Details of the Target
General Information of Target
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
6.40 | LDD0402 | [1] | |
|
Nap-2 Probe Info |
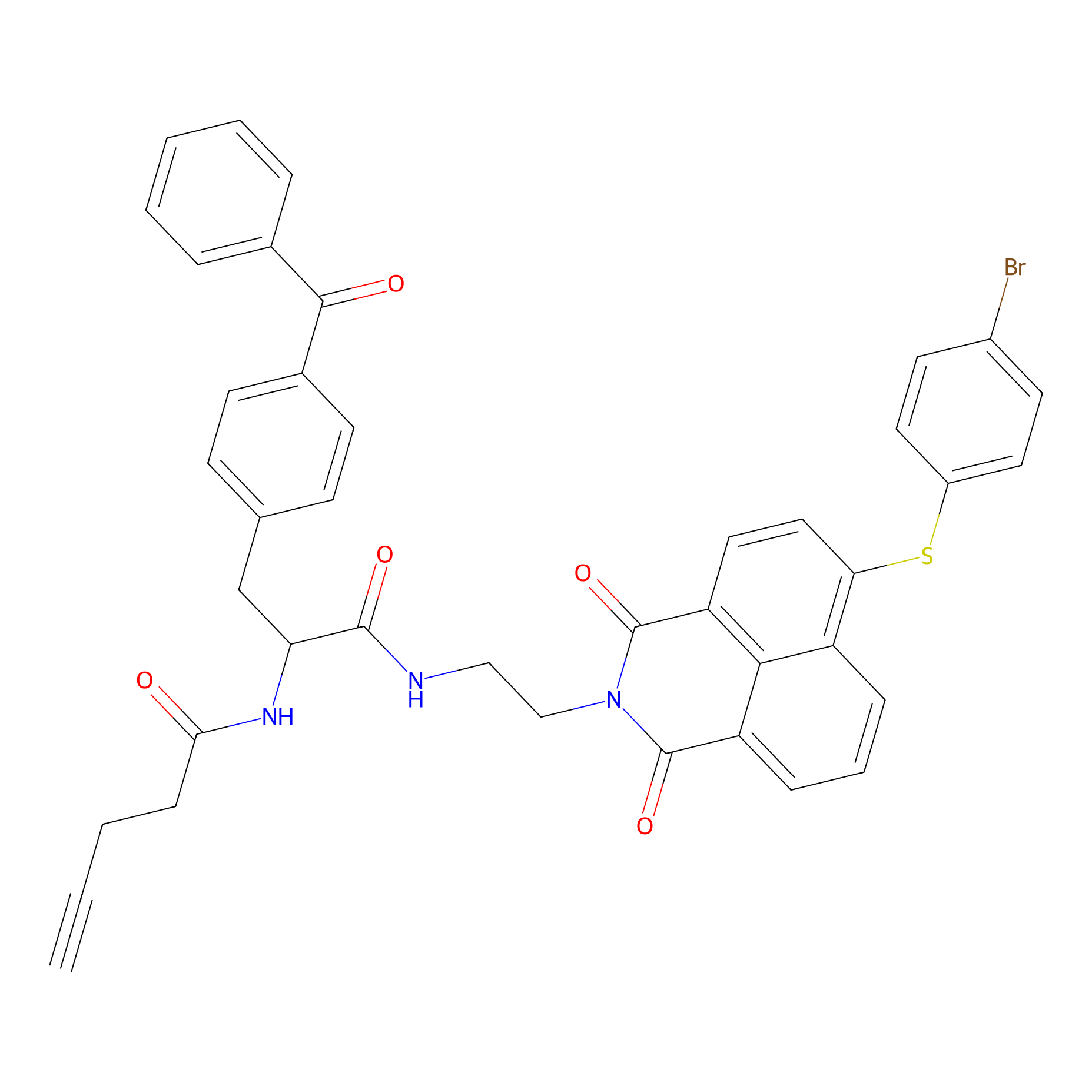 |
N.A. | LDD0435 | [2] | |
|
STPyne Probe Info |
 |
K70(9.96) | LDD2217 | [3] | |
|
Probe 1 Probe Info |
 |
Y71(27.63) | LDD3495 | [4] | |
|
DBIA Probe Info |
 |
C219(2.42) | LDD3314 | [5] | |
|
THZ1-DTB Probe Info |
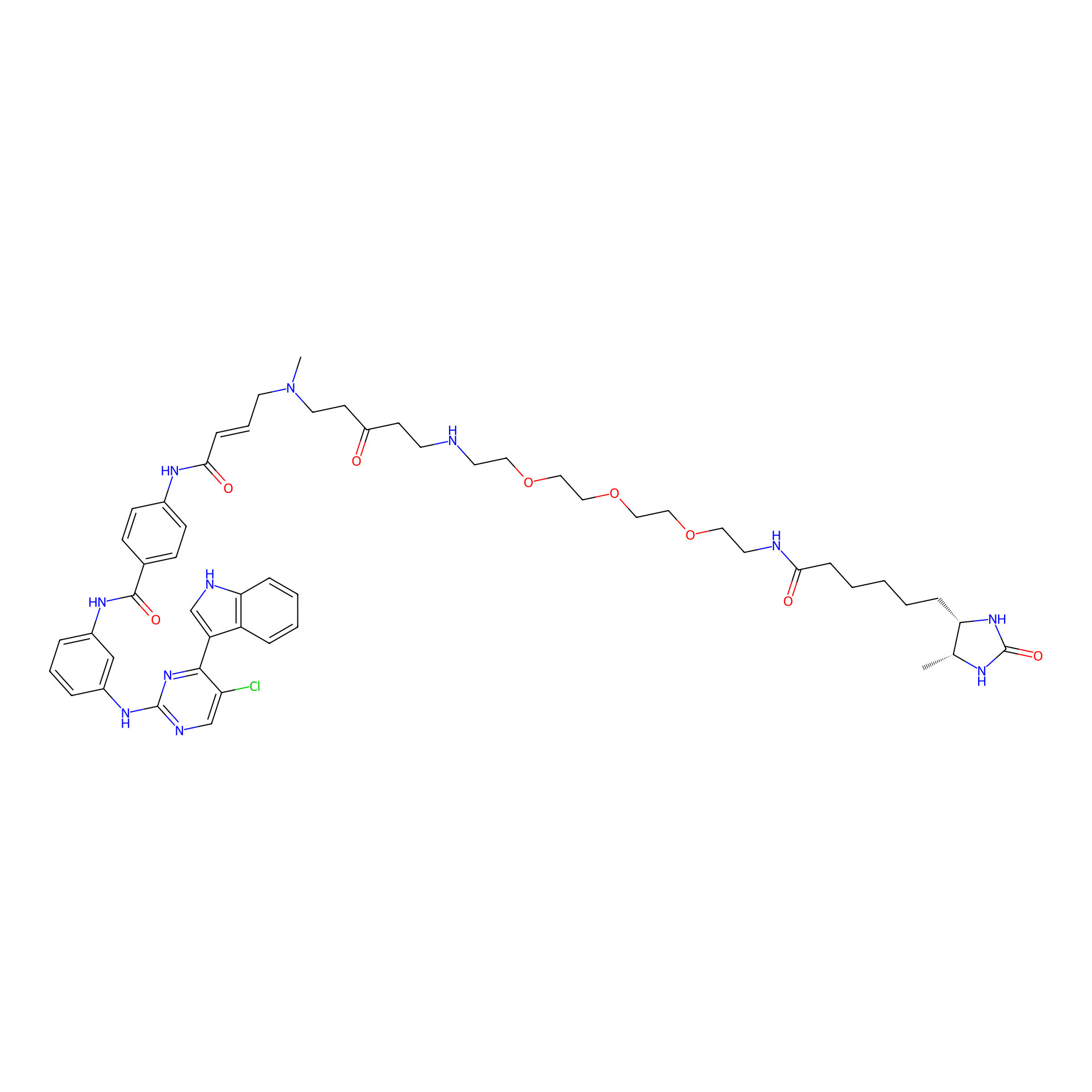 |
C219(1.08) | LDD0460 | [6] | |
|
HHS-475 Probe Info |
 |
Y242(0.75); Y55(0.77); Y71(0.92); Y339(1.09) | LDD0264 | [7] | |
|
IA-alkyne Probe Info |
 |
C287(0.37) | LDD2182 | [8] | |
|
HHS-465 Probe Info |
 |
Y190(10.00); Y242(10.00); Y55(10.00); Y71(9.20) | LDD2237 | [9] | |
|
5E-2FA Probe Info |
 |
H89(0.00); H90(0.00); H373(0.00); H103(0.00) | LDD2235 | [10] | |
|
ATP probe Probe Info |
 |
K52(0.00); K63(0.00); K330(0.00) | LDD0035 | [11] | |
|
AZ-9 Probe Info |
 |
D27(0.00); E243(0.00) | LDD0395 | [12] | |
|
NAIA_4 Probe Info |
 |
C219(0.00); C259(0.00) | LDD2226 | [13] | |
|
1c-yne Probe Info |
 |
K293(0.00); K70(0.00); C219(0.00) | LDD0228 | [14] | |
|
Acrolein Probe Info |
 |
C12(0.00); C19(0.00); K20(0.00) | LDD0217 | [15] | |
|
Methacrolein Probe Info |
 |
C19(0.00); K20(0.00) | LDD0218 | [15] | |
|
AOyne Probe Info |
 |
8.30 | LDD0443 | [16] | |
|
NAIA_5 Probe Info |
 |
C287(0.00); C259(0.00); C219(0.00) | LDD2223 | [13] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
FFF probe11 Probe Info |
 |
5.06 | LDD0472 | [17] | |
|
FFF probe13 Probe Info |
 |
5.85 | LDD0475 | [17] | |
|
FFF probe14 Probe Info |
 |
7.95 | LDD0477 | [17] | |
|
JN0003 Probe Info |
 |
7.95 | LDD0469 | [17] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0156 | Aniline | NCI-H1299 | 8.42 | LDD0403 | [1] |
| LDCM0625 | F8 | Ramos | C287(0.44) | LDD2187 | [8] |
| LDCM0572 | Fragment10 | Ramos | C287(0.49) | LDD2189 | [8] |
| LDCM0573 | Fragment11 | Ramos | C287(0.01) | LDD2190 | [8] |
| LDCM0574 | Fragment12 | Ramos | C287(0.43) | LDD2191 | [8] |
| LDCM0575 | Fragment13 | Ramos | C287(0.12) | LDD2192 | [8] |
| LDCM0576 | Fragment14 | Ramos | C287(0.74) | LDD2193 | [8] |
| LDCM0579 | Fragment20 | Ramos | C287(0.89) | LDD2194 | [8] |
| LDCM0580 | Fragment21 | Ramos | C287(0.51) | LDD2195 | [8] |
| LDCM0582 | Fragment23 | Ramos | C287(0.50) | LDD2196 | [8] |
| LDCM0578 | Fragment27 | Ramos | C287(0.54) | LDD2197 | [8] |
| LDCM0586 | Fragment28 | Ramos | C287(0.14) | LDD2198 | [8] |
| LDCM0588 | Fragment30 | Ramos | C287(0.17) | LDD2199 | [8] |
| LDCM0589 | Fragment31 | Ramos | C287(0.68) | LDD2200 | [8] |
| LDCM0590 | Fragment32 | Ramos | C287(0.75) | LDD2201 | [8] |
| LDCM0468 | Fragment33 | Ramos | C287(0.23) | LDD2202 | [8] |
| LDCM0596 | Fragment38 | Ramos | C287(0.18) | LDD2203 | [8] |
| LDCM0566 | Fragment4 | Ramos | C287(0.64) | LDD2184 | [8] |
| LDCM0610 | Fragment52 | Ramos | C287(0.09) | LDD2204 | [8] |
| LDCM0614 | Fragment56 | Ramos | C287(0.35) | LDD2205 | [8] |
| LDCM0569 | Fragment7 | Ramos | C287(0.61) | LDD2186 | [8] |
| LDCM0571 | Fragment9 | Ramos | C287(0.50) | LDD2188 | [8] |
| LDCM0116 | HHS-0101 | DM93 | Y242(0.75); Y55(0.77); Y71(0.92); Y339(1.09) | LDD0264 | [7] |
| LDCM0117 | HHS-0201 | DM93 | Y339(0.60); Y242(0.62); Y55(0.67); Y71(0.87) | LDD0265 | [7] |
| LDCM0118 | HHS-0301 | DM93 | Y55(0.71); Y71(0.85); Y190(1.11); Y242(1.16) | LDD0266 | [7] |
| LDCM0119 | HHS-0401 | DM93 | Y339(0.63); Y242(0.73); Y55(0.75); Y71(0.97) | LDD0267 | [7] |
| LDCM0120 | HHS-0701 | DM93 | Y242(0.52); Y55(0.71); Y71(0.72); Y190(1.00) | LDD0268 | [7] |
| LDCM0022 | KB02 | Ramos | C287(0.37) | LDD2182 | [8] |
| LDCM0023 | KB03 | Ramos | C287(0.35) | LDD2183 | [8] |
| LDCM0024 | KB05 | IGR37 | C219(2.42) | LDD3314 | [5] |
| LDCM0109 | NEM | HeLa | N.A. | LDD0223 | [15] |
| LDCM0021 | THZ1 | HeLa S3 | C219(1.08) | LDD0460 | [6] |
The Interaction Atlas With This Target
References
