Details of the Target
General Information of Target
| Target ID | LDTP05809 | |||||
|---|---|---|---|---|---|---|
| Target Name | Dehydrogenase/reductase SDR family member 2, mitochondrial (DHRS2) | |||||
| Gene Name | DHRS2 | |||||
| Gene ID | 10202 | |||||
| Synonyms |
SDR25C1; Dehydrogenase/reductase SDR family member 2, mitochondrial; EC 1.1.1.-; Dicarbonyl reductase HEP27; Protein D; Short chain dehydrogenase/reductase family 25C member 1; Protein SDR25C1 |
|||||
| 3D Structure | ||||||
| Sequence |
MLSAVARGYQGWFHPCARLSVRMSSTGIDRKGVLANRVAVVTGSTSGIGFAIARRLARDG
AHVVISSRKQQNVDRAMAKLQGEGLSVAGIVCHVGKAEDREQLVAKALEHCGGVDFLVCS AGVNPLVGSTLGTSEQIWDKILSVNVKSPALLLSQLLPYMENRRGAVILVSSIAAYNPVV ALGVYNVSKTALLGLTRTLALELAPKDIRVNCVVPGIIKTDFSKVFHGNESLWKNFKEHH QLQRIGESEDCAGIVSFLCSPDASYVNGENIAVAGYSTRL |
|||||
| Target Bioclass |
Enzyme
|
|||||
| Family |
Short-chain dehydrogenases/reductases (SDR) family
|
|||||
| Subcellular location |
Mitochondrion matrix
|
|||||
| Function |
NADPH-dependent oxidoreductase which catalyzes the reduction of dicarbonyl compounds. Displays reductase activity in vitro with 3,4-hexanedione, 2,3-heptanedione and 1-phenyl-1,2-propanedione as substrates. May function as a dicarbonyl reductase in the enzymatic inactivation of reactive carbonyls involved in covalent modification of cellular components. Also displays a minor hydroxysteroid dehydrogenase activity toward bile acids such as ursodeoxycholic acid (UDCA) and isoursodeoxycholic acid (isoUDCA), which makes it unlikely to control hormone levels. Doesn't show any activity in vitro with retinoids and sugars as substrates. Attenuates MDM2-mediated p53/TP53 degradation, leading to p53/TP53 stabilization and increased transcription activity, resulting in the accumulation of MDM2 and CDKN1A/p21. Reduces proliferation, migration and invasion of cancer cells and well as the production of ROS in cancer.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| ABC1 | SNV: p.G43R | DBIA Probe Info | |||
| COLO320 | SNV: p.V38L | DBIA Probe Info | |||
| COLO678 | SNV: p.V38L | DBIA Probe Info | |||
| COLO792 | SNV: p.G253R | DBIA Probe Info | |||
| COLO800 | SNV: p.V38L | . | |||
| HUH7 | SNV: p.S260Y | . | |||
| LS123 | SNV: p.V38L | DBIA Probe Info | |||
| MCC26 | Substitution: p.R279Q | . | |||
| NCIH716 | SNV: p.V38L | . | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
W1 Probe Info |
 |
22.31 | LDD0235 | [1] | |
|
YN-1 Probe Info |
 |
100.00 | LDD0444 | [2] | |
|
YN-4 Probe Info |
 |
100.00 | LDD0445 | [2] | |
|
IPM Probe Info |
 |
C92(0.00); C212(0.00); C251(0.00); C111(0.00) | LDD0241 | [1] | |
|
DBIA Probe Info |
 |
C92(1.19); C212(1.51) | LDD3310 | [3] | |
|
Curcusone 37 Probe Info |
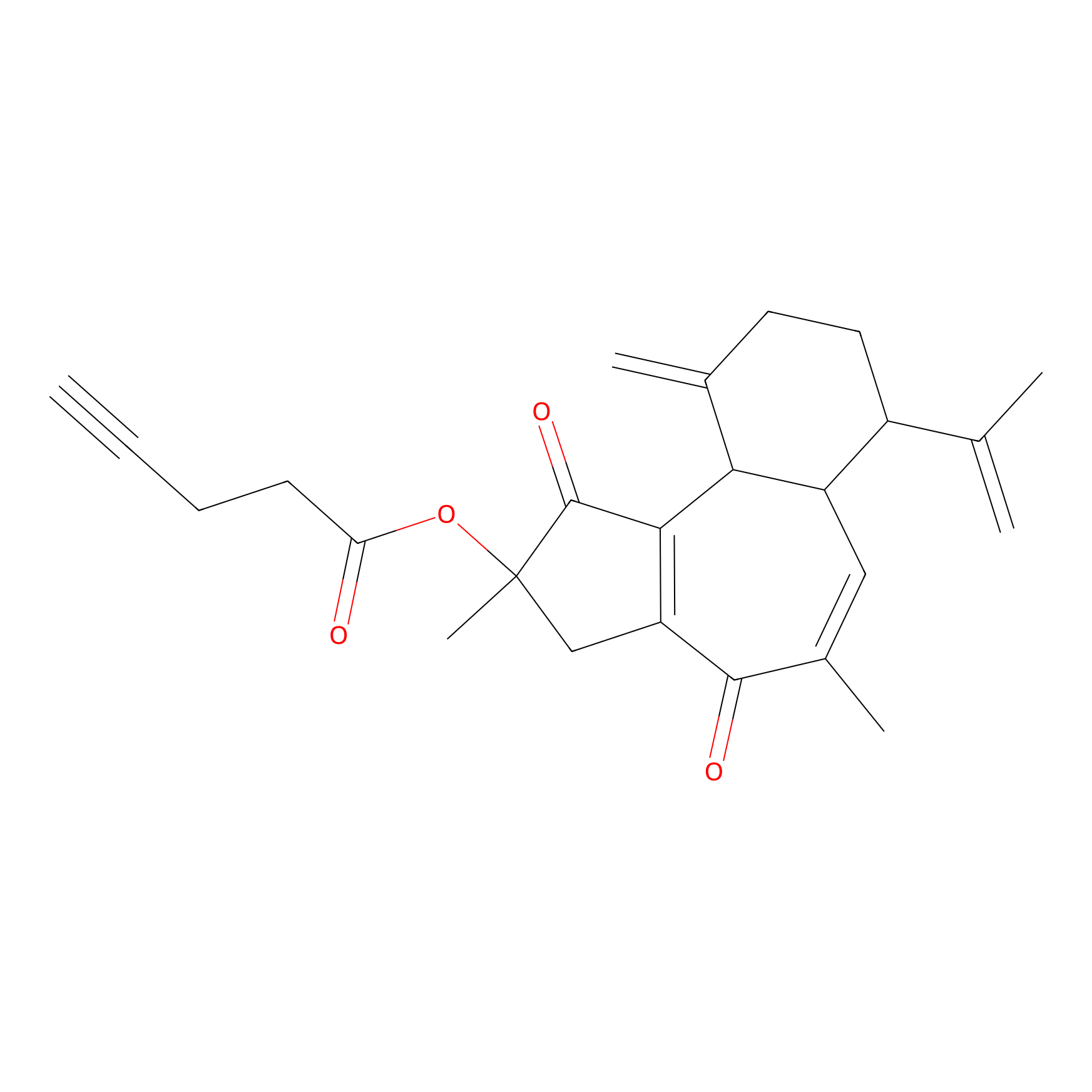 |
2.34 | LDD0188 | [4] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0237 | AC12 | HEK-293T | C212(0.92) | LDD1510 | [5] |
| LDCM0280 | AC20 | HEK-293T | C212(0.91) | LDD1519 | [5] |
| LDCM0288 | AC28 | HEK-293T | C212(1.01) | LDD1527 | [5] |
| LDCM0297 | AC36 | HEK-293T | C212(1.00) | LDD1536 | [5] |
| LDCM0301 | AC4 | HEK-293T | C212(0.81) | LDD1540 | [5] |
| LDCM0306 | AC44 | HEK-293T | C212(1.02) | LDD1545 | [5] |
| LDCM0315 | AC52 | HEK-293T | C212(1.19) | LDD1554 | [5] |
| LDCM0324 | AC60 | HEK-293T | C212(0.82) | LDD1563 | [5] |
| LDCM0408 | CL20 | HEK-293T | C212(1.49) | LDD1612 | [5] |
| LDCM0421 | CL32 | HEK-293T | C212(1.08) | LDD1625 | [5] |
| LDCM0434 | CL44 | HEK-293T | C212(1.31) | LDD1638 | [5] |
| LDCM0447 | CL56 | HEK-293T | C212(1.66) | LDD1650 | [5] |
| LDCM0460 | CL68 | HEK-293T | C212(1.33) | LDD1663 | [5] |
| LDCM0473 | CL8 | HEK-293T | C212(1.72) | LDD1676 | [5] |
| LDCM0474 | CL80 | HEK-293T | C212(1.23) | LDD1677 | [5] |
| LDCM0487 | CL92 | HEK-293T | C212(1.66) | LDD1690 | [5] |
| LDCM0033 | Curcusone1d | MCF-7 | 2.34 | LDD0188 | [4] |
| LDCM0022 | KB02 | 22RV1 | C92(1.55) | LDD2243 | [3] |
| LDCM0023 | KB03 | 22RV1 | C92(1.92) | LDD2660 | [3] |
| LDCM0024 | KB05 | COLO792 | C92(1.19); C212(1.51) | LDD3310 | [3] |
| LDCM0110 | W12 | Hep-G2 | E238(0.67); H227(1.79); E230(2.05); N229(2.05) | LDD0237 | [1] |
| LDCM0111 | W14 | Hep-G2 | K31(0.61); H227(1.52) | LDD0238 | [1] |
| LDCM0112 | W16 | Hep-G2 | R209(0.70); K206(0.80); D207(0.80); H227(0.80) | LDD0239 | [1] |
| LDCM0113 | W17 | Hep-G2 | K31(0.51); H239(13.46); H240(13.84) | LDD0240 | [1] |
The Interaction Atlas With This Target
References
