Details of the Target
General Information of Target
| Target ID | LDTP04980 | |||||
|---|---|---|---|---|---|---|
| Target Name | U6 snRNA-associated Sm-like protein LSm6 (LSM6) | |||||
| Gene Name | LSM6 | |||||
| Gene ID | 11157 | |||||
| Synonyms |
U6 snRNA-associated Sm-like protein LSm6 |
|||||
| 3D Structure | ||||||
| Sequence |
MSLRKQTPSDFLKQIIGRPVVVKLNSGVDYRGVLACLDGYMNIALEQTEEYVNGQLKNKY
GDAFIRGNNVLYISTQKRRM |
|||||
| Target Bioclass |
Other
|
|||||
| Family |
SnRNP Sm proteins family, SmF/LSm6 subfamily
|
|||||
| Subcellular location |
Cytoplasm
|
|||||
| Function |
Plays a role in pre-mRNA splicing as component of the U4/U6-U5 tri-snRNP complex that is involved in spliceosome assembly, and as component of the precatalytic spliceosome (spliceosome B complex). The heptameric LSM2-8 complex binds specifically to the 3'-terminal U-tract of U6 snRNA. Component of LSm protein complexes, which are involved in RNA processing and may function in a chaperone-like manner, facilitating the efficient association of RNA processing factors with their substrates. Component of the cytoplasmic LSM1-LSM7 complex, which is thought to be involved in mRNA degradation by activating the decapping step in the 5'-to-3' mRNA decay pathway (Probable).
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
12.37 | LDD0402 | [1] | |
|
C-Sul Probe Info |
 |
2.99 | LDD0066 | [2] | |
|
TH211 Probe Info |
 |
Y40(20.00) | LDD0257 | [3] | |
|
STPyne Probe Info |
 |
K5(6.25) | LDD0277 | [4] | |
|
DBIA Probe Info |
 |
C36(0.94) | LDD1492 | [5] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
N.A. | LDD0038 | [6] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0036 | [6] | |
|
NAIA_5 Probe Info |
 |
N.A. | LDD2223 | [7] | |
|
HHS-482 Probe Info |
 |
Y40(1.14) | LDD2239 | [8] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C134 Probe Info |
 |
20.68 | LDD1816 | [9] | |
|
C305 Probe Info |
 |
9.71 | LDD1974 | [9] | |
|
C423 Probe Info |
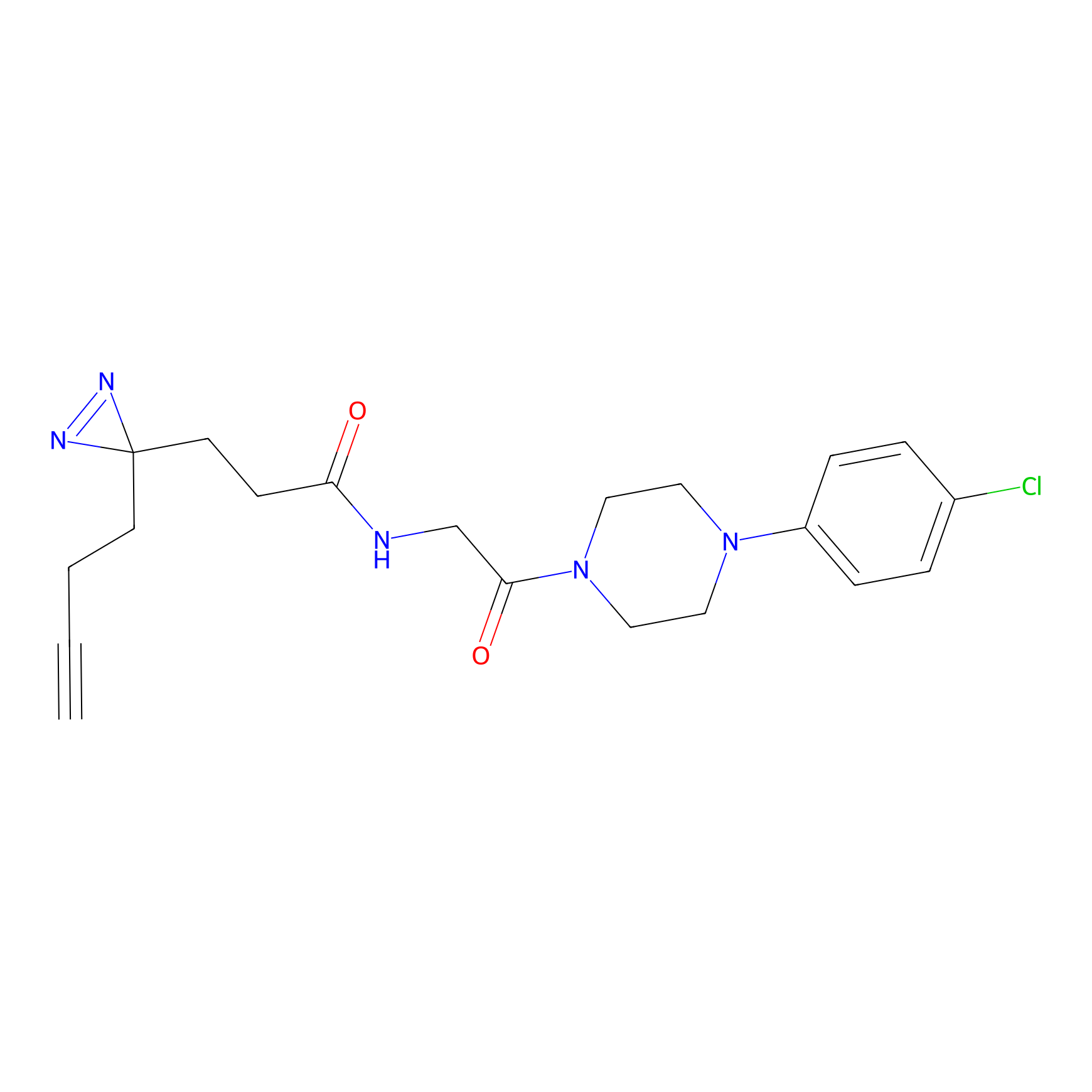 |
10.41 | LDD2078 | [9] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0214 | AC1 | HEK-293T | C36(1.07) | LDD1507 | [5] |
| LDCM0237 | AC12 | HEK-293T | C36(0.87) | LDD1510 | [5] |
| LDCM0276 | AC17 | HEK-293T | C36(0.94) | LDD1515 | [5] |
| LDCM0280 | AC20 | HEK-293T | C36(1.10) | LDD1519 | [5] |
| LDCM0281 | AC21 | HEK-293T | C36(0.96) | LDD1520 | [5] |
| LDCM0284 | AC24 | HEK-293T | C36(1.07) | LDD1523 | [5] |
| LDCM0285 | AC25 | HEK-293T | C36(0.97) | LDD1524 | [5] |
| LDCM0288 | AC28 | HEK-293T | C36(0.85) | LDD1527 | [5] |
| LDCM0289 | AC29 | HEK-293T | C36(1.00) | LDD1528 | [5] |
| LDCM0293 | AC32 | HEK-293T | C36(0.99) | LDD1532 | [5] |
| LDCM0294 | AC33 | HEK-293T | C36(1.09) | LDD1533 | [5] |
| LDCM0297 | AC36 | HEK-293T | C36(0.97) | LDD1536 | [5] |
| LDCM0298 | AC37 | HEK-293T | C36(1.08) | LDD1537 | [5] |
| LDCM0301 | AC4 | HEK-293T | C36(1.15) | LDD1540 | [5] |
| LDCM0302 | AC40 | HEK-293T | C36(0.97) | LDD1541 | [5] |
| LDCM0303 | AC41 | HEK-293T | C36(0.94) | LDD1542 | [5] |
| LDCM0306 | AC44 | HEK-293T | C36(0.85) | LDD1545 | [5] |
| LDCM0307 | AC45 | HEK-293T | C36(1.03) | LDD1546 | [5] |
| LDCM0310 | AC48 | HEK-293T | C36(1.01) | LDD1549 | [5] |
| LDCM0311 | AC49 | HEK-293T | C36(0.97) | LDD1550 | [5] |
| LDCM0312 | AC5 | HEK-293T | C36(0.96) | LDD1551 | [5] |
| LDCM0315 | AC52 | HEK-293T | C36(1.11) | LDD1554 | [5] |
| LDCM0316 | AC53 | HEK-293T | C36(0.92) | LDD1555 | [5] |
| LDCM0319 | AC56 | HEK-293T | C36(1.03) | LDD1558 | [5] |
| LDCM0320 | AC57 | HEK-293T | C36(1.09) | LDD1559 | [5] |
| LDCM0324 | AC60 | HEK-293T | C36(0.79) | LDD1563 | [5] |
| LDCM0325 | AC61 | HEK-293T | C36(1.01) | LDD1564 | [5] |
| LDCM0328 | AC64 | HEK-293T | C36(0.97) | LDD1567 | [5] |
| LDCM0345 | AC8 | HEK-293T | C36(0.89) | LDD1569 | [5] |
| LDCM0248 | AKOS034007472 | HEK-293T | C36(1.09) | LDD1511 | [5] |
| LDCM0356 | AKOS034007680 | HEK-293T | C36(1.05) | LDD1570 | [5] |
| LDCM0275 | AKOS034007705 | HEK-293T | C36(1.11) | LDD1514 | [5] |
| LDCM0367 | CL1 | HEK-293T | C36(1.01) | LDD1571 | [5] |
| LDCM0369 | CL100 | HEK-293T | C36(0.87) | LDD1573 | [5] |
| LDCM0370 | CL101 | HEK-293T | C36(0.80) | LDD1574 | [5] |
| LDCM0373 | CL104 | HEK-293T | C36(1.26) | LDD1577 | [5] |
| LDCM0374 | CL105 | HEK-293T | C36(1.10) | LDD1578 | [5] |
| LDCM0377 | CL108 | HEK-293T | C36(0.91) | LDD1581 | [5] |
| LDCM0378 | CL109 | HEK-293T | C36(1.06) | LDD1582 | [5] |
| LDCM0382 | CL112 | HEK-293T | C36(0.95) | LDD1586 | [5] |
| LDCM0383 | CL113 | HEK-293T | C36(1.20) | LDD1587 | [5] |
| LDCM0386 | CL116 | HEK-293T | C36(0.97) | LDD1590 | [5] |
| LDCM0387 | CL117 | HEK-293T | C36(0.85) | LDD1591 | [5] |
| LDCM0390 | CL12 | HEK-293T | C36(0.66) | LDD1594 | [5] |
| LDCM0391 | CL120 | HEK-293T | C36(1.11) | LDD1595 | [5] |
| LDCM0392 | CL121 | HEK-293T | C36(1.21) | LDD1596 | [5] |
| LDCM0395 | CL124 | HEK-293T | C36(1.19) | LDD1599 | [5] |
| LDCM0396 | CL125 | HEK-293T | C36(0.82) | LDD1600 | [5] |
| LDCM0399 | CL128 | HEK-293T | C36(1.32) | LDD1603 | [5] |
| LDCM0400 | CL13 | HEK-293T | C36(0.97) | LDD1604 | [5] |
| LDCM0403 | CL16 | HEK-293T | C36(0.96) | LDD1607 | [5] |
| LDCM0404 | CL17 | HEK-293T | C36(0.88) | LDD1608 | [5] |
| LDCM0408 | CL20 | HEK-293T | C36(1.07) | LDD1612 | [5] |
| LDCM0409 | CL21 | HEK-293T | C36(0.79) | LDD1613 | [5] |
| LDCM0412 | CL24 | HEK-293T | C36(0.66) | LDD1616 | [5] |
| LDCM0413 | CL25 | HEK-293T | C36(0.78) | LDD1617 | [5] |
| LDCM0416 | CL28 | HEK-293T | C36(0.85) | LDD1620 | [5] |
| LDCM0417 | CL29 | HEK-293T | C36(1.01) | LDD1621 | [5] |
| LDCM0421 | CL32 | HEK-293T | C36(0.73) | LDD1625 | [5] |
| LDCM0422 | CL33 | HEK-293T | C36(0.77) | LDD1626 | [5] |
| LDCM0425 | CL36 | HEK-293T | C36(0.66) | LDD1629 | [5] |
| LDCM0426 | CL37 | HEK-293T | C36(0.92) | LDD1630 | [5] |
| LDCM0429 | CL4 | HEK-293T | C36(0.98) | LDD1633 | [5] |
| LDCM0430 | CL40 | HEK-293T | C36(1.02) | LDD1634 | [5] |
| LDCM0431 | CL41 | HEK-293T | C36(0.89) | LDD1635 | [5] |
| LDCM0434 | CL44 | HEK-293T | C36(0.89) | LDD1638 | [5] |
| LDCM0435 | CL45 | HEK-293T | C36(0.96) | LDD1639 | [5] |
| LDCM0438 | CL48 | HEK-293T | C36(0.55) | LDD1642 | [5] |
| LDCM0439 | CL49 | HEK-293T | C36(1.09) | LDD1643 | [5] |
| LDCM0440 | CL5 | HEK-293T | C36(0.89) | LDD1644 | [5] |
| LDCM0443 | CL52 | HEK-293T | C36(0.88) | LDD1646 | [5] |
| LDCM0444 | CL53 | HEK-293T | C36(0.89) | LDD1647 | [5] |
| LDCM0447 | CL56 | HEK-293T | C36(0.89) | LDD1650 | [5] |
| LDCM0448 | CL57 | HEK-293T | C36(0.78) | LDD1651 | [5] |
| LDCM0452 | CL60 | HEK-293T | C36(0.64) | LDD1655 | [5] |
| LDCM0453 | CL61 | HEK-293T | C36(0.85) | LDD1656 | [5] |
| LDCM0456 | CL64 | HEK-293T | C36(0.90) | LDD1659 | [5] |
| LDCM0457 | CL65 | HEK-293T | C36(0.92) | LDD1660 | [5] |
| LDCM0460 | CL68 | HEK-293T | C36(0.96) | LDD1663 | [5] |
| LDCM0461 | CL69 | HEK-293T | C36(0.88) | LDD1664 | [5] |
| LDCM0465 | CL72 | HEK-293T | C36(0.68) | LDD1668 | [5] |
| LDCM0466 | CL73 | HEK-293T | C36(1.36) | LDD1669 | [5] |
| LDCM0469 | CL76 | HEK-293T | C36(1.09) | LDD1672 | [5] |
| LDCM0470 | CL77 | HEK-293T | C36(0.78) | LDD1673 | [5] |
| LDCM0473 | CL8 | HEK-293T | C36(0.79) | LDD1676 | [5] |
| LDCM0474 | CL80 | HEK-293T | C36(0.85) | LDD1677 | [5] |
| LDCM0475 | CL81 | HEK-293T | C36(0.98) | LDD1678 | [5] |
| LDCM0478 | CL84 | HEK-293T | C36(0.97) | LDD1681 | [5] |
| LDCM0479 | CL85 | HEK-293T | C36(1.36) | LDD1682 | [5] |
| LDCM0482 | CL88 | HEK-293T | C36(0.95) | LDD1685 | [5] |
| LDCM0483 | CL89 | HEK-293T | C36(0.91) | LDD1686 | [5] |
| LDCM0484 | CL9 | HEK-293T | C36(0.95) | LDD1687 | [5] |
| LDCM0487 | CL92 | HEK-293T | C36(0.72) | LDD1690 | [5] |
| LDCM0488 | CL93 | HEK-293T | C36(0.92) | LDD1691 | [5] |
| LDCM0491 | CL96 | HEK-293T | C36(0.87) | LDD1694 | [5] |
| LDCM0492 | CL97 | HEK-293T | C36(0.92) | LDD1695 | [5] |
| LDCM0022 | KB02 | HEK-293T | C36(0.94) | LDD1492 | [5] |
| LDCM0023 | KB03 | HEK-293T | C36(1.20) | LDD1497 | [5] |
| LDCM0024 | KB05 | HEK-293T | C36(1.00) | LDD1502 | [5] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Transporter and channel
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Methylosome subunit pICln (CLNS1A) | PICln (TC 1.A.47) family | P54105 | |||
| Syntenin-1 (SDCBP) | . | O00560 | |||
Other
References
