Details of the Target
General Information of Target
| Target ID | LDTP10999 | |||||
|---|---|---|---|---|---|---|
| Target Name | Histone H2A type 1-J (H2AC14) | |||||
| Gene Name | H2AC14 | |||||
| Gene ID | 8331 | |||||
| Synonyms |
H2AFE; HIST1H2AJ; Histone H2A type 1-J; Histone H2A/e |
|||||
| 3D Structure | ||||||
| Sequence |
MPEPSKSAPAPKKGSKKAVTKAQKKDGKKRKRSRKESYSVYVYKVLKQVHPDTGISSKAM
GIMNSFVNDIFERIAGEASRLAHYNKRSTITSREIQTAVRLLLPGELAKHAVSEGTKAVT KYTSSK |
|||||
| Target Bioclass |
Other
|
|||||
| Family |
Histone H2A family
|
|||||
| Subcellular location |
Nucleus
|
|||||
| Function |
Core component of nucleosome. Nucleosomes wrap and compact DNA into chromatin, limiting DNA accessibility to the cellular machineries which require DNA as a template. Histones thereby play a central role in transcription regulation, DNA repair, DNA replication and chromosomal stability. DNA accessibility is regulated via a complex set of post-translational modifications of histones, also called histone code, and nucleosome remodeling.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
P1 Probe Info |
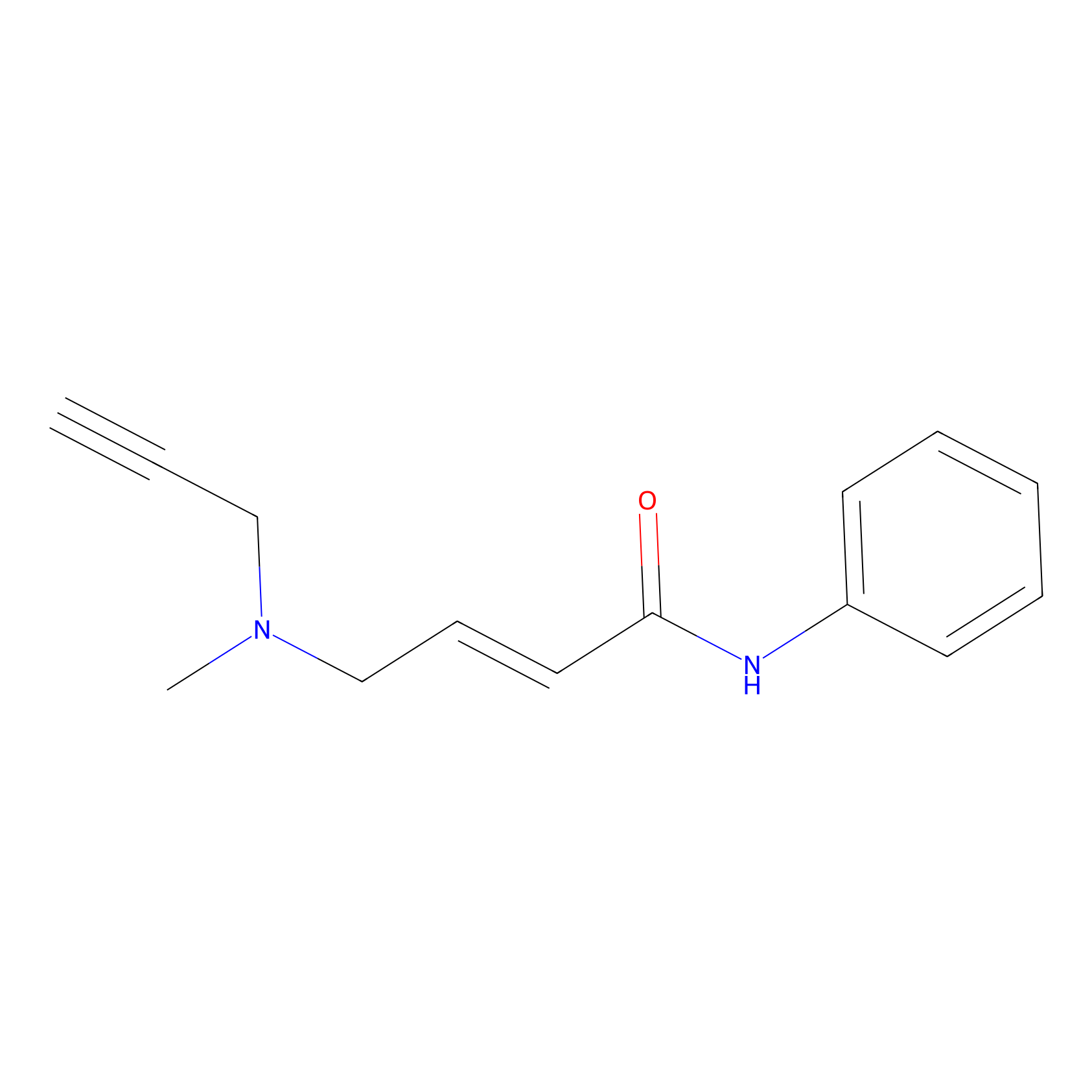 |
2.89 | LDD0448 | [1] | |
|
P2 Probe Info |
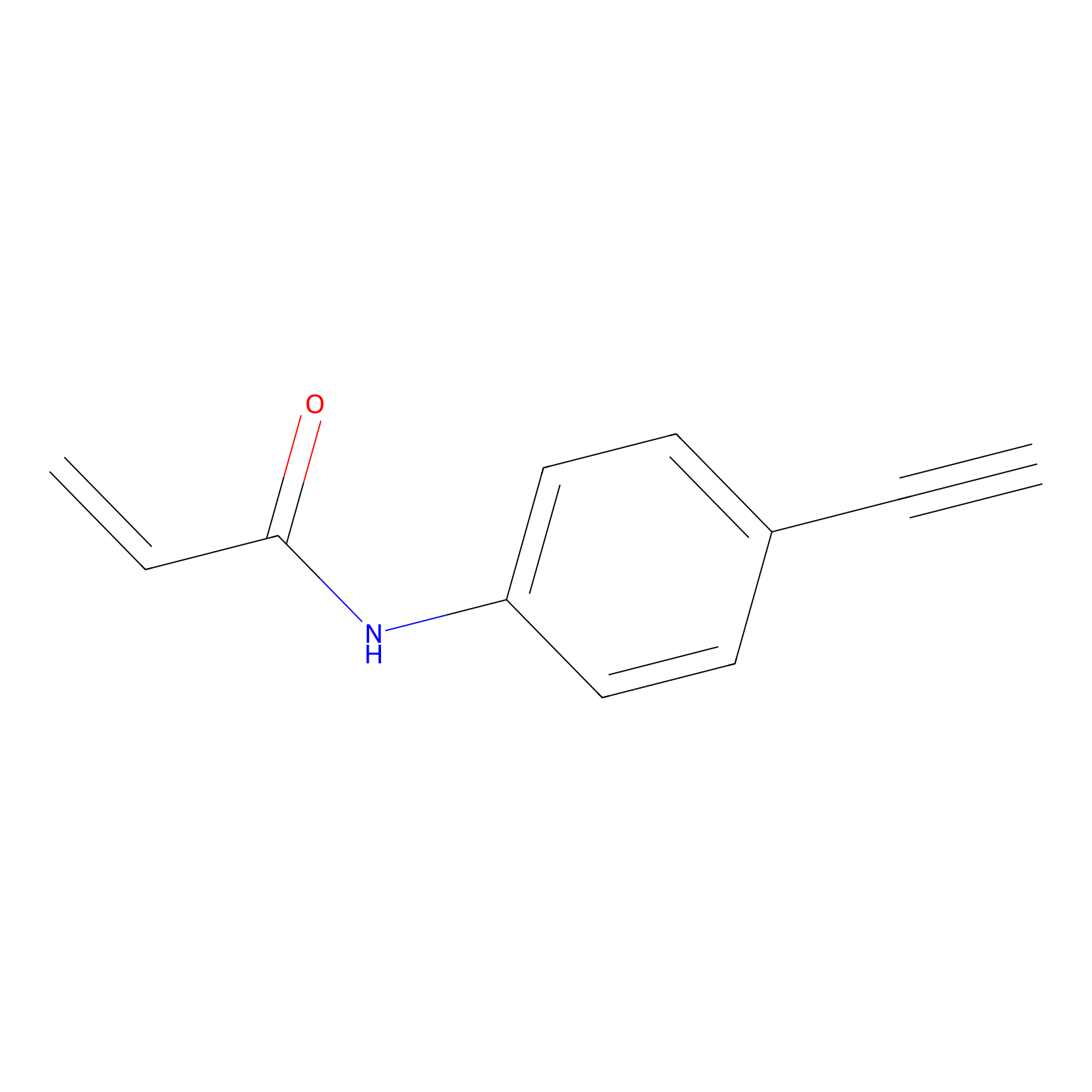 |
3.03 | LDD0449 | [1] | |
|
P8 Probe Info |
 |
1.99 | LDD0451 | [1] | |
|
FBPP2 Probe Info |
 |
2.50 | LDD0318 | [2] | |
|
YN-1 Probe Info |
 |
100.00 | LDD0444 | [3] | |
|
YN-4 Probe Info |
 |
100.00 | LDD0445 | [3] | |
|
AZ-9 Probe Info |
 |
E92(1.09) | LDD2208 | [4] | |
|
ATP probe Probe Info |
 |
K96(0.00); K119(0.00); K120(0.00) | LDD0199 | [5] | |
|
W1 Probe Info |
 |
K120(0.00); K96(0.00); K119(0.00); H83(0.00) | LDD0236 | [6] | |
Competitor(s) Related to This Target
References
