Details of the Target
General Information of Target
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
P3 Probe Info |
 |
5.81 | LDD0450 | [1] | |
|
P8 Probe Info |
 |
1.98 | LDD0451 | [1] | |
|
AZ-5 Probe Info |
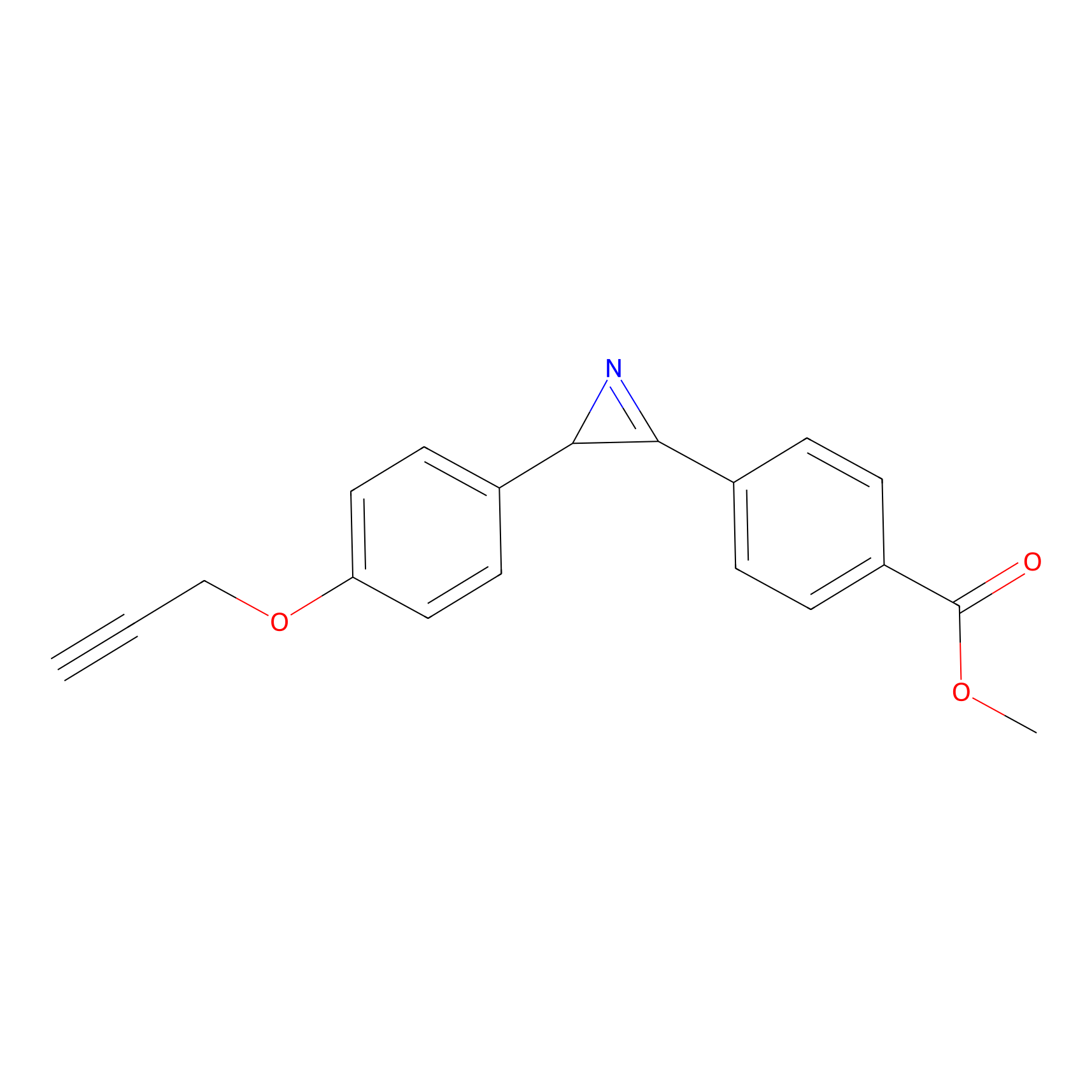 |
5.04 | LDD0394 | [2] | |
|
YN-1 Probe Info |
 |
100.00 | LDD0444 | [3] | |
|
DBIA Probe Info |
 |
C336(4.59); C40(2.42) | LDD3313 | [4] | |
|
Jackson_14 Probe Info |
 |
8.33 | LDD0123 | [5] | |
|
W1 Probe Info |
 |
C40(1.30) | LDD0239 | [6] | |
|
m-APA Probe Info |
 |
H428(0.00); H394(0.00) | LDD2231 | [7] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0149 | [8] | |
|
IPM Probe Info |
 |
C40(0.00); C358(0.00) | LDD0147 | [9] | |
|
TFBX Probe Info |
 |
N.A. | LDD0148 | [9] | |
|
Acrolein Probe Info |
 |
H394(0.00); H222(0.00); C40(0.00); C358(0.00) | LDD0217 | [10] | |
|
Cinnamaldehyde Probe Info |
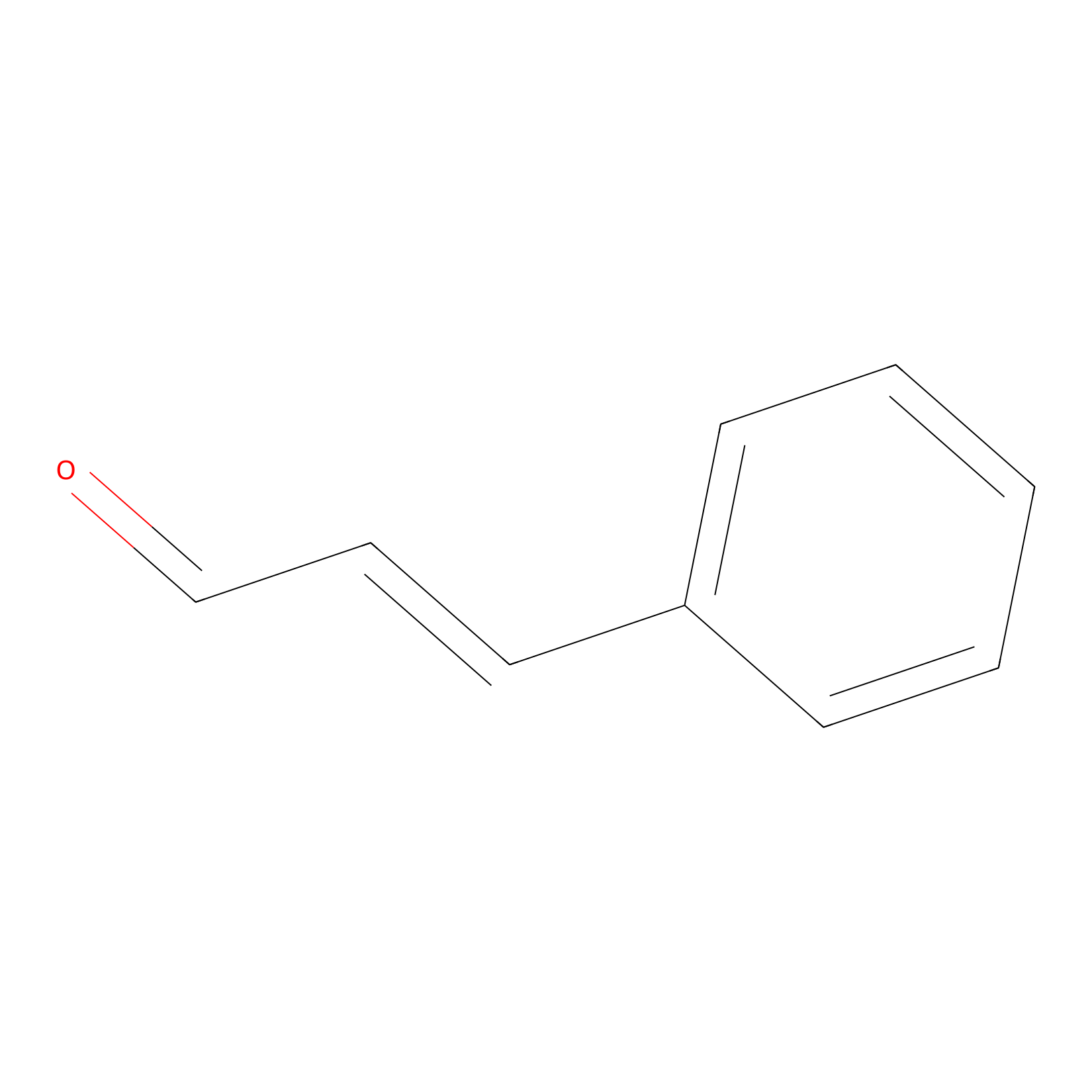 |
C40(0.00); C358(0.00) | LDD0220 | [10] | |
|
Crotonaldehyde Probe Info |
 |
C40(0.00); C358(0.00); H428(0.00); H394(0.00) | LDD0219 | [10] | |
|
Methacrolein Probe Info |
 |
H222(0.00); H394(0.00); C358(0.00); C40(0.00) | LDD0218 | [10] | |
|
HHS-465 Probe Info |
 |
Y398(0.00); Y69(0.00); Y61(0.00); K399(0.00) | LDD2240 | [11] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
STS-1 Probe Info |
 |
1.14 | LDD0136 | [12] | |
|
STS-2 Probe Info |
 |
1.05 | LDD0138 | [12] | |
|
Dasatinib-CA-8PAP Probe Info |
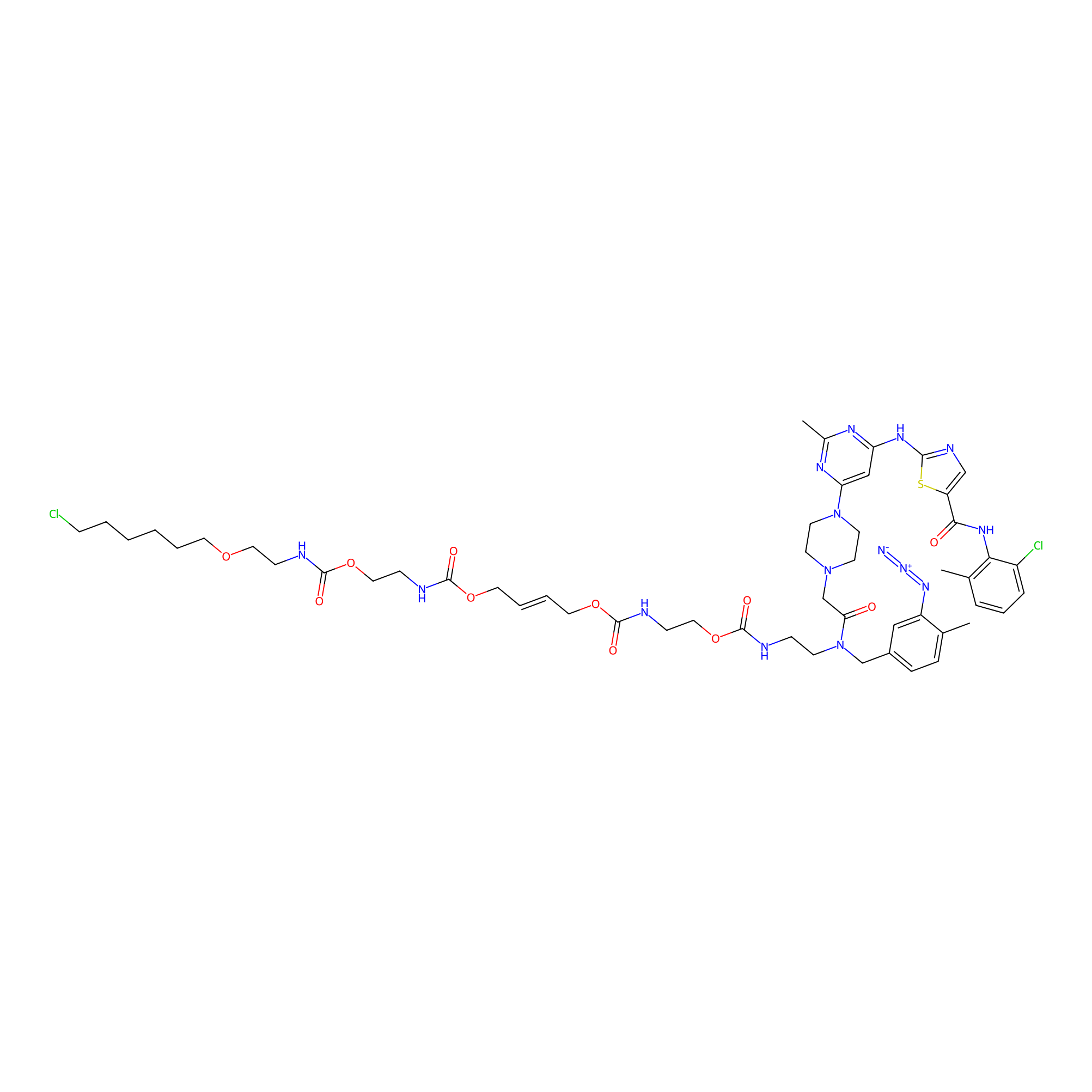 |
N.A. | LDD0364 | [13] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0108 | Chloroacetamide | HeLa | H222(0.00); C40(0.00); C358(0.00); H394(0.00) | LDD0222 | [10] |
| LDCM0097 | Dasatinib | K562 | N.A. | LDD0364 | [13] |
| LDCM0107 | IAA | HeLa | H394(0.00); H222(0.00); C40(0.00); H428(0.00) | LDD0221 | [10] |
| LDCM0022 | KB02 | 8305C | C389(1.89); C40(2.16) | LDD2248 | [4] |
| LDCM0023 | KB03 | 8305C | C389(4.48) | LDD2665 | [4] |
| LDCM0024 | KB05 | Hs 936.T | C336(4.59); C40(2.42) | LDD3313 | [4] |
| LDCM0109 | NEM | HeLa | H394(0.00); H222(0.00); C40(0.00); H428(0.00) | LDD0223 | [10] |
| LDCM0016 | Ranjitkar_cp1 | MDA-MB-231 | 8.33 | LDD0123 | [5] |
| LDCM0112 | W16 | Hep-G2 | C40(1.30) | LDD0239 | [6] |
The Interaction Atlas With This Target
References
