Details of the Target
General Information of Target
| Target ID | LDTP02884 | |||||
|---|---|---|---|---|---|---|
| Target Name | Heat shock 70 kDa protein 6 (HSPA6) | |||||
| Gene Name | HSPA6 | |||||
| Gene ID | 3310 | |||||
| Synonyms |
HSP70B'; Heat shock 70 kDa protein 6; Heat shock 70 kDa protein B' |
|||||
| 3D Structure | ||||||
| Sequence |
MQAPRELAVGIDLGTTYSCVGVFQQGRVEILANDQGNRTTPSYVAFTDTERLVGDAAKSQ
AALNPHNTVFDAKRLIGRKFADTTVQSDMKHWPFRVVSEGGKPKVRVCYRGEDKTFYPEE ISSMVLSKMKETAEAYLGQPVKHAVITVPAYFNDSQRQATKDAGAIAGLNVLRIINEPTA AAIAYGLDRRGAGERNVLIFDLGGGTFDVSVLSIDAGVFEVKATAGDTHLGGEDFDNRLV NHFMEEFRRKHGKDLSGNKRALRRLRTACERAKRTLSSSTQATLEIDSLFEGVDFYTSIT RARFEELCSDLFRSTLEPVEKALRDAKLDKAQIHDVVLVGGSTRIPKVQKLLQDFFNGKE LNKSINPDEAVAYGAAVQAAVLMGDKCEKVQDLLLLDVAPLSLGLETAGGVMTTLIQRNA TIPTKQTQTFTTYSDNQPGVFIQVYEGERAMTKDNNLLGRFELSGIPPAPRGVPQIEVTF DIDANGILSVTATDRSTGKANKITITNDKGRLSKEEVERMVHEAEQYKAEDEAQRDRVAA KNSLEAHVFHVKGSLQEESLRDKIPEEDRRKMQDKCREVLAWLEHNQLAEKEEYEHQKRE LEQICRPIFSRLYGGPGVPGGSSCGTQARQGDPSTGPIIEEVD |
|||||
| Target Bioclass |
Other
|
|||||
| Family |
Heat shock protein 70 family
|
|||||
| Function |
Molecular chaperone implicated in a wide variety of cellular processes, including protection of the proteome from stress, folding and transport of newly synthesized polypeptides, activation of proteolysis of misfolded proteins and the formation and dissociation of protein complexes. Plays a pivotal role in the protein quality control system, ensuring the correct folding of proteins, the re-folding of misfolded proteins and controlling the targeting of proteins for subsequent degradation. This is achieved through cycles of ATP binding, ATP hydrolysis and ADP release, mediated by co-chaperones. The affinity for polypeptides is regulated by its nucleotide bound state. In the ATP-bound form, it has a low affinity for substrate proteins. However, upon hydrolysis of the ATP to ADP, it undergoes a conformational change that increases its affinity for substrate proteins. It goes through repeated cycles of ATP hydrolysis and nucleotide exchange, which permits cycles of substrate binding and release.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
P2 Probe Info |
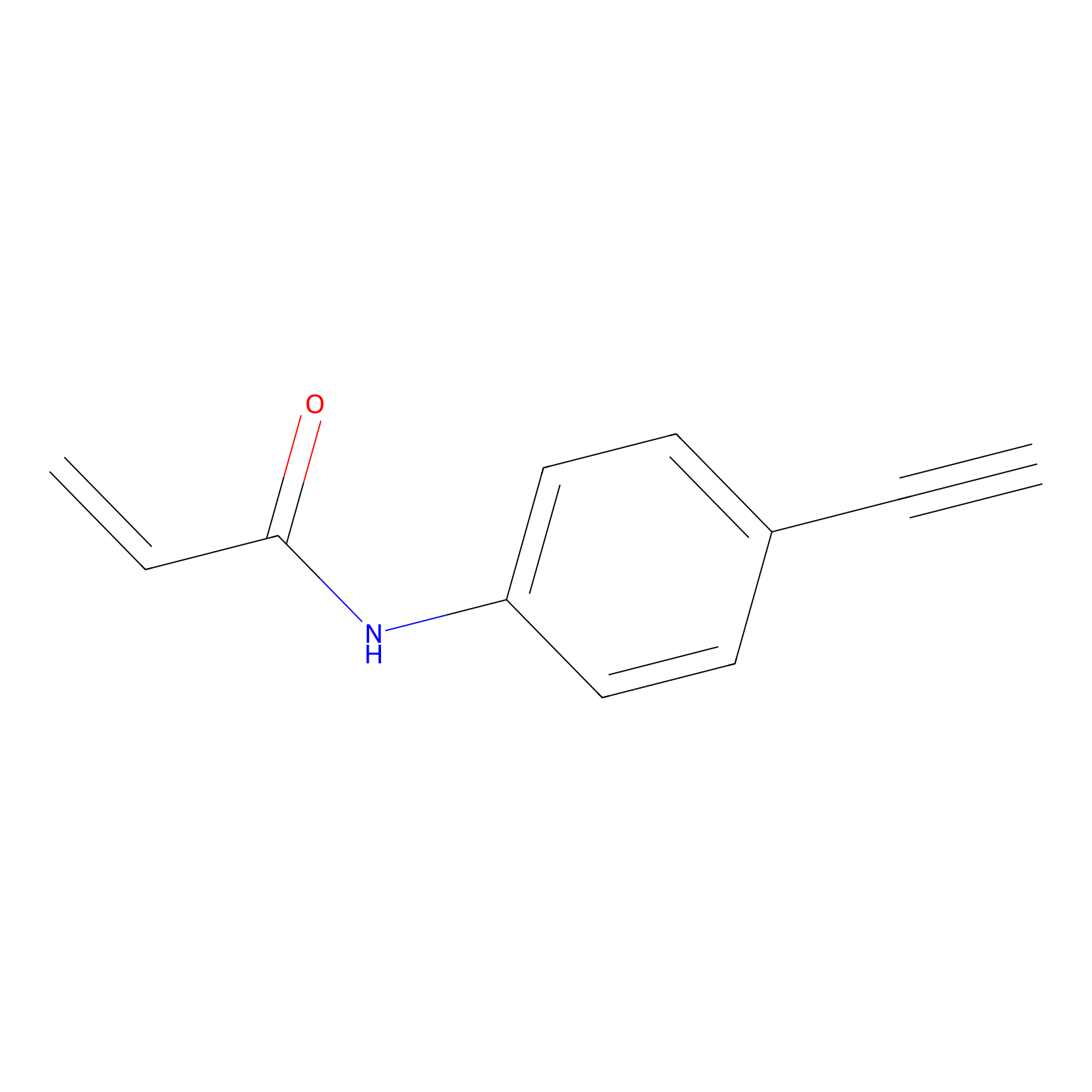 |
1.88 | LDD0449 | [1] | |
|
P3 Probe Info |
 |
1.71 | LDD0450 | [1] | |
|
P8 Probe Info |
 |
2.27 | LDD0451 | [1] | |
|
YN-1 Probe Info |
 |
100.00 | LDD0444 | [2] | |
|
YN-4 Probe Info |
 |
100.00 | LDD0445 | [2] | |
|
AZ-9 Probe Info |
 |
E317(1.00); D354(0.89); D48(10.00); Y185(0.78) | LDD2208 | [3] | |
|
ONAyne Probe Info |
 |
K514(0.00); K330(0.00) | LDD0273 | [4] | |
|
HHS-475 Probe Info |
 |
Y43(0.96); Y373(1.91) | LDD0264 | [5] | |
|
HHS-465 Probe Info |
 |
Y185(5.12); Y43(10.00); Y433(4.63) | LDD2237 | [6] | |
|
DBIA Probe Info |
 |
C308(1.13) | LDD2171 | [7] | |
|
5E-2FA Probe Info |
 |
N.A. | LDD2235 | [8] | |
|
IA-alkyne Probe Info |
 |
C308(0.00); C269(0.00) | LDD0166 | [9] | |
|
HHS-482 Probe Info |
 |
Y373(2.39) | LDD2239 | [6] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
FFF probe13 Probe Info |
 |
6.13 | LDD0475 | [10] | |
|
FFF probe14 Probe Info |
 |
5.47 | LDD0477 | [10] | |
|
FFF probe3 Probe Info |
 |
5.61 | LDD0465 | [10] | |
|
OEA-DA Probe Info |
 |
3.43 | LDD0046 | [11] | |
|
STS-1 Probe Info |
 |
N.A. | LDD0069 | [12] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0020 | ARS-1620 | HCC44 | C308(1.13) | LDD2171 | [7] |
| LDCM0116 | HHS-0101 | DM93 | Y43(0.96); Y373(1.91) | LDD0264 | [5] |
| LDCM0117 | HHS-0201 | DM93 | Y43(0.84); Y373(1.06) | LDD0265 | [5] |
| LDCM0118 | HHS-0301 | DM93 | Y43(0.85); Y373(6.31) | LDD0266 | [5] |
| LDCM0119 | HHS-0401 | DM93 | Y43(0.75); Y373(1.95) | LDD0267 | [5] |
| LDCM0120 | HHS-0701 | DM93 | Y43(0.48); Y373(1.18) | LDD0268 | [5] |
| LDCM0021 | THZ1 | HCT 116 | C308(1.13) | LDD2173 | [7] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Other
References
