Details of the Target
General Information of Target
Target Site Mutations in Different Cell Lines
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
FBPP2 Probe Info |
 |
5.75 | LDD0324 | [1] | |
|
FBP2 Probe Info |
 |
2.70 | LDD0323 | [1] | |
|
STPyne Probe Info |
 |
K123(7.23) | LDD0277 | [2] | |
|
IPM Probe Info |
 |
C313(0.00); C302(0.00) | LDD0241 | [3] | |
|
OPA-S-S-alkyne Probe Info |
 |
K222(0.58); K62(1.13); K322(1.38) | LDD3494 | [4] | |
|
DBIA Probe Info |
 |
C365(2.74) | LDD3373 | [5] | |
|
HHS-475 Probe Info |
 |
Y341(0.81) | LDD0264 | [6] | |
|
5E-2FA Probe Info |
 |
H171(0.00); H392(0.00) | LDD2235 | [7] | |
|
m-APA Probe Info |
 |
H171(0.00); H392(0.00) | LDD2231 | [7] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0162 | [8] | |
|
Acrolein Probe Info |
 |
H392(0.00); H217(0.00) | LDD0217 | [9] | |
|
Crotonaldehyde Probe Info |
 |
H392(0.00); C313(0.00) | LDD0219 | [9] | |
|
NAIA_5 Probe Info |
 |
N.A. | LDD2223 | [10] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C275 Probe Info |
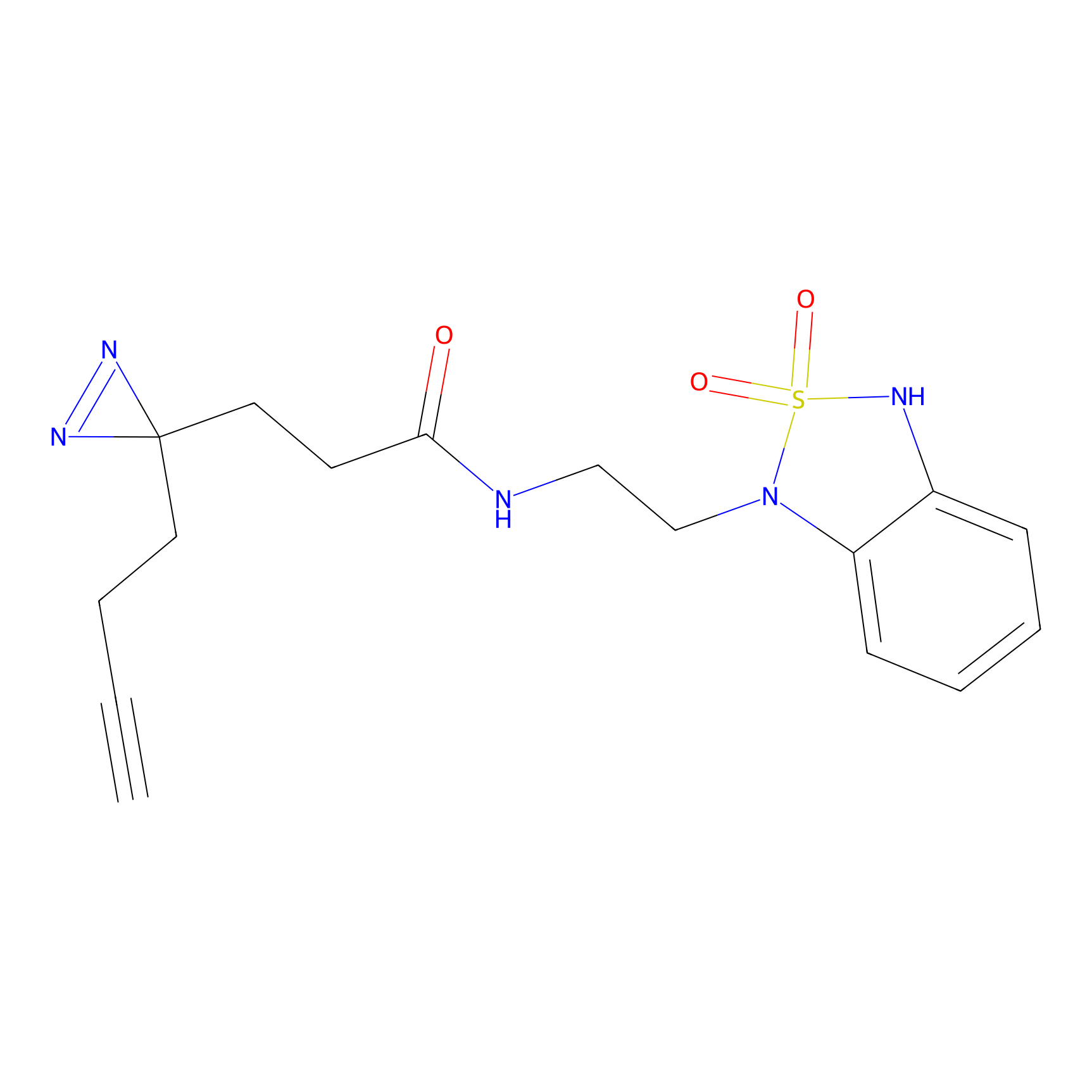 |
5.21 | LDD1945 | [11] | |
|
C313 Probe Info |
 |
13.64 | LDD1980 | [11] | |
|
C344 Probe Info |
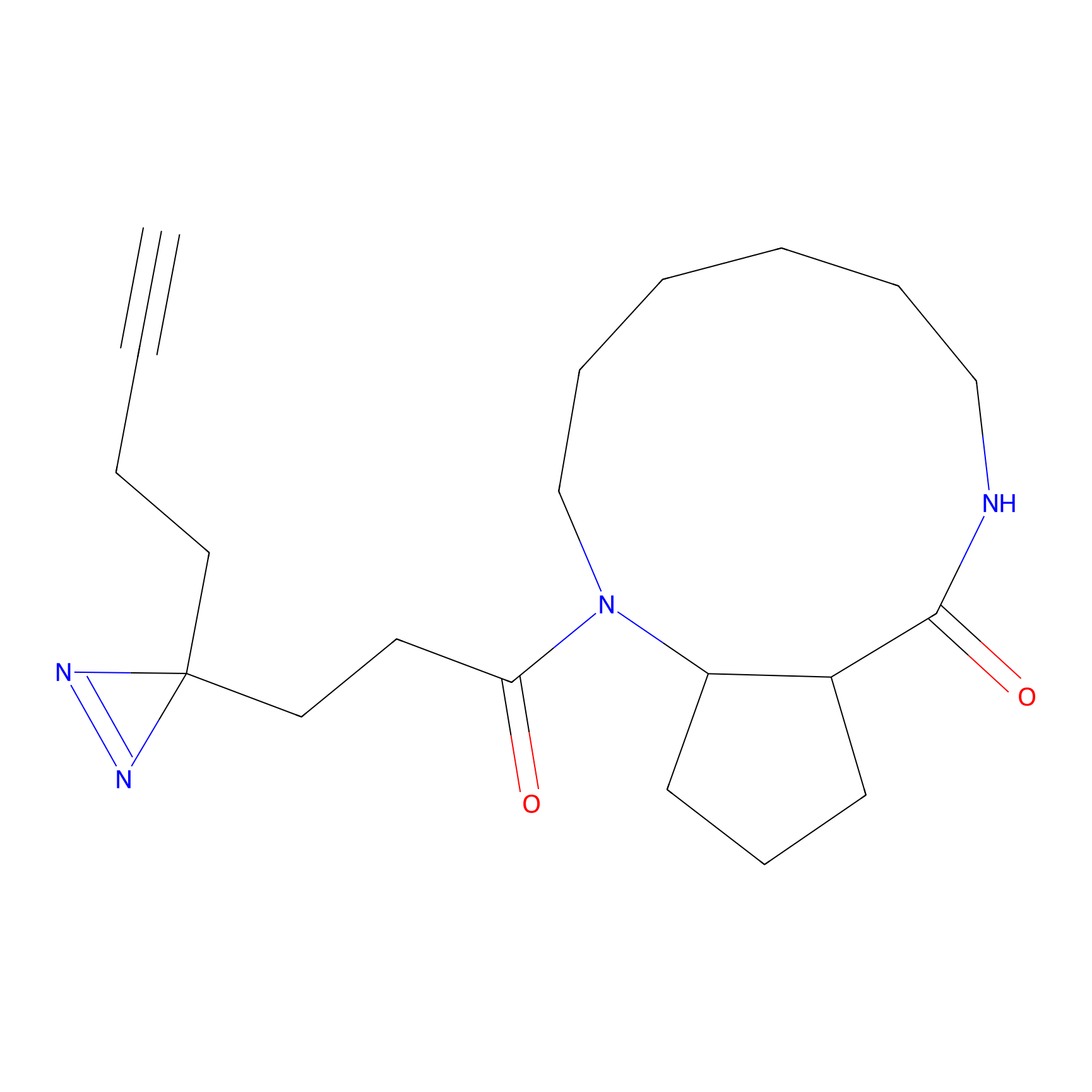 |
5.03 | LDD2006 | [11] | |
|
Alk-rapa Probe Info |
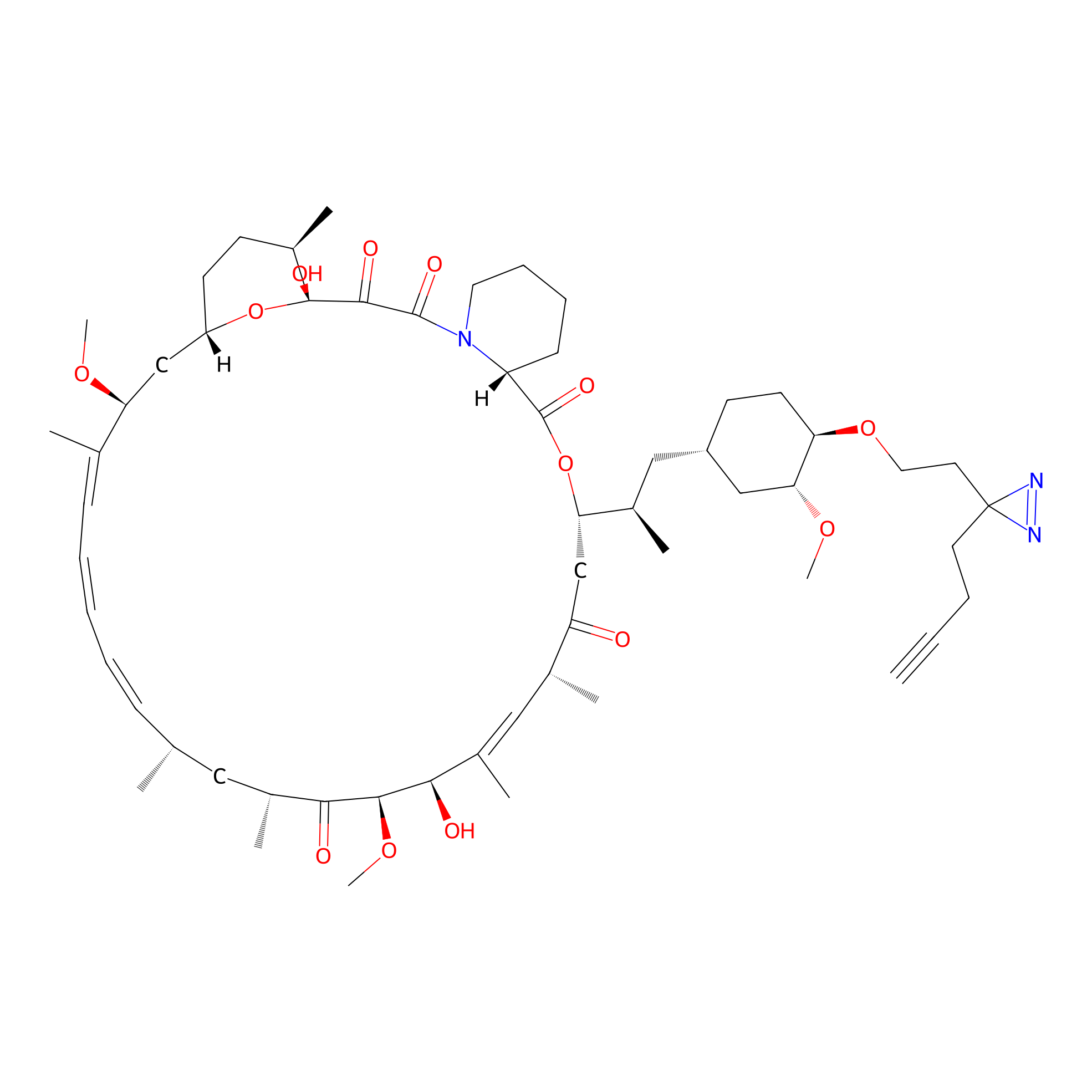 |
2.05 | LDD0213 | [12] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0108 | Chloroacetamide | HeLa | C313(0.00); H392(0.00); H217(0.00) | LDD0222 | [9] |
| LDCM0116 | HHS-0101 | DM93 | Y341(0.81) | LDD0264 | [6] |
| LDCM0117 | HHS-0201 | DM93 | Y341(0.60) | LDD0265 | [6] |
| LDCM0118 | HHS-0301 | DM93 | Y341(0.73) | LDD0266 | [6] |
| LDCM0119 | HHS-0401 | DM93 | Y341(0.55) | LDD0267 | [6] |
| LDCM0120 | HHS-0701 | DM93 | Y341(1.13) | LDD0268 | [6] |
| LDCM0107 | IAA | HeLa | H392(0.00); H217(0.00) | LDD0221 | [9] |
| LDCM0022 | KB02 | 22RV1 | C365(1.96) | LDD2243 | [5] |
| LDCM0023 | KB03 | 22RV1 | C365(2.69) | LDD2660 | [5] |
| LDCM0024 | KB05 | OCUG-1 | C365(2.74) | LDD3373 | [5] |
| LDCM0109 | NEM | HeLa | H392(0.00); H217(0.00) | LDD0223 | [9] |
| LDCM0090 | Rapamycin | JHH-7 | 2.05 | LDD0213 | [12] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Methionine-R-sulfoxide reductase B1 (MSRB1) | MsrB Met sulfoxide reductase family | Q9NZV6 | |||
| Protein disulfide-isomerase A3 (PDIA3) | Protein disulfide isomerase family | P30101 | |||
Transporter and channel
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Amyloid-beta precursor protein (APP) | APP family | P05067 | |||
| Alpha-synuclein (SNCA) | Synuclein family | P37840 | |||
Transcription factor
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Protein c-Fos (FOS) | BZIP family | P01100 | |||
| Peroxisome proliferator-activated receptor gamma (PPARG) | Nuclear hormone receptor family | P37231 | |||
Other
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Pituitary adenylate cyclase-activating polypeptide (ADCYAP1) | Glucagon family | P18509 | |||
| Disrupted in schizophrenia 1 protein (DISC1) | . | Q9NRI5 | |||
References
