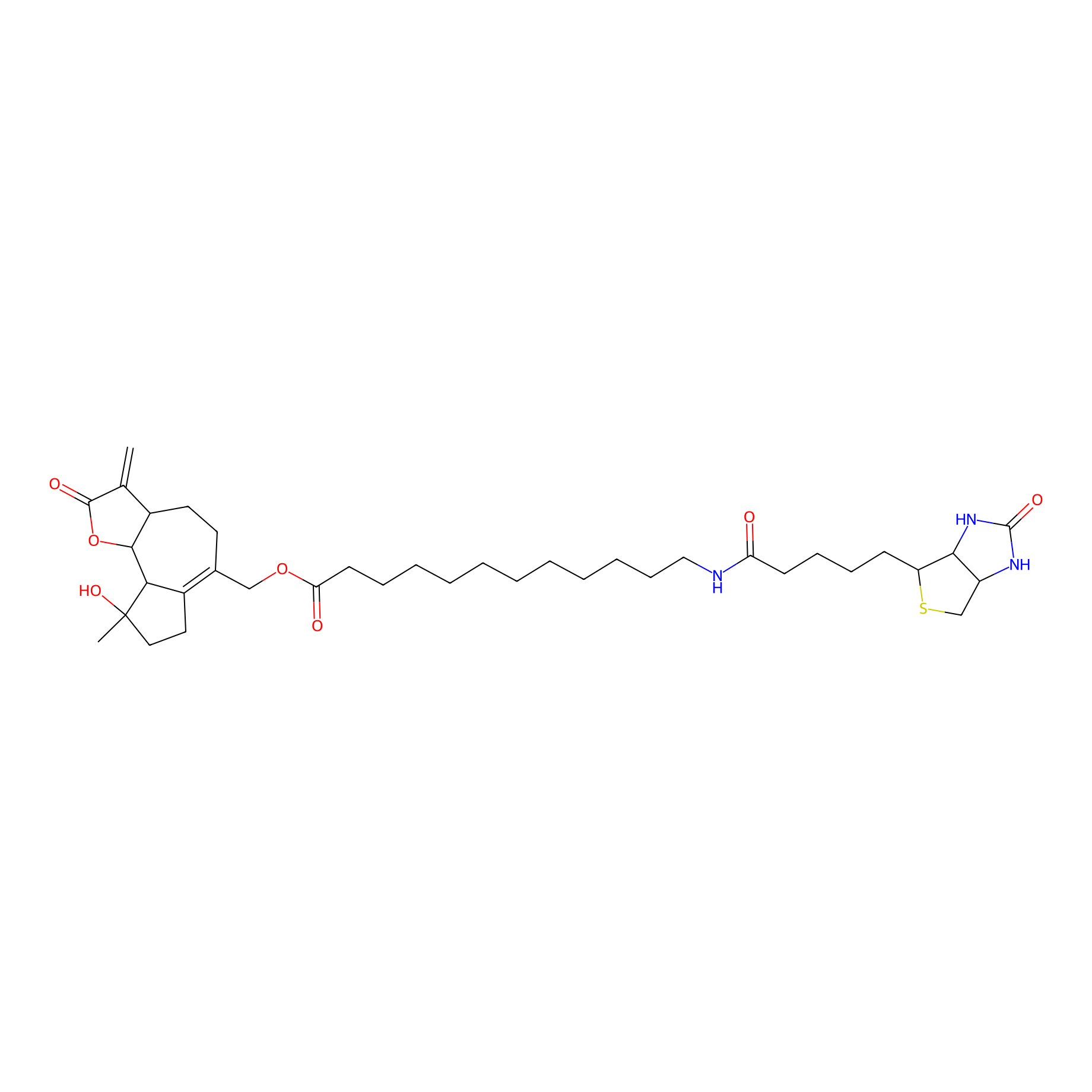 |
3.70 | LDD0050 | [5] | |
|
AHL-Pu-1 Probe Info |
 |
C1277(7.28) | LDD0168 | [6] | |
|
DBIA Probe Info |
 |
C555(12.92); C1262(5.53); C1959(32.95) | LDD0209 | [7] | |
|
CY-1 Probe Info |
 |
Q2134(0.00); Q2138(0.00) | LDD0246 | [8] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0162 | [9] | |
|
NAIA_4 Probe Info |
 |
C511(0.00); C587(0.00); C997(0.00); C1915(0.00) | LDD2226 | [10] | |
|
TFBX Probe Info |
 |
C1229(0.00); C1227(0.00); C997(0.00); C1262(0.00) | LDD0027 | [11] | |
|
WYneN Probe Info |
 |
N.A. | LDD0021 | [12] | |
|
IPM Probe Info |
 |
C555(0.00); C688(0.00) | LDD0005 | [12] | |
|
AOyne Probe Info |
 |
12.70 | LDD0443 | [13] | |
|
NAIA_5 Probe Info |
 |
C1123(0.00); C2646(0.00); C511(0.00); C2451(0.00) | LDD2223 | [10] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0524 | 2-Cyano-N-(2-morpholin-4-yl-ethyl)-acetamide | MDA-MB-231 | C511(1.09) | LDD2117 | [4] |
| LDCM0558 | 2-Cyano-N-phenylacetamide | MDA-MB-231 | C1959(0.71) | LDD2152 | [4] |
| LDCM0025 | 4SU-RNA | HEK-293T | C1277(7.28) | LDD0168 | [6] |
| LDCM0026 | 4SU-RNA+native RNA | HEK-293T | C352(3.30); C1277(7.70) | LDD0169 | [6] |
| LDCM0214 | AC1 | HEK-293T | C1368(1.09); C1915(1.12); C688(0.98); C195(1.07) | LDD1507 | [14] |
| LDCM0215 | AC10 | HEK-293T | C1277(1.00); C997(1.12); C1959(1.01); C1915(0.77) | LDD1508 | [14] |
| LDCM0226 | AC11 | HEK-293T | C1277(1.00); C1368(1.05); C1959(0.99); C1915(1.06) | LDD1509 | [14] |
| LDCM0237 | AC12 | HEK-293T | C1277(0.88); C997(1.05); C1368(1.04); C1959(1.02) | LDD1510 | [14] |
| LDCM0259 | AC14 | HEK-293T | C1277(1.19); C997(0.84); C1368(1.25); C1959(1.03) | LDD1512 | [14] |
| LDCM0270 | AC15 | HEK-293T | C1277(0.94); C997(1.21); C1368(0.91); C1959(1.10) | LDD1513 | [14] |
| LDCM0276 | AC17 | HEK-293T | C1368(1.00); C1915(1.12); C688(1.05); C195(0.91) | LDD1515 | [14] |
| LDCM0277 | AC18 | HEK-293T | C1277(0.92); C997(0.90); C1959(1.11); C1915(0.91) | LDD1516 | [14] |
| LDCM0278 | AC19 | HEK-293T | C1277(1.59); C1368(1.25); C1959(1.22); C1915(1.07) | LDD1517 | [14] |
| LDCM0279 | AC2 | HEK-293T | C1277(1.00); C997(1.27); C1959(1.09); C1915(0.81) | LDD1518 | [14] |
| LDCM0280 | AC20 | HEK-293T | C1277(0.84); C997(0.87); C1368(1.13); C1959(0.91) | LDD1519 | [14] |
| LDCM0281 | AC21 | HEK-293T | C1277(1.02); C1368(1.03); C1959(1.01); C1797(1.00) | LDD1520 | [14] |
| LDCM0282 | AC22 | HEK-293T | C1277(1.04); C997(1.16); C1368(1.08); C1959(1.01) | LDD1521 | [14] |
| LDCM0283 | AC23 | HEK-293T | C1277(1.05); C997(1.07); C1368(1.07); C1959(1.02) | LDD1522 | [14] |
| LDCM0284 | AC24 | HEK-293T | C1277(0.88); C1368(0.79); C1959(1.01); C2618(1.10) | LDD1523 | [14] |
| LDCM0285 | AC25 | HEK-293T | C1368(0.97); C1915(1.04); C688(1.09); C195(0.82) | LDD1524 | [14] |
| LDCM0286 | AC26 | HEK-293T | C1277(1.01); C997(1.34); C1959(1.09); C1915(0.87) | LDD1525 | [14] |
| LDCM0287 | AC27 | HEK-293T | C1277(1.03); C1368(0.99); C1959(0.98); C1915(1.11) | LDD1526 | [14] |
| LDCM0288 | AC28 | HEK-293T | C1277(0.90); C997(1.06); C1368(0.97); C1959(0.86) | LDD1527 | [14] |
| LDCM0289 | AC29 | HEK-293T | C1277(1.15); C1368(0.90); C1959(1.02); C1797(0.98) | LDD1528 | [14] |
| LDCM0290 | AC3 | HEK-293T | C1277(1.16); C1368(1.05); C1959(0.96); C1915(1.07) | LDD1529 | [14] |
| LDCM0291 | AC30 | HEK-293T | C1277(1.04); C997(0.96); C1368(1.40); C1959(0.99) | LDD1530 | [14] |
| LDCM0292 | AC31 | HEK-293T | C1277(1.02); C997(1.18); C1368(0.94); C1959(0.96) | LDD1531 | [14] |
| LDCM0293 | AC32 | HEK-293T | C1277(0.89); C1368(0.79); C1959(1.02); C2618(1.05) | LDD1532 | [14] |
| LDCM0294 | AC33 | HEK-293T | C1368(1.18); C1915(0.99); C688(1.02); C195(0.85) | LDD1533 | [14] |
| LDCM0295 | AC34 | HEK-293T | C1277(0.95); C997(1.32); C1959(0.91); C1915(0.82) | LDD1534 | [14] |
| LDCM0296 | AC35 | HEK-293T | C1277(1.01); C1368(1.07); C1959(0.94); C1915(1.27) | LDD1535 | [14] |
| LDCM0297 | AC36 | HEK-293T | C1277(1.00); C997(1.02); C1368(1.08); C1959(0.93) | LDD1536 | [14] |
| LDCM0298 | AC37 | HEK-293T | C1277(1.05); C1368(0.75); C1959(1.12); C1797(1.00) | LDD1537 | [14] |
| LDCM0299 | AC38 | HEK-293T | C1277(1.07); C997(0.91); C1368(1.20); C1959(1.05) | LDD1538 | [14] |
| LDCM0300 | AC39 | HEK-293T | C1277(0.96); C997(1.14); C1368(0.99); C1959(1.16) | LDD1539 | [14] |
| LDCM0301 | AC4 | HEK-293T | C1277(0.95); C997(1.12); C1368(1.20); C1959(0.95) | LDD1540 | [14] |
| LDCM0302 | AC40 | HEK-293T | C1277(0.89); C1368(1.03); C1959(1.19); C2618(1.34) | LDD1541 | [14] |
| LDCM0303 | AC41 | HEK-293T | C1368(0.94); C1915(1.17); C688(0.98); C195(1.24) | LDD1542 | [14] |
| LDCM0304 | AC42 | HEK-293T | C1277(1.03); C997(1.13); C1959(1.06); C1915(0.93) | LDD1543 | [14] |
| LDCM0305 | AC43 | HEK-293T | C1277(1.00); C1368(1.10); C1959(0.95); C1915(1.06) | LDD1544 | [14] |
| LDCM0306 | AC44 | HEK-293T | C1277(0.91); C997(1.04); C1368(1.16); C1959(0.90) | LDD1545 | [14] |
| LDCM0307 | AC45 | HEK-293T | C1277(1.05); C1368(0.93); C1959(1.11); C1797(1.11) | LDD1546 | [14] |
| LDCM0308 | AC46 | HEK-293T | C1277(1.06); C997(0.89); C1368(1.12); C1959(1.19) | LDD1547 | [14] |
| LDCM0309 | AC47 | HEK-293T | C1277(0.89); C997(0.98); C1368(0.93); C1959(1.11) | LDD1548 | [14] |
| LDCM0310 | AC48 | HEK-293T | C1277(0.90); C1368(0.80); C1959(0.99); C2618(1.03) | LDD1549 | [14] |
| LDCM0311 | AC49 | HEK-293T | C1368(0.92); C1915(0.98); C688(0.99); C195(0.98) | LDD1550 | [14] |
| LDCM0312 | AC5 | HEK-293T | C1277(0.95); C1368(1.03); C1959(1.05); C1797(1.08) | LDD1551 | [14] |
| LDCM0313 | AC50 | HEK-293T | C1277(0.98); C997(1.40); C1959(1.04); C1915(0.91) | LDD1552 | [14] |
| LDCM0314 | AC51 | HEK-293T | C1277(1.00); C1368(1.05); C1959(0.96); C1915(0.99) | LDD1553 | [14] |
| LDCM0315 | AC52 | HEK-293T | C1277(1.06); C997(0.96); C1368(1.08); C1959(1.01) | LDD1554 | [14] |
| LDCM0316 | AC53 | HEK-293T | C1277(1.00); C1368(0.96); C1959(1.19); C1797(1.15) | LDD1555 | [14] |
| LDCM0317 | AC54 | HEK-293T | C1277(0.98); C997(1.24); C1368(0.81); C1959(1.08) | LDD1556 | [14] |
| LDCM0318 | AC55 | HEK-293T | C1277(0.89); C997(0.96); C1368(0.98); C1959(1.04) | LDD1557 | [14] |
| LDCM0319 | AC56 | HEK-293T | C1277(0.96); C1368(1.02); C1959(1.00); C2618(1.15) | LDD1558 | [14] |
| LDCM0320 | AC57 | HEK-293T | C1368(0.99); C1915(1.27); C688(0.99); C195(0.97) | LDD1559 | [14] |
| LDCM0321 | AC58 | HEK-293T | C1277(1.00); C997(1.09); C1959(0.97); C1915(0.89) | LDD1560 | [14] |
| LDCM0322 | AC59 | HEK-293T | C1277(1.03); C1368(1.01); C1959(0.90); C1915(1.10) | LDD1561 | [14] |
| LDCM0323 | AC6 | HEK-293T | C1277(1.08); C997(0.93); C1368(1.12); C1959(1.07) | LDD1562 | [14] |
| LDCM0324 | AC60 | HEK-293T | C1277(0.89); C997(1.08); C1368(1.18); C1959(1.01) | LDD1563 | [14] |
| LDCM0325 | AC61 | HEK-293T | C1277(0.95); C1368(0.91); C1959(1.08); C1797(1.07) | LDD1564 | [14] |
| LDCM0326 | AC62 | HEK-293T | C1277(1.06); C997(0.97); C1368(0.90); C1959(1.03) | LDD1565 | [14] |
| LDCM0327 | AC63 | HEK-293T | C1277(0.98); C997(0.96); C1368(0.93); C1959(1.11) | LDD1566 | [14] |
| LDCM0328 | AC64 | HEK-293T | C1277(0.82); C1368(0.95); C1959(1.09); C2618(1.15) | LDD1567 | [14] |
| LDCM0334 | AC7 | HEK-293T | C1277(0.93); C997(0.81); C1368(0.97); C1959(1.04) | LDD1568 | [14] |
| LDCM0345 | AC8 | HEK-293T | C1277(0.99); C1368(0.67); C1959(0.97); C2618(1.09) | LDD1569 | [14] |
| LDCM0248 | AKOS034007472 | HEK-293T | C1277(1.06); C1368(1.13); C1959(1.12); C1797(1.14) | LDD1511 | [14] |
| LDCM0356 | AKOS034007680 | HEK-293T | C1368(1.06); C1915(1.07); C688(1.03); C195(1.13) | LDD1570 | [14] |
| LDCM0275 | AKOS034007705 | HEK-293T | C1277(0.95); C1368(0.86); C1959(1.00); C2618(1.24) | LDD1514 | [14] |
| LDCM0632 | CL-Sc | Hep-G2 | C788(1.06); C2646(0.62); C2451(0.62) | LDD2227 | [10] |
| LDCM0367 | CL1 | HEK-293T | C1277(0.92); C1368(0.98); C1959(0.98); C1797(1.10) | LDD1571 | [14] |
| LDCM0368 | CL10 | HEK-293T | C1277(1.11); C997(0.92); C1368(1.13); C1959(1.13) | LDD1572 | [14] |
| LDCM0369 | CL100 | HEK-293T | C1277(0.93); C1368(1.12); C1959(1.29); C1915(1.20) | LDD1573 | [14] |
| LDCM0370 | CL101 | HEK-293T | C1277(0.99); C1368(1.00); C1959(1.05); C1797(1.08) | LDD1574 | [14] |
| LDCM0371 | CL102 | HEK-293T | C1277(1.10); C1368(2.36); C1959(0.97); C1915(1.05) | LDD1575 | [14] |
| LDCM0372 | CL103 | HEK-293T | C997(0.96); C1368(0.92); C1959(0.94); C1915(1.08) | LDD1576 | [14] |
| LDCM0373 | CL104 | HEK-293T | C1277(0.98); C1368(1.15); C1959(1.17); C1915(1.05) | LDD1577 | [14] |
| LDCM0374 | CL105 | HEK-293T | C1277(0.91); C1368(0.97); C1959(0.99); C1797(0.96) | LDD1578 | [14] |
| LDCM0375 | CL106 | HEK-293T | C1277(0.95); C1368(1.32); C1959(1.06); C1915(0.97) | LDD1579 | [14] |
| LDCM0376 | CL107 | HEK-293T | C997(1.11); C1368(1.00); C1959(0.89); C1915(0.98) | LDD1580 | [14] |
| LDCM0377 | CL108 | HEK-293T | C1277(0.89); C1368(1.18); C1959(0.89); C1915(1.01) | LDD1581 | [14] |
| LDCM0378 | CL109 | HEK-293T | C1277(1.08); C1368(0.87); C1959(0.93); C1797(0.95) | LDD1582 | [14] |
| LDCM0379 | CL11 | HEK-293T | C1277(1.22); C997(1.02); C1368(1.20); C1959(1.31) | LDD1583 | [14] |
| LDCM0380 | CL110 | HEK-293T | C1277(0.91); C1368(0.90); C1959(1.13); C1915(0.89) | LDD1584 | [14] |
| LDCM0381 | CL111 | HEK-293T | C997(0.73); C1368(0.18); C1959(0.88); C1915(1.02) | LDD1585 | [14] |
| LDCM0382 | CL112 | HEK-293T | C1277(1.02); C1368(1.05); C1959(1.11); C1915(1.25) | LDD1586 | [14] |
| LDCM0383 | CL113 | HEK-293T | C1277(0.88); C1368(1.21); C1959(1.11); C1797(1.28) | LDD1587 | [14] |
| LDCM0384 | CL114 | HEK-293T | C1277(1.08); C1368(1.22); C1959(0.90); C1915(1.08) | LDD1588 | [14] |
| LDCM0385 | CL115 | HEK-293T | C997(1.03); C1368(1.05); C1959(0.91); C1915(1.02) | LDD1589 | [14] |
| LDCM0386 | CL116 | HEK-293T | C1277(0.98); C1368(1.13); C1959(1.15); C1915(0.98) | LDD1590 | [14] |
| LDCM0387 | CL117 | HEK-293T | C1277(0.95); C1368(0.96); C1959(1.10); C1797(1.17) | LDD1591 | [14] |
| LDCM0388 | CL118 | HEK-293T | C1277(0.98); C1368(1.27); C1959(0.95); C1915(1.01) | LDD1592 | [14] |
| LDCM0389 | CL119 | HEK-293T | C997(1.08); C1368(1.03); C1959(0.89); C1915(1.07) | LDD1593 | [14] |
| LDCM0390 | CL12 | HEK-293T | C1277(1.12); C1368(1.08); C1959(1.04); C2618(1.19) | LDD1594 | [14] |
| LDCM0391 | CL120 | HEK-293T | C1277(1.00); C1368(1.16); C1959(1.20); C1915(1.07) | LDD1595 | [14] |
| LDCM0392 | CL121 | HEK-293T | C1277(0.96); C1368(0.75); C1959(1.10); C1797(0.90) | LDD1596 | [14] |
| LDCM0393 | CL122 | HEK-293T | C1277(0.90); C1368(0.96); C1959(1.10); C1915(1.01) | LDD1597 | [14] |
| LDCM0394 | CL123 | HEK-293T | C997(0.84); C1368(1.00); C1959(0.95); C1915(1.17) | LDD1598 | [14] |
| LDCM0395 | CL124 | HEK-293T | C1277(0.95); C1368(1.04); C1959(1.14); C1915(1.18) | LDD1599 | [14] |
| LDCM0396 | CL125 | HEK-293T | C1277(0.91); C1368(0.79); C1959(0.97); C1797(1.07) | LDD1600 | [14] |
| LDCM0397 | CL126 | HEK-293T | C1277(0.93); C1368(0.99); C1959(1.07); C1915(1.06) | LDD1601 | [14] |
| LDCM0398 | CL127 | HEK-293T | C997(1.02); C1368(0.94); C1959(0.99); C1915(1.13) | LDD1602 | [14] |
| LDCM0399 | CL128 | HEK-293T | C1277(1.03); C1368(0.98); C1959(1.22); C1915(1.11) | LDD1603 | [14] |
| LDCM0400 | CL13 | HEK-293T | C1277(1.01); C1368(0.92); C1959(1.11); C1797(1.04) | LDD1604 | [14] |
| LDCM0401 | CL14 | HEK-293T | C1277(1.00); C1368(0.83); C1959(1.03); C1915(1.04) | LDD1605 | [14] |
| LDCM0402 | CL15 | HEK-293T | C997(0.86); C1368(1.01); C1959(1.07); C1915(1.18) | LDD1606 | [14] |
| LDCM0403 | CL16 | HEK-293T | C1277(1.06); C1368(1.05); C1959(0.99); C1915(1.20) | LDD1607 | [14] |
| LDCM0404 | CL17 | HEK-293T | C1368(0.91); C1915(1.14); C688(1.13); C195(0.77) | LDD1608 | [14] |
| LDCM0405 | CL18 | HEK-293T | C1277(1.17); C997(0.96); C1959(1.22); C1915(0.90) | LDD1609 | [14] |
| LDCM0406 | CL19 | HEK-293T | C1277(1.04); C1368(1.06); C1959(0.93); C1915(1.00) | LDD1610 | [14] |
| LDCM0407 | CL2 | HEK-293T | C1277(1.07); C1368(1.32); C1959(1.18); C1915(1.01) | LDD1611 | [14] |
| LDCM0408 | CL20 | HEK-293T | C1277(1.00); C997(0.95); C1368(1.17); C1959(1.08) | LDD1612 | [14] |
| LDCM0409 | CL21 | HEK-293T | C1277(0.90); C1368(1.05); C1959(1.27); C1797(1.03) | LDD1613 | [14] |
| LDCM0410 | CL22 | HEK-293T | C1277(1.13); C997(1.11); C1368(1.15); C1959(1.05) | LDD1614 | [14] |
| LDCM0411 | CL23 | HEK-293T | C1277(1.02); C997(1.11); C1368(1.17); C1959(1.22) | LDD1615 | [14] |
| LDCM0412 | CL24 | HEK-293T | C1277(1.31); C1368(1.13); C1959(1.03); C2618(1.31) | LDD1616 | [14] |
| LDCM0413 | CL25 | HEK-293T | C1277(0.93); C1368(1.00); C1959(0.87); C1797(1.00) | LDD1617 | [14] |
| LDCM0414 | CL26 | HEK-293T | C1277(0.77); C1368(1.08); C1959(0.95); C1915(1.00) | LDD1618 | [14] |
| LDCM0415 | CL27 | HEK-293T | C997(1.34); C1368(0.96); C1959(0.84); C1915(1.09) | LDD1619 | [14] |
| LDCM0416 | CL28 | HEK-293T | C1277(0.94); C1368(1.01); C1959(1.01); C1915(1.07) | LDD1620 | [14] |
| LDCM0417 | CL29 | HEK-293T | C1368(0.93); C1915(1.11); C688(1.04); C195(0.92) | LDD1621 | [14] |
| LDCM0418 | CL3 | HEK-293T | C997(1.03); C1368(1.05); C1959(0.98); C1915(1.22) | LDD1622 | [14] |
| LDCM0419 | CL30 | HEK-293T | C1277(1.09); C997(0.95); C1959(1.07); C1915(1.08) | LDD1623 | [14] |
| LDCM0420 | CL31 | HEK-293T | C1277(1.10); C1368(1.03); C1959(0.83); C1915(1.20) | LDD1624 | [14] |
| LDCM0421 | CL32 | HEK-293T | C1277(0.90); C997(1.16); C1368(1.23); C1959(0.97) | LDD1625 | [14] |
| LDCM0422 | CL33 | HEK-293T | C1277(0.96); C1368(1.18); C1959(1.08); C1797(0.81) | LDD1626 | [14] |
| LDCM0423 | CL34 | HEK-293T | C1277(1.04); C997(1.12); C1368(1.00); C1959(1.07) | LDD1627 | [14] |
| LDCM0424 | CL35 | HEK-293T | C1277(0.85); C997(1.04); C1368(1.33); C1959(1.06) | LDD1628 | [14] |
| LDCM0425 | CL36 | HEK-293T | C1277(1.18); C1368(0.94); C1959(1.08); C2618(1.14) | LDD1629 | [14] |
| LDCM0426 | CL37 | HEK-293T | C1277(0.95); C1368(1.05); C1959(1.12); C1797(1.03) | LDD1630 | [14] |
| LDCM0428 | CL39 | HEK-293T | C997(1.04); C1368(0.97); C1959(0.91); C1915(1.30) | LDD1632 | [14] |
| LDCM0429 | CL4 | HEK-293T | C1277(1.02); C1368(1.05); C1959(1.15); C1915(1.14) | LDD1633 | [14] |
| LDCM0430 | CL40 | HEK-293T | C1277(0.97); C1368(1.22); C1959(1.08); C1915(1.17) | LDD1634 | [14] |
| LDCM0431 | CL41 | HEK-293T | C1368(1.08); C1915(1.18); C688(1.00); C195(0.99) | LDD1635 | [14] |
| LDCM0432 | CL42 | HEK-293T | C1277(1.07); C997(1.16); C1959(1.15); C1915(0.79) | LDD1636 | [14] |
| LDCM0433 | CL43 | HEK-293T | C1277(0.95); C1368(1.07); C1959(1.00); C1915(1.15) | LDD1637 | [14] |
| LDCM0434 | CL44 | HEK-293T | C1277(0.92); C997(1.11); C1368(1.09); C1959(1.06) | LDD1638 | [14] |
| LDCM0435 | CL45 | HEK-293T | C1277(1.05); C1368(0.98); C1959(1.07); C1797(1.08) | LDD1639 | [14] |
| LDCM0436 | CL46 | HEK-293T | C1277(1.04); C997(1.01); C1368(1.12); C1959(0.97) | LDD1640 | [14] |
| LDCM0437 | CL47 | HEK-293T | C1277(1.05); C997(0.93); C1368(1.28); C1959(0.99) | LDD1641 | [14] |
| LDCM0438 | CL48 | HEK-293T | C1277(1.19); C1368(1.14); C1959(1.07); C2618(1.17) | LDD1642 | [14] |
| LDCM0439 | CL49 | HEK-293T | C1277(0.91); C1368(0.81); C1959(1.18); C1797(1.04) | LDD1643 | [14] |
| LDCM0440 | CL5 | HEK-293T | C1368(0.86); C1915(1.07); C688(1.08); C195(1.03) | LDD1644 | [14] |
| LDCM0441 | CL50 | HEK-293T | C1277(0.97); C1368(1.04); C1959(0.90); C1915(0.97) | LDD1645 | [14] |
| LDCM0443 | CL52 | HEK-293T | C1277(0.92); C1368(0.91); C1959(1.13); C1915(1.13) | LDD1646 | [14] |
| LDCM0444 | CL53 | HEK-293T | C1368(1.12); C1915(0.90); C688(1.00); C195(1.03) | LDD1647 | [14] |
| LDCM0445 | CL54 | HEK-293T | C1277(1.02); C997(0.92); C1959(1.13); C1915(0.74) | LDD1648 | [14] |
| LDCM0446 | CL55 | HEK-293T | C1277(1.29); C1368(1.23); C1959(0.95); C1915(1.21) | LDD1649 | [14] |
| LDCM0447 | CL56 | HEK-293T | C1277(0.84); C997(1.03); C1368(1.42); C1959(0.95) | LDD1650 | [14] |
| LDCM0448 | CL57 | HEK-293T | C1277(1.05); C1368(0.98); C1959(1.24); C1797(1.08) | LDD1651 | [14] |
| LDCM0449 | CL58 | HEK-293T | C1277(1.06); C997(1.10); C1368(1.06); C1959(1.15) | LDD1652 | [14] |
| LDCM0450 | CL59 | HEK-293T | C1277(0.83); C997(1.15); C1368(1.06); C1959(1.13) | LDD1653 | [14] |
| LDCM0451 | CL6 | HEK-293T | C1277(1.20); C997(1.43); C1959(1.28); C1915(0.99) | LDD1654 | [14] |
| LDCM0452 | CL60 | HEK-293T | C1277(1.10); C1368(1.09); C1959(1.09); C2618(1.22) | LDD1655 | [14] |
| LDCM0453 | CL61 | HEK-293T | C1277(0.92); C1368(1.11); C1959(0.95); C1797(0.89) | LDD1656 | [14] |
| LDCM0454 | CL62 | HEK-293T | C1277(0.95); C1368(0.97); C1959(0.97); C1915(1.04) | LDD1657 | [14] |
| LDCM0455 | CL63 | HEK-293T | C997(0.76); C1368(1.13); C1959(0.74); C1915(1.14) | LDD1658 | [14] |
| LDCM0456 | CL64 | HEK-293T | C1277(1.02); C1368(1.18); C1959(1.24); C1915(0.98) | LDD1659 | [14] |
| LDCM0457 | CL65 | HEK-293T | C1368(1.10); C1915(1.03); C688(1.07); C195(1.06) | LDD1660 | [14] |
| LDCM0458 | CL66 | HEK-293T | C1277(1.07); C997(1.19); C1959(0.94); C1915(0.77) | LDD1661 | [14] |
| LDCM0459 | CL67 | HEK-293T | C1277(1.15); C1368(1.12); C1959(0.82); C1915(1.07) | LDD1662 | [14] |
| LDCM0460 | CL68 | HEK-293T | C1277(0.86); C997(0.89); C1368(1.17); C1959(1.01) | LDD1663 | [14] |
| LDCM0461 | CL69 | HEK-293T | C1277(1.23); C1368(1.23); C1959(1.36); C1797(1.06) | LDD1664 | [14] |
| LDCM0462 | CL7 | HEK-293T | C1277(0.99); C1368(1.14); C1959(1.11); C1915(1.12) | LDD1665 | [14] |
| LDCM0463 | CL70 | HEK-293T | C1277(1.19); C997(0.99); C1368(1.03); C1959(1.05) | LDD1666 | [14] |
| LDCM0464 | CL71 | HEK-293T | C1277(0.92); C997(1.30); C1368(1.10); C1959(1.11) | LDD1667 | [14] |
| LDCM0465 | CL72 | HEK-293T | C1277(1.19); C1368(1.01); C1959(1.17); C2618(1.26) | LDD1668 | [14] |
| LDCM0466 | CL73 | HEK-293T | C1277(1.06); C1368(1.01); C1959(1.01); C1797(1.05) | LDD1669 | [14] |
| LDCM0467 | CL74 | HEK-293T | C1277(0.87); C1368(1.65); C1959(1.00); C1915(1.07) | LDD1670 | [14] |
| LDCM0469 | CL76 | HEK-293T | C1277(1.06); C1368(1.18); C1959(1.22); C1915(1.04) | LDD1672 | [14] |
| LDCM0470 | CL77 | HEK-293T | C1368(0.98); C1915(1.20); C688(1.03); C195(0.98) | LDD1673 | [14] |
| LDCM0471 | CL78 | HEK-293T | C1277(1.01); C997(1.81); C1959(1.08); C1915(0.85) | LDD1674 | [14] |
| LDCM0472 | CL79 | HEK-293T | C1277(1.09); C1368(1.03); C1959(1.02); C1915(1.01) | LDD1675 | [14] |
| LDCM0473 | CL8 | HEK-293T | C1277(0.97); C997(0.92); C1368(1.04); C1959(1.26) | LDD1676 | [14] |
| LDCM0474 | CL80 | HEK-293T | C1277(0.88); C997(0.94); C1368(1.16); C1959(1.01) | LDD1677 | [14] |
| LDCM0475 | CL81 | HEK-293T | C1277(1.16); C1368(1.09); C1959(1.17); C1797(0.83) | LDD1678 | [14] |
| LDCM0476 | CL82 | HEK-293T | C1277(1.09); C997(1.12); C1368(0.84); C1959(1.08) | LDD1679 | [14] |
| LDCM0477 | CL83 | HEK-293T | C1277(0.94); C997(0.98); C1368(1.15); C1959(1.11) | LDD1680 | [14] |
| LDCM0478 | CL84 | HEK-293T | C1277(1.16); C1368(1.07); C1959(0.97); C2618(1.11) | LDD1681 | [14] |
| LDCM0479 | CL85 | HEK-293T | C1277(0.99); C1368(0.92); C1959(0.96); C1797(1.25) | LDD1682 | [14] |
| LDCM0480 | CL86 | HEK-293T | C1277(0.91); C1368(0.99); C1959(1.00); C1915(0.95) | LDD1683 | [14] |
| LDCM0481 | CL87 | HEK-293T | C997(0.94); C1368(0.91); C1959(0.73); C1915(0.90) | LDD1684 | [14] |
| LDCM0482 | CL88 | HEK-293T | C1277(1.00); C1368(1.12); C1959(1.48); C1915(1.06) | LDD1685 | [14] |
| LDCM0483 | CL89 | HEK-293T | C1368(1.05); C1915(1.07); C688(0.95); C195(0.87) | LDD1686 | [14] |
| LDCM0484 | CL9 | HEK-293T | C1277(1.07); C1368(1.09); C1959(1.23); C1797(1.10) | LDD1687 | [14] |
| LDCM0485 | CL90 | HEK-293T | C1277(1.13); C997(1.41); C1959(1.15); C1915(0.97) | LDD1688 | [14] |
| LDCM0486 | CL91 | HEK-293T | C1277(0.95); C1368(0.97); C1959(1.03); C1915(1.12) | LDD1689 | [14] |
| LDCM0487 | CL92 | HEK-293T | C1277(1.05); C997(1.07); C1368(1.06); C1959(0.95) | LDD1690 | [14] |
| LDCM0488 | CL93 | HEK-293T | C1277(0.98); C1368(0.89); C1959(1.07); C1797(0.92) | LDD1691 | [14] |
| LDCM0489 | CL94 | HEK-293T | C1277(1.06); C997(0.77); C1368(1.01); C1959(1.06) | LDD1692 | [14] |
| LDCM0490 | CL95 | HEK-293T | C1277(0.91); C997(1.07); C1368(1.05); C1959(1.10) | LDD1693 | [14] |
| LDCM0491 | CL96 | HEK-293T | C1277(1.06); C1368(0.95); C1959(1.18); C2618(1.32) | LDD1694 | [14] |
| LDCM0492 | CL97 | HEK-293T | C1277(0.95); C1368(0.81); C1959(1.21); C1797(1.06) | LDD1695 | [14] |
| LDCM0493 | CL98 | HEK-293T | C1277(0.99); C1368(1.00); C1959(0.94); C1915(0.96) | LDD1696 | [14] |
| LDCM0494 | CL99 | HEK-293T | C997(0.85); C1368(0.99); C1959(0.85); C1915(1.13) | LDD1697 | [14] |
| LDCM0495 | E2913 | HEK-293T | C997(0.98); C1368(1.05); C1959(1.03); C1915(1.32) | LDD1698 | [14] |
| LDCM0213 | Electrophilic fragment 2 | MDA-MB-231 | C1229(2.41); C1262(0.94) | LDD1702 | [4] |
| LDCM0625 | F8 | Ramos | C511(0.69) | LDD2187 | [15] |
| LDCM0575 | Fragment13 | Ramos | C511(0.91) | LDD2192 | [15] |
| LDCM0580 | Fragment21 | Ramos | C511(0.97) | LDD2195 | [15] |
| LDCM0582 | Fragment23 | Ramos | C511(1.05) | LDD2196 | [15] |
| LDCM0578 | Fragment27 | Ramos | C511(0.83) | LDD2197 | [15] |
| LDCM0586 | Fragment28 | Ramos | C511(0.62) | LDD2198 | [15] |
| LDCM0588 | Fragment30 | Ramos | C511(1.34) | LDD2199 | [15] |
| LDCM0589 | Fragment31 | Ramos | C511(0.77) | LDD2200 | [15] |
| LDCM0468 | Fragment33 | HEK-293T | C997(0.90); C1368(0.99); C1959(0.87); C1915(1.11) | LDD1671 | [14] |
| LDCM0596 | Fragment38 | Ramos | C511(0.94) | LDD2203 | [15] |
| LDCM0427 | Fragment51 | HEK-293T | C1277(0.95); C1368(1.26); C1959(0.97); C1915(1.02) | LDD1631 | [14] |
| LDCM0610 | Fragment52 | Ramos | C511(1.17) | LDD2204 | [15] |
| LDCM0614 | Fragment56 | Ramos | C511(1.19) | LDD2205 | [15] |
| LDCM0022 | KB02 | HEK-293T | C1277(1.05); C1368(1.04); C555(0.85); C1959(0.99) | LDD1492 | [14] |
| LDCM0023 | KB03 | Jurkat | C555(12.92); C1262(5.53); C1959(32.95) | LDD0209 | [7] |
| LDCM0024 | KB05 | COLO792 | C1959(2.61) | LDD3310 | [16] |
| LDCM0006 | Micheliolide | M9-ENL1 | 3.70 | LDD0050 | [5] |
| LDCM0506 | Nucleophilic fragment 16a | MDA-MB-231 | C511(0.67) | LDD2099 | [4] |
| LDCM0532 | Nucleophilic fragment 29a | MDA-MB-231 | C511(0.66) | LDD2125 | [4] |
| LDCM0550 | Nucleophilic fragment 5a | MDA-MB-231 | C1959(1.64) | LDD2144 | [4] |
| LDCM0556 | Nucleophilic fragment 8a | MDA-MB-231 | C1959(0.35) | LDD2150 | [4] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Other
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Survival motor neuron protein (SMN1; SMN2) | SMN family | Q16637 | |||
| Superkiller complex protein 8 (SKIC8) | . | Q9GZS3 | |||
References
