Details of the Target
General Information of Target
| Target ID | LDTP01922 | |||||
|---|---|---|---|---|---|---|
| Target Name | Hemoglobin subunit epsilon (HBE1) | |||||
| Gene Name | HBE1 | |||||
| Gene ID | 3046 | |||||
| Synonyms |
HBE; Hemoglobin subunit epsilon; Epsilon-globin; Hemoglobin epsilon chain |
|||||
| 3D Structure | ||||||
| Sequence |
MVHFTAEEKAAVTSLWSKMNVEEAGGEALGRLLVVYPWTQRFFDSFGNLSSPSAILGNPK
VKAHGKKVLTSFGDAIKNMDNLKPAFAKLSELHCDKLHVDPENFKLLGNVMVIILATHFG KEFTPEVQAAWQKLVSAVAIALAHKYH |
|||||
| Target Bioclass |
Other
|
|||||
| Family |
Globin family
|
|||||
| Function | The epsilon chain is a beta-type chain of early mammalian embryonic hemoglobin. | |||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
A-EBA Probe Info |
 |
3.87 | LDD0215 | [1] | |
|
STPyne Probe Info |
 |
K62(6.56) | LDD0277 | [2] | |
|
HPAP Probe Info |
 |
3.00 | LDD0063 | [3] | |
|
DBIA Probe Info |
 |
C94(1.33) | LDD1507 | [4] | |
|
2PCA Probe Info |
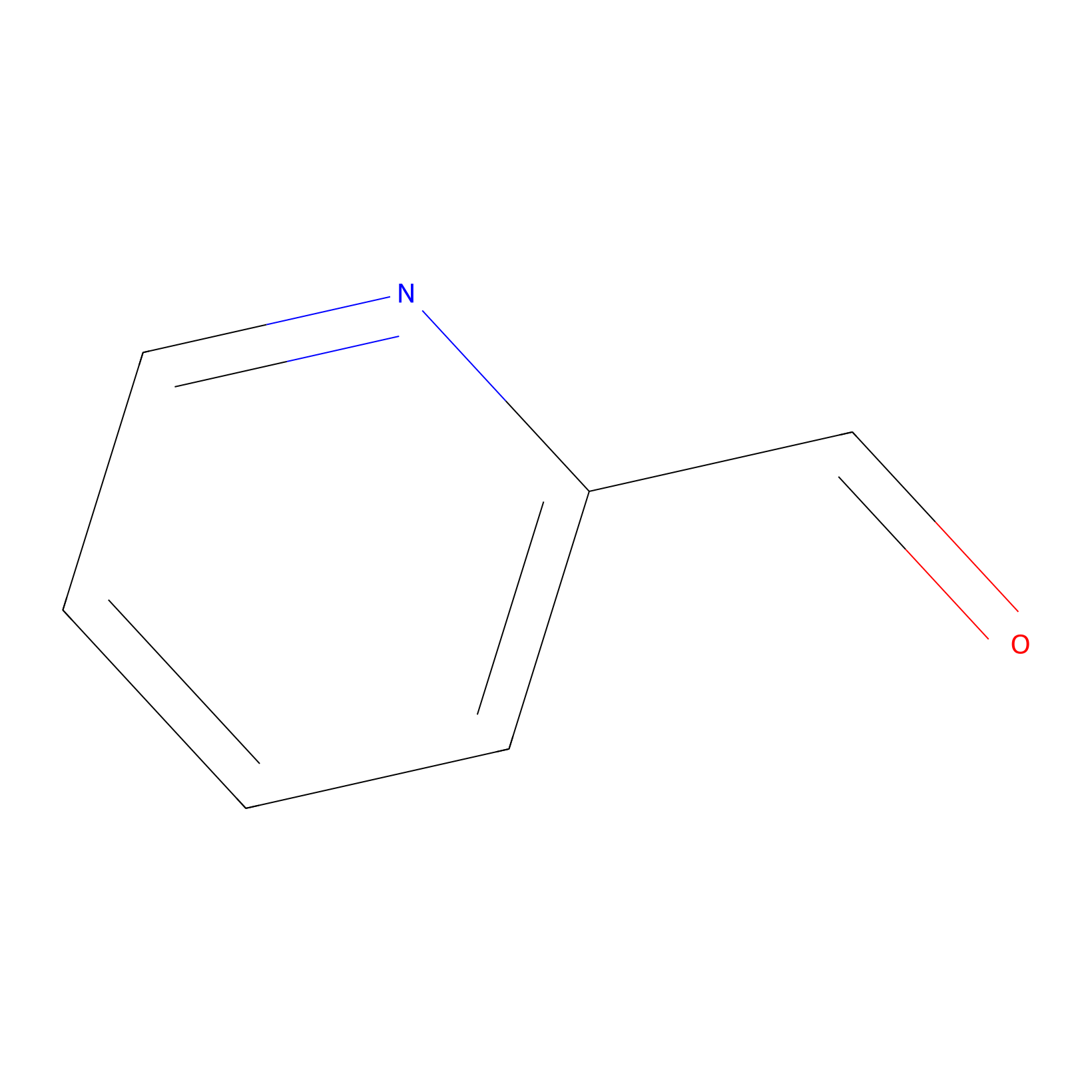 |
N.A. | LDD0034 | [5] | |
|
VSF Probe Info |
 |
N.A. | LDD0007 | [6] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0217 | [7] | |
|
TER-AC Probe Info |
 |
N.A. | LDD0426 | [8] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
FFF probe3 Probe Info |
 |
7.51 | LDD0465 | [9] | |
|
DA-2 Probe Info |
 |
N.A. | LDD0070 | [10] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0214 | AC1 | HEK-293T | C94(1.33) | LDD1507 | [4] |
| LDCM0215 | AC10 | HEK-293T | C94(1.14) | LDD1508 | [4] |
| LDCM0276 | AC17 | HEK-293T | C94(1.32) | LDD1515 | [4] |
| LDCM0277 | AC18 | HEK-293T | C94(1.16) | LDD1516 | [4] |
| LDCM0279 | AC2 | HEK-293T | C94(1.26) | LDD1518 | [4] |
| LDCM0284 | AC24 | HEK-293T | C94(1.07) | LDD1523 | [4] |
| LDCM0285 | AC25 | HEK-293T | C94(1.26) | LDD1524 | [4] |
| LDCM0286 | AC26 | HEK-293T | C94(1.27) | LDD1525 | [4] |
| LDCM0293 | AC32 | HEK-293T | C94(0.95) | LDD1532 | [4] |
| LDCM0294 | AC33 | HEK-293T | C94(1.55) | LDD1533 | [4] |
| LDCM0295 | AC34 | HEK-293T | C94(1.22) | LDD1534 | [4] |
| LDCM0302 | AC40 | HEK-293T | C94(1.02) | LDD1541 | [4] |
| LDCM0303 | AC41 | HEK-293T | C94(1.33) | LDD1542 | [4] |
| LDCM0304 | AC42 | HEK-293T | C94(1.18) | LDD1543 | [4] |
| LDCM0310 | AC48 | HEK-293T | C94(0.98) | LDD1549 | [4] |
| LDCM0311 | AC49 | HEK-293T | C94(0.98) | LDD1550 | [4] |
| LDCM0313 | AC50 | HEK-293T | C94(0.95) | LDD1552 | [4] |
| LDCM0319 | AC56 | HEK-293T | C94(0.99) | LDD1558 | [4] |
| LDCM0320 | AC57 | HEK-293T | C94(0.29) | LDD1559 | [4] |
| LDCM0321 | AC58 | HEK-293T | C94(0.41) | LDD1560 | [4] |
| LDCM0328 | AC64 | HEK-293T | C94(0.84) | LDD1567 | [4] |
| LDCM0345 | AC8 | HEK-293T | C94(0.87) | LDD1569 | [4] |
| LDCM0356 | AKOS034007680 | HEK-293T | C94(1.46) | LDD1570 | [4] |
| LDCM0275 | AKOS034007705 | HEK-293T | C94(0.95) | LDD1514 | [4] |
| LDCM0108 | Chloroacetamide | HeLa | N.A. | LDD0222 | [7] |
| LDCM0369 | CL100 | HEK-293T | C94(1.09) | LDD1573 | [4] |
| LDCM0373 | CL104 | HEK-293T | C94(1.01) | LDD1577 | [4] |
| LDCM0377 | CL108 | HEK-293T | C94(1.00) | LDD1581 | [4] |
| LDCM0382 | CL112 | HEK-293T | C94(1.17) | LDD1586 | [4] |
| LDCM0386 | CL116 | HEK-293T | C94(1.06) | LDD1590 | [4] |
| LDCM0390 | CL12 | HEK-293T | C94(1.11) | LDD1594 | [4] |
| LDCM0391 | CL120 | HEK-293T | C94(1.02) | LDD1595 | [4] |
| LDCM0395 | CL124 | HEK-293T | C94(1.00) | LDD1599 | [4] |
| LDCM0399 | CL128 | HEK-293T | C94(0.74) | LDD1603 | [4] |
| LDCM0403 | CL16 | HEK-293T | C94(0.92) | LDD1607 | [4] |
| LDCM0404 | CL17 | HEK-293T | C94(0.85) | LDD1608 | [4] |
| LDCM0405 | CL18 | HEK-293T | C94(0.99) | LDD1609 | [4] |
| LDCM0412 | CL24 | HEK-293T | C94(1.26) | LDD1616 | [4] |
| LDCM0416 | CL28 | HEK-293T | C94(1.10) | LDD1620 | [4] |
| LDCM0417 | CL29 | HEK-293T | C94(0.94) | LDD1621 | [4] |
| LDCM0419 | CL30 | HEK-293T | C94(0.83) | LDD1623 | [4] |
| LDCM0425 | CL36 | HEK-293T | C94(1.21) | LDD1629 | [4] |
| LDCM0429 | CL4 | HEK-293T | C94(1.04) | LDD1633 | [4] |
| LDCM0430 | CL40 | HEK-293T | C94(1.01) | LDD1634 | [4] |
| LDCM0431 | CL41 | HEK-293T | C94(0.77) | LDD1635 | [4] |
| LDCM0432 | CL42 | HEK-293T | C94(0.85) | LDD1636 | [4] |
| LDCM0438 | CL48 | HEK-293T | C94(1.38) | LDD1642 | [4] |
| LDCM0440 | CL5 | HEK-293T | C94(0.76) | LDD1644 | [4] |
| LDCM0443 | CL52 | HEK-293T | C94(1.12) | LDD1646 | [4] |
| LDCM0444 | CL53 | HEK-293T | C94(1.16) | LDD1647 | [4] |
| LDCM0445 | CL54 | HEK-293T | C94(1.24) | LDD1648 | [4] |
| LDCM0451 | CL6 | HEK-293T | C94(0.81) | LDD1654 | [4] |
| LDCM0452 | CL60 | HEK-293T | C94(1.31) | LDD1655 | [4] |
| LDCM0456 | CL64 | HEK-293T | C94(1.33) | LDD1659 | [4] |
| LDCM0457 | CL65 | HEK-293T | C94(1.42) | LDD1660 | [4] |
| LDCM0458 | CL66 | HEK-293T | C94(1.39) | LDD1661 | [4] |
| LDCM0465 | CL72 | HEK-293T | C94(1.33) | LDD1668 | [4] |
| LDCM0469 | CL76 | HEK-293T | C94(0.94) | LDD1672 | [4] |
| LDCM0470 | CL77 | HEK-293T | C94(1.32) | LDD1673 | [4] |
| LDCM0471 | CL78 | HEK-293T | C94(1.07) | LDD1674 | [4] |
| LDCM0478 | CL84 | HEK-293T | C94(1.22) | LDD1681 | [4] |
| LDCM0482 | CL88 | HEK-293T | C94(0.90) | LDD1685 | [4] |
| LDCM0483 | CL89 | HEK-293T | C94(0.96) | LDD1686 | [4] |
| LDCM0485 | CL90 | HEK-293T | C94(1.04) | LDD1688 | [4] |
| LDCM0491 | CL96 | HEK-293T | C94(1.11) | LDD1694 | [4] |
| LDCM0107 | IAA | HeLa | N.A. | LDD0221 | [7] |
| LDCM0109 | NEM | HeLa | N.A. | LDD0229 | [7] |
| LDCM0014 | Panhematin | K562 | 3.00 | LDD0063 | [3] |
The Interaction Atlas With This Target
References
