Details of the Target
General Information of Target
| Target ID | LDTP07760 | |||||
|---|---|---|---|---|---|---|
| Target Name | CD109 antigen (CD109) | |||||
| Gene Name | CD109 | |||||
| Gene ID | 135228 | |||||
| Synonyms |
CPAMD7; CD109 antigen; 150 kDa TGF-beta-1-binding protein; C3 and PZP-like alpha-2-macroglobulin domain-containing protein 7; Platelet-specific Gov antigen; p180; r150; CD antigen CD109 |
|||||
| 3D Structure | ||||||
| Sequence |
MQGPPLLTAAHLLCVCTAALAVAPGPRFLVTAPGIIRPGGNVTIGVELLEHCPSQVTVKA
ELLKTASNLTVSVLEAEGVFEKGSFKTLTLPSLPLNSADEIYELRVTGRTQDEILFSNST RLSFETKRISVFIQTDKALYKPKQEVKFRIVTLFSDFKPYKTSLNILIKDPKSNLIQQWL SQQSDLGVISKTFQLSSHPILGDWSIQVQVNDQTYYQSFQVSEYVLPKFEVTLQTPLYCS MNSKHLNGTITAKYTYGKPVKGDVTLTFLPLSFWGKKKNITKTFKINGSANFSFNDEEMK NVMDSSNGLSEYLDLSSPGPVEILTTVTESVTGISRNVSTNVFFKQHDYIIEFFDYTTVL KPSLNFTATVKVTRADGNQLTLEERRNNVVITVTQRNYTEYWSGSNSGNQKMEAVQKINY TVPQSGTFKIEFPILEDSSELQLKAYFLGSKSSMAVHSLFKSPSKTYIQLKTRDENIKVG SPFELVVSGNKRLKELSYMVVSRGQLVAVGKQNSTMFSLTPENSWTPKACVIVYYIEDDG EIISDVLKIPVQLVFKNKIKLYWSKVKAEPSEKVSLRISVTQPDSIVGIVAVDKSVNLMN ASNDITMENVVHELELYNTGYYLGMFMNSFAVFQECGLWVLTDANLTKDYIDGVYDNAEY AERFMEENEGHIVDIHDFSLGSSPHVRKHFPETWIWLDTNMGYRIYQEFEVTVPDSITSW VATGFVISEDLGLGLTTTPVELQAFQPFFIFLNLPYSVIRGEEFALEITIFNYLKDATEV KVIIEKSDKFDILMTSNEINATGHQQTLLVPSEDGATVLFPIRPTHLGEIPITVTALSPT ASDAVTQMILVKAEGIEKSYSQSILLDLTDNRLQSTLKTLSFSFPPNTVTGSERVQITAI GDVLGPSINGLASLIRMPYGCGEQNMINFAPNIYILDYLTKKKQLTDNLKEKALSFMRQG YQRELLYQREDGSFSAFGNYDPSGSTWLSAFVLRCFLEADPYIDIDQNVLHRTYTWLKGH QKSNGEFWDPGRVIHSELQGGNKSPVTLTAYIVTSLLGYRKYQPNIDVQESIHFLESEFS RGISDNYTLALITYALSSVGSPKAKEALNMLTWRAEQEGGMQFWVSSESKLSDSWQPRSL DIEVAAYALLSHFLQFQTSEGIPIMRWLSRQRNSLGGFASTQDTTVALKALSEFAALMNT ERTNIQVTVTGPSSPSPVKFLIDTHNRLLLQTAELAVVQPTAVNISANGFGFAICQLNVV YNVKASGSSRRRRSIQNQEAFDLDVAVKENKDDLNHVDLNVCTSFSGPGRSGMALMEVNL LSGFMVPSEAISLSETVKKVEYDHGKLNLYLDSVNETQFCVNIPAVRNFKVSNTQDASVS IVDYYEPRRQAVRSYNSEVKLSSCDLCSDVQGCRPCEDGASGSHHHSSVIFIFCFKLLYF MELWL |
|||||
| Target Bioclass |
Other
|
|||||
| Family |
Protease inhibitor I39 (alpha-2-macroglobulin) family
|
|||||
| Subcellular location |
Cell membrane
|
|||||
| Function | Modulates negatively TGFB1 signaling in keratinocytes. | |||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| A3KAW | SNV: p.D1183Y | . | |||
| BT549 | SNV: p.S1332P | . | |||
| CALU1 | SNV: p.M625T | . | |||
| DU145 | SNV: p.G202C | . | |||
| EFO27 | SNV: p.Q1122Ter | . | |||
| G361 | Insertion: p.L752FfsTer11 | . | |||
| KMRC20 | SNV: p.P527L | . | |||
| KMS27 | SNV: p.A953T | . | |||
| KYSE410 | SNV: p.R1273T; p.V1285D | DBIA Probe Info | |||
| LNCaP clone FGC | SNV: p.V546A | . | |||
| OAW42 | Deletion: p.L441YfsTer3 | . | |||
| ONS76 | SNV: p.F1156L | DBIA Probe Info | |||
| SHP77 | SNV: p.G653V | . | |||
| TE4 | SNV: p.E1329K | DBIA Probe Info | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
FBPP2 Probe Info |
 |
2.39 | LDD0324 | [1] | |
|
OPA-S-S-alkyne Probe Info |
 |
K952(0.64); K1022(0.98); K1105(1.09); K86(1.68) | LDD3494 | [2] | |
|
DBIA Probe Info |
 |
C239(1.77) | LDD3312 | [3] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0225 | [4] | |
|
5E-2FA Probe Info |
 |
H1296(0.00); H1344(0.00) | LDD2235 | [5] | |
|
m-APA Probe Info |
 |
N.A. | LDD2231 | [5] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0165 | [6] | |
|
TFBX Probe Info |
 |
C995(0.00); C239(0.00) | LDD0148 | [7] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C149 Probe Info |
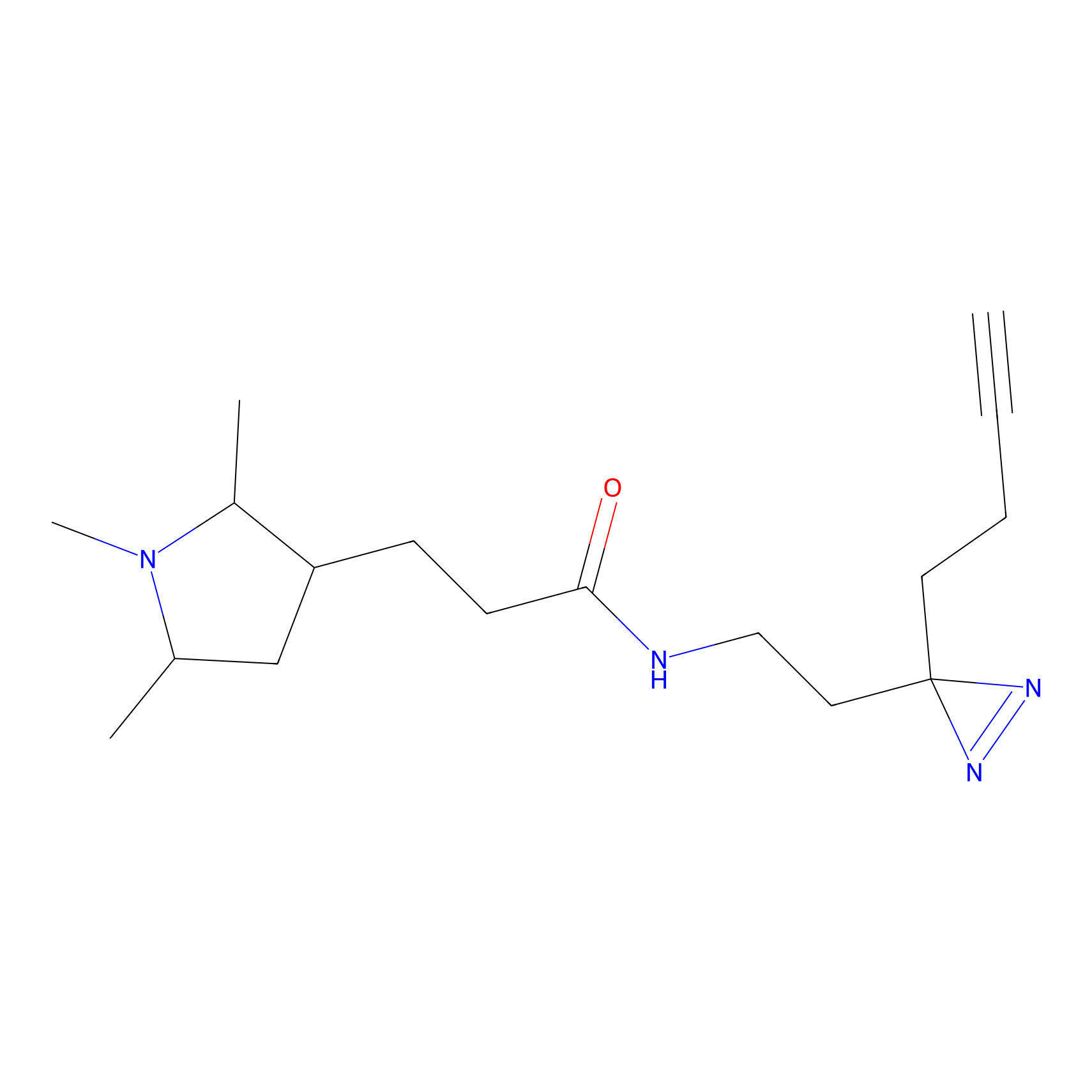 |
6.32 | LDD1830 | [8] | |
|
C158 Probe Info |
 |
18.00 | LDD1838 | [8] | |
|
C170 Probe Info |
 |
15.24 | LDD1850 | [8] | |
|
C171 Probe Info |
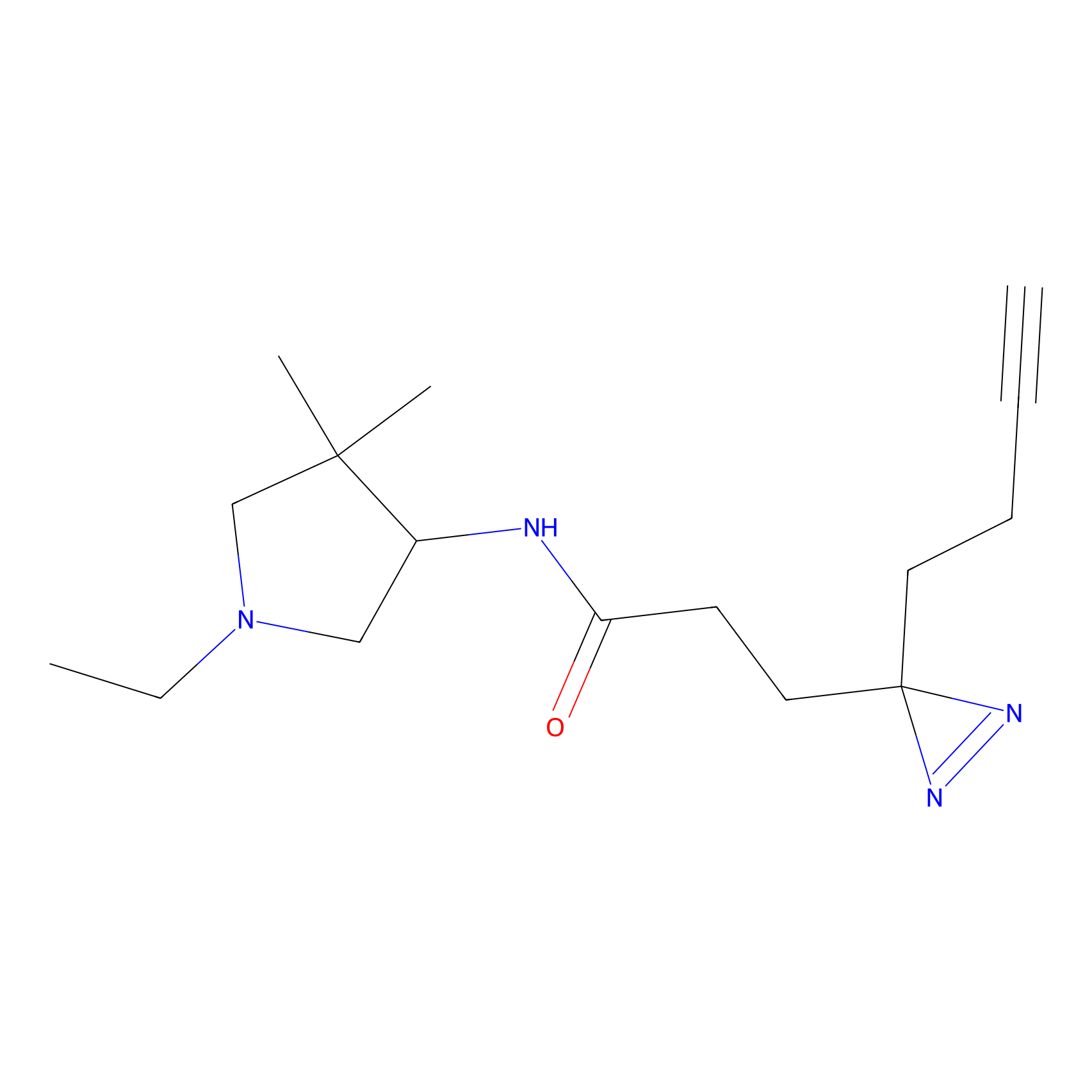 |
5.62 | LDD1851 | [8] | |
|
C223 Probe Info |
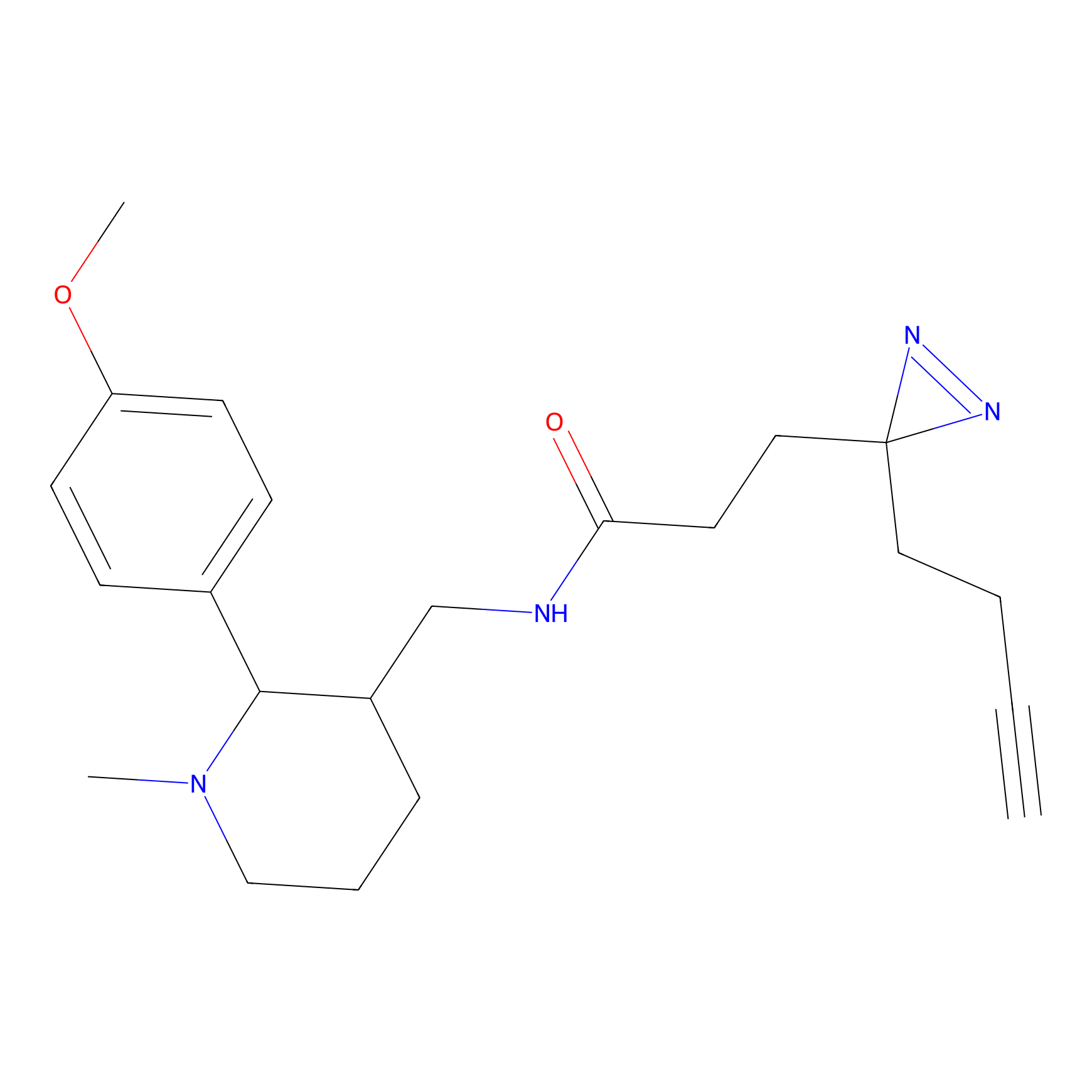 |
6.15 | LDD1897 | [8] | |
Competitor(s) Related to This Target
References
