Details of the Target
General Information of Target
| Target ID | LDTP11336 | |||||
|---|---|---|---|---|---|---|
| Target Name | Chitinase domain-containing protein 1 (CHID1) | |||||
| Gene Name | CHID1 | |||||
| Gene ID | 66005 | |||||
| Synonyms |
Chitinase domain-containing protein 1; Stabilin-1-interacting chitinase-like protein; SI-CLP |
|||||
| 3D Structure | ||||||
| Sequence |
MTQGKLSVANKAPGTEGQQQVHGEKKEAPAVPSAPPSYEEATSGEGMKAGAFPPAPTAVP
LHPSWAYVDPSSSSSYDNGFPTGDHELFTTFSWDDQKVRRVFVRKVYTILLIQLLVTLAV VALFTFCDPVKDYVQANPGWYWASYAVFFATYLTLACCSGPRRHFPWNLILLTVFTLSMA YLTGMLSSYYNTTSVLLCLGITALVCLSVTVFSFQTKFDFTSCQGVLFVLLMTLFFSGLI LAILLPFQYVPWLHAVYAALGAGVFTLFLALDTQLLMGNRRHSLSPEEYIFGALNIYLDI IYIFTFFLQLFGTNRE |
|||||
| Target Bioclass |
Enzyme
|
|||||
| Family |
Glycosyl hydrolase 18 family
|
|||||
| Subcellular location |
Secreted
|
|||||
| Function |
Saccharide- and LPS-binding protein with possible roles in pathogen sensing and endotoxin neutralization. Ligand-binding specificity relates to the length of the oligosaccharides, with preference for chitotetraose (in vitro).
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| A375 | SNV: p.P269S | DBIA Probe Info | |||
| HCC1143 | SNV: p.V180L | DBIA Probe Info | |||
| IGROV1 | Deletion: p.V375_G383del | DBIA Probe Info | |||
| KASUMI2 | SNV: p.S257G | . | |||
| MOLT4 | SNV: p.R116C; p.R292Ter | . | |||
| RL952 | SNV: p.D262N | DBIA Probe Info | |||
| TOV21G | SNV: p.L224I | . | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
12.00 | LDD0403 | [1] | |
|
DBIA Probe Info |
 |
C306(3.62) | LDD3314 | [2] | |
|
BTD Probe Info |
 |
C68(0.51) | LDD2109 | [3] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0224 | [4] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0032 | [5] | |
|
NAIA_5 Probe Info |
 |
N.A. | LDD2223 | [6] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C017 Probe Info |
 |
11.63 | LDD1725 | [7] | |
|
C102 Probe Info |
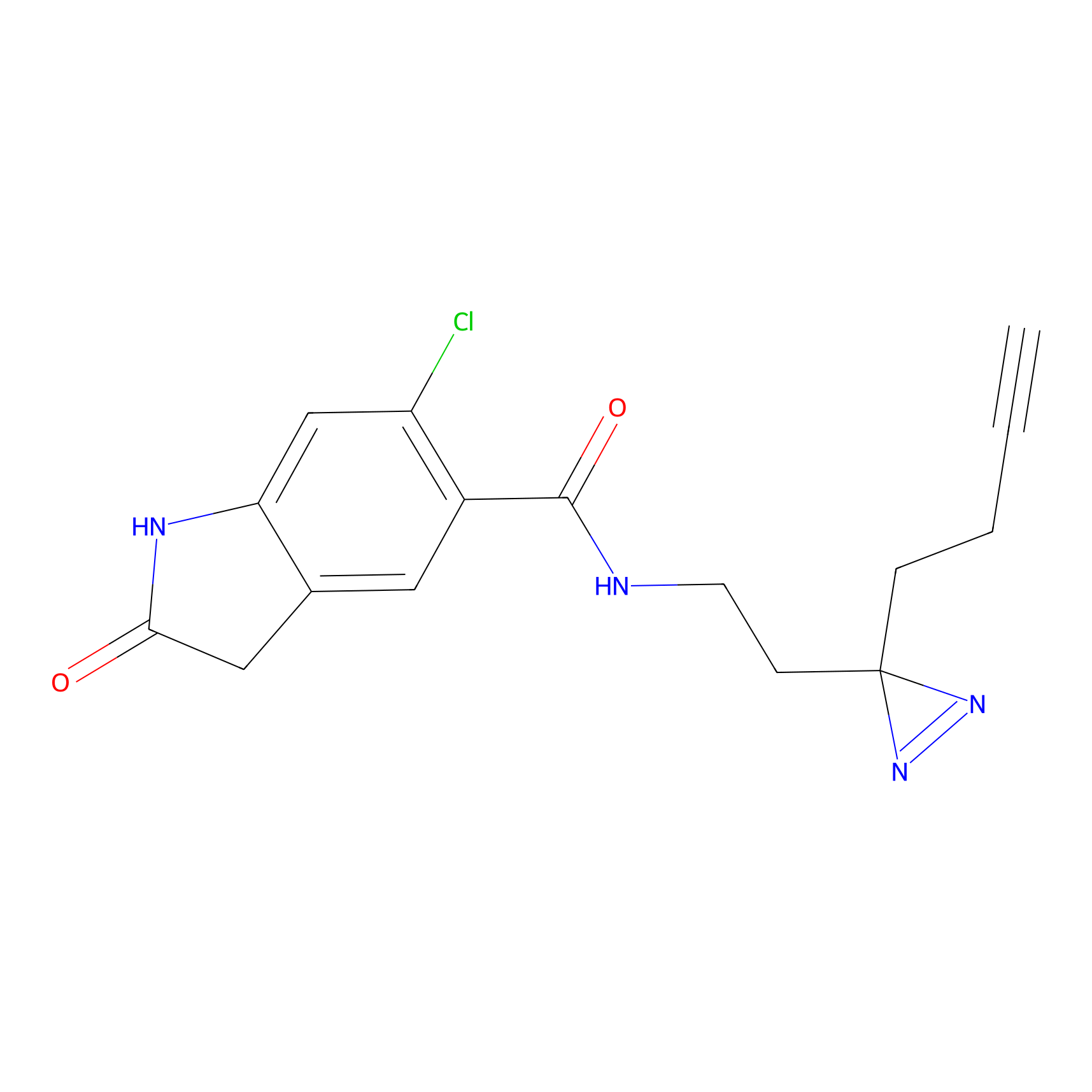 |
6.15 | LDD1790 | [7] | |
|
C116 Probe Info |
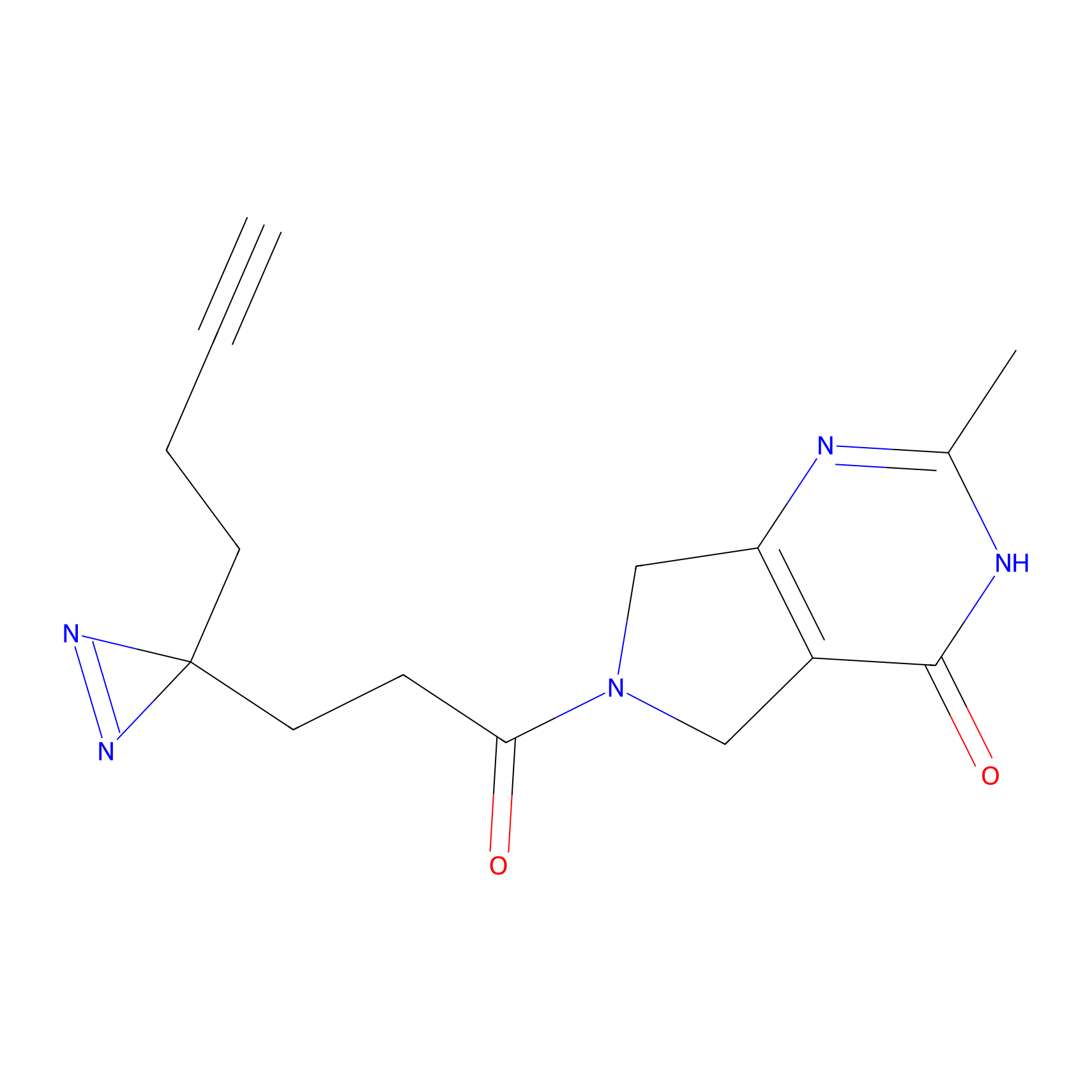 |
5.43 | LDD1803 | [7] | |
|
C338 Probe Info |
 |
29.24 | LDD2001 | [7] | |
|
C355 Probe Info |
 |
39.95 | LDD2016 | [7] | |
|
VE-P Probe Info |
 |
N.A. | LDD0396 | [8] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0270 | AC15 | HEK-293T | C281(1.06) | LDD1513 | [9] |
| LDCM0281 | AC21 | HEK-293T | C281(0.98) | LDD1520 | [9] |
| LDCM0283 | AC23 | HEK-293T | C281(1.18) | LDD1522 | [9] |
| LDCM0289 | AC29 | HEK-293T | C281(1.05) | LDD1528 | [9] |
| LDCM0292 | AC31 | HEK-293T | C281(1.03) | LDD1531 | [9] |
| LDCM0298 | AC37 | HEK-293T | C281(1.21) | LDD1537 | [9] |
| LDCM0300 | AC39 | HEK-293T | C281(1.18) | LDD1539 | [9] |
| LDCM0307 | AC45 | HEK-293T | C281(1.10) | LDD1546 | [9] |
| LDCM0309 | AC47 | HEK-293T | C281(1.09) | LDD1548 | [9] |
| LDCM0312 | AC5 | HEK-293T | C281(1.09) | LDD1551 | [9] |
| LDCM0316 | AC53 | HEK-293T | C281(1.13) | LDD1555 | [9] |
| LDCM0318 | AC55 | HEK-293T | C281(1.08) | LDD1557 | [9] |
| LDCM0325 | AC61 | HEK-293T | C281(1.10) | LDD1564 | [9] |
| LDCM0327 | AC63 | HEK-293T | C281(1.10) | LDD1566 | [9] |
| LDCM0334 | AC7 | HEK-293T | C281(1.23) | LDD1568 | [9] |
| LDCM0248 | AKOS034007472 | HEK-293T | C281(1.04) | LDD1511 | [9] |
| LDCM0156 | Aniline | NCI-H1299 | 12.00 | LDD0403 | [1] |
| LDCM0369 | CL100 | HEK-293T | C281(0.90) | LDD1573 | [9] |
| LDCM0372 | CL103 | HEK-293T | C281(0.98) | LDD1576 | [9] |
| LDCM0373 | CL104 | HEK-293T | C281(0.97) | LDD1577 | [9] |
| LDCM0376 | CL107 | HEK-293T | C281(0.97) | LDD1580 | [9] |
| LDCM0377 | CL108 | HEK-293T | C281(0.99) | LDD1581 | [9] |
| LDCM0379 | CL11 | HEK-293T | C281(1.19) | LDD1583 | [9] |
| LDCM0381 | CL111 | HEK-293T | C281(0.90) | LDD1585 | [9] |
| LDCM0382 | CL112 | HEK-293T | C281(0.94) | LDD1586 | [9] |
| LDCM0385 | CL115 | HEK-293T | C281(1.09) | LDD1589 | [9] |
| LDCM0386 | CL116 | HEK-293T | C281(1.00) | LDD1590 | [9] |
| LDCM0389 | CL119 | HEK-293T | C281(1.08) | LDD1593 | [9] |
| LDCM0391 | CL120 | HEK-293T | C281(1.01) | LDD1595 | [9] |
| LDCM0394 | CL123 | HEK-293T | C281(0.85) | LDD1598 | [9] |
| LDCM0395 | CL124 | HEK-293T | C281(0.96) | LDD1599 | [9] |
| LDCM0398 | CL127 | HEK-293T | C281(1.02) | LDD1602 | [9] |
| LDCM0399 | CL128 | HEK-293T | C281(1.00) | LDD1603 | [9] |
| LDCM0402 | CL15 | HEK-293T | C281(0.97) | LDD1606 | [9] |
| LDCM0403 | CL16 | HEK-293T | C281(0.98) | LDD1607 | [9] |
| LDCM0409 | CL21 | HEK-293T | C281(1.05) | LDD1613 | [9] |
| LDCM0411 | CL23 | HEK-293T | C281(1.19) | LDD1615 | [9] |
| LDCM0415 | CL27 | HEK-293T | C281(1.00) | LDD1619 | [9] |
| LDCM0416 | CL28 | HEK-293T | C281(0.87) | LDD1620 | [9] |
| LDCM0418 | CL3 | HEK-293T | C281(1.09) | LDD1622 | [9] |
| LDCM0422 | CL33 | HEK-293T | C281(0.99) | LDD1626 | [9] |
| LDCM0424 | CL35 | HEK-293T | C281(1.15) | LDD1628 | [9] |
| LDCM0428 | CL39 | HEK-293T | C281(1.01) | LDD1632 | [9] |
| LDCM0429 | CL4 | HEK-293T | C281(0.97) | LDD1633 | [9] |
| LDCM0430 | CL40 | HEK-293T | C281(0.94) | LDD1634 | [9] |
| LDCM0435 | CL45 | HEK-293T | C281(1.00) | LDD1639 | [9] |
| LDCM0437 | CL47 | HEK-293T | C281(1.26) | LDD1641 | [9] |
| LDCM0443 | CL52 | HEK-293T | C281(1.03) | LDD1646 | [9] |
| LDCM0448 | CL57 | HEK-293T | C281(1.09) | LDD1651 | [9] |
| LDCM0450 | CL59 | HEK-293T | C281(1.01) | LDD1653 | [9] |
| LDCM0455 | CL63 | HEK-293T | C281(0.99) | LDD1658 | [9] |
| LDCM0456 | CL64 | HEK-293T | C281(0.97) | LDD1659 | [9] |
| LDCM0461 | CL69 | HEK-293T | C281(1.02) | LDD1664 | [9] |
| LDCM0464 | CL71 | HEK-293T | C281(1.03) | LDD1667 | [9] |
| LDCM0469 | CL76 | HEK-293T | C281(0.99) | LDD1672 | [9] |
| LDCM0475 | CL81 | HEK-293T | C281(1.21) | LDD1678 | [9] |
| LDCM0477 | CL83 | HEK-293T | C281(1.08) | LDD1680 | [9] |
| LDCM0481 | CL87 | HEK-293T | C281(1.03) | LDD1684 | [9] |
| LDCM0482 | CL88 | HEK-293T | C281(0.99) | LDD1685 | [9] |
| LDCM0484 | CL9 | HEK-293T | C281(1.21) | LDD1687 | [9] |
| LDCM0488 | CL93 | HEK-293T | C281(1.26) | LDD1691 | [9] |
| LDCM0490 | CL95 | HEK-293T | C281(0.96) | LDD1693 | [9] |
| LDCM0494 | CL99 | HEK-293T | C281(0.99) | LDD1697 | [9] |
| LDCM0495 | E2913 | HEK-293T | C281(0.95) | LDD1698 | [9] |
| LDCM0468 | Fragment33 | HEK-293T | C281(1.00) | LDD1671 | [9] |
| LDCM0022 | KB02 | 786-O | C306(2.19) | LDD2247 | [2] |
| LDCM0023 | KB03 | 786-O | C306(2.73) | LDD2664 | [2] |
| LDCM0024 | KB05 | IGR37 | C306(3.62) | LDD3314 | [2] |
| LDCM0109 | NEM | HeLa | N.A. | LDD0224 | [4] |
| LDCM0516 | Nucleophilic fragment 21a | MDA-MB-231 | C68(0.51) | LDD2109 | [3] |
| LDCM0534 | Nucleophilic fragment 30a | MDA-MB-231 | C68(0.93) | LDD2127 | [3] |
| LDCM0543 | Nucleophilic fragment 38 | MDA-MB-231 | C68(1.14) | LDD2136 | [3] |
| LDCM0549 | Nucleophilic fragment 43 | MDA-MB-231 | C68(0.99) | LDD2143 | [3] |
References
