Details of the Target
General Information of Target
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
SAA-alkyne Probe Info |
 |
1.27 | LDD0252 | [1] | |
|
TH211 Probe Info |
 |
Y294(20.00) | LDD0260 | [2] | |
|
ONAyne Probe Info |
 |
K304(5.88) | LDD0275 | [3] | |
|
OPA-S-S-alkyne Probe Info |
 |
K182(0.73); K340(1.47); K107(1.78); K323(2.24) | LDD3494 | [4] | |
|
Probe 1 Probe Info |
 |
Y246(12.73); Y336(72.13) | LDD3495 | [5] | |
|
HHS-475 Probe Info |
 |
Y336(0.94); Y294(1.11) | LDD0264 | [6] | |
|
HHS-465 Probe Info |
 |
Y294(6.70) | LDD2237 | [7] | |
|
5E-2FA Probe Info |
 |
H184(0.00); H341(0.00); H243(0.00) | LDD2235 | [8] | |
|
ATP probe Probe Info |
 |
K287(0.00); K182(0.00); K323(0.00); K65(0.00) | LDD0199 | [9] | |
|
m-APA Probe Info |
 |
H184(0.00); H243(0.00); H341(0.00) | LDD2231 | [8] | |
|
ATP probe Probe Info |
 |
N.A. | LDD0035 | [10] | |
|
SF Probe Info |
 |
N.A. | LDD0028 | [11] | |
|
STPyne Probe Info |
 |
N.A. | LDD0009 | [12] | |
|
Acrolein Probe Info |
 |
H243(0.00); H184(0.00); H341(0.00); H307(0.00) | LDD0217 | [13] | |
|
Crotonaldehyde Probe Info |
 |
H307(0.00); H243(0.00) | LDD0219 | [13] | |
|
Methacrolein Probe Info |
 |
N.A. | LDD0218 | [13] | |
|
AOyne Probe Info |
 |
14.50 | LDD0443 | [14] | |
|
HHS-482 Probe Info |
 |
Y294(0.60) | LDD2239 | [7] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C017 Probe Info |
 |
6.87 | LDD1725 | [15] | |
|
C052 Probe Info |
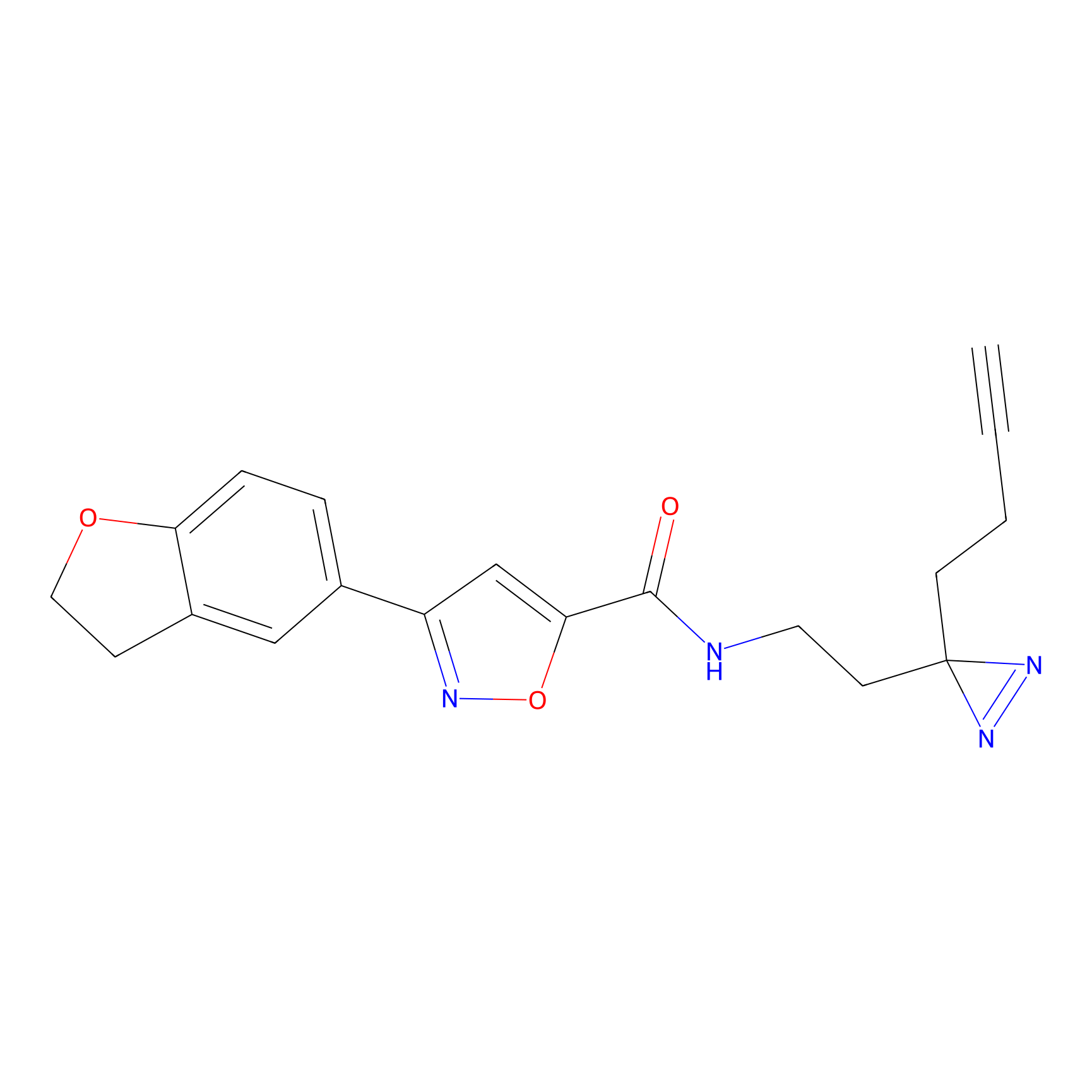 |
5.74 | LDD1750 | [15] | |
|
C158 Probe Info |
 |
11.39 | LDD1838 | [15] | |
|
C187 Probe Info |
 |
15.56 | LDD1865 | [15] | |
|
C191 Probe Info |
 |
10.06 | LDD1868 | [15] | |
|
C207 Probe Info |
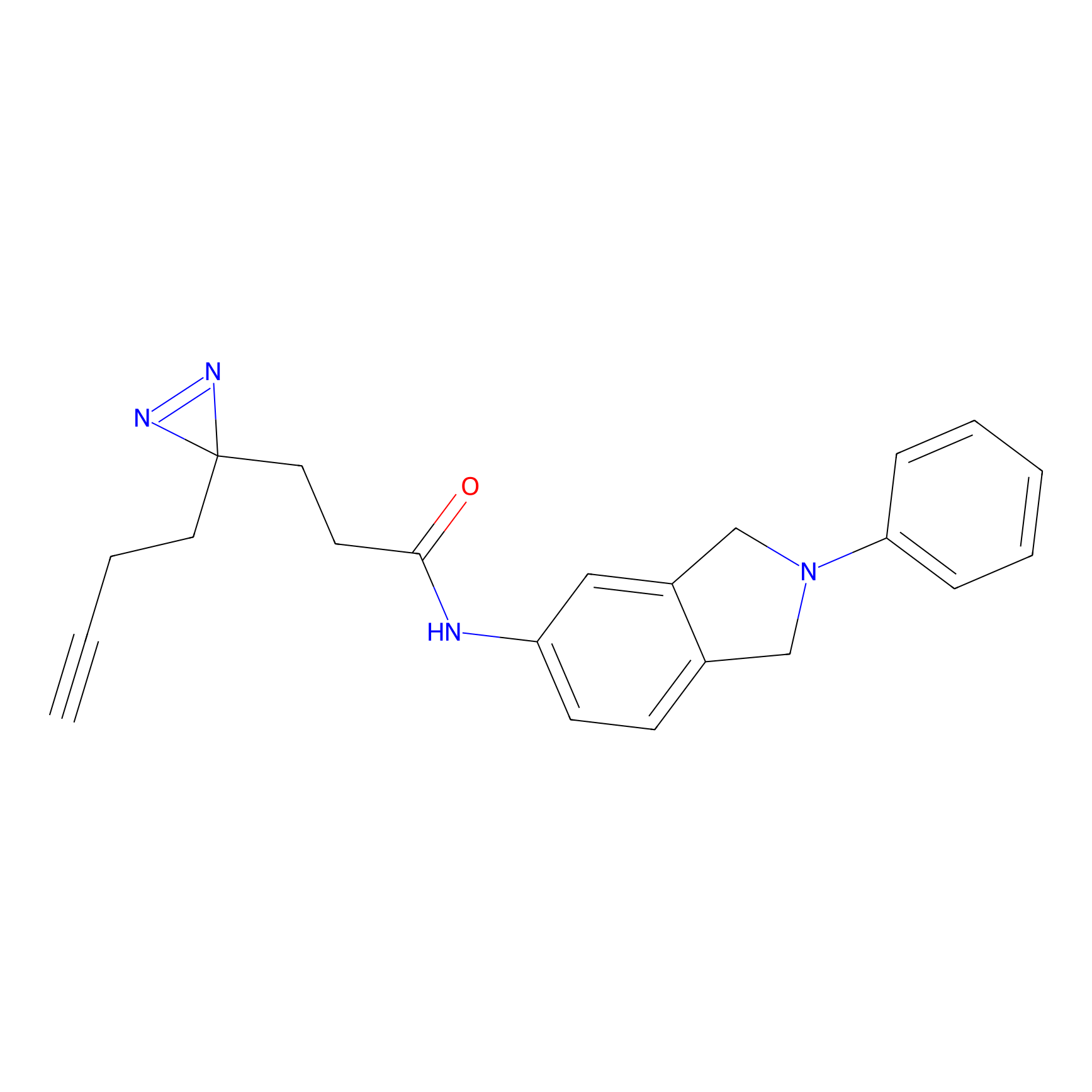 |
18.13 | LDD1882 | [15] | |
|
C338 Probe Info |
 |
10.70 | LDD2001 | [15] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0108 | Chloroacetamide | HeLa | H243(0.00); H184(0.00); H341(0.00); H307(0.00) | LDD0222 | [13] |
| LDCM0116 | HHS-0101 | DM93 | Y336(0.94); Y294(1.11) | LDD0264 | [6] |
| LDCM0117 | HHS-0201 | DM93 | Y336(0.66); Y294(0.76) | LDD0265 | [6] |
| LDCM0118 | HHS-0301 | DM93 | Y294(0.50); Y336(9.07) | LDD0266 | [6] |
| LDCM0119 | HHS-0401 | DM93 | Y294(0.84); Y336(4.27) | LDD0267 | [6] |
| LDCM0120 | HHS-0701 | DM93 | Y294(0.62); Y336(12.96) | LDD0268 | [6] |
| LDCM0107 | IAA | HeLa | H243(0.00); H184(0.00); H307(0.00); H341(0.00) | LDD0221 | [13] |
| LDCM0109 | NEM | HeLa | H243(0.00); H184(0.00); H307(0.00); H341(0.00) | LDD0223 | [13] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Transporter and channel
Other
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Very low-density lipoprotein receptor (VLDLR) | . | P98155 | |||
References
