Details of the Target
General Information of Target
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
AZ-9 Probe Info |
 |
3.29 | LDD0393 | [1] | |
|
FBPP2 Probe Info |
 |
3.48 | LDD0318 | [2] | |
|
CHEMBL5175495 Probe Info |
 |
7.48 | LDD0196 | [3] | |
|
TG42 Probe Info |
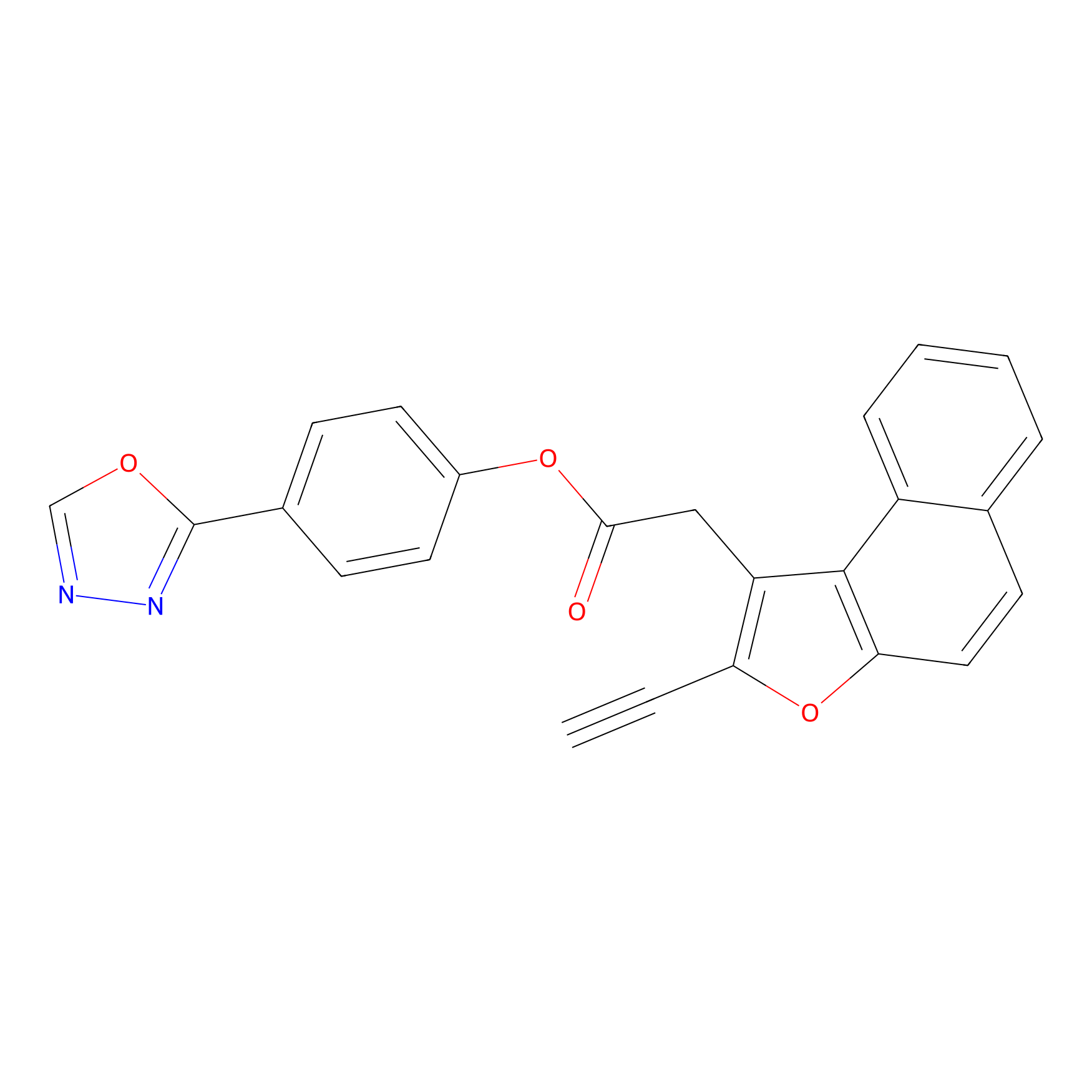 |
7.24 | LDD0326 | [4] | |
|
FBP2 Probe Info |
 |
2.45 | LDD0317 | [2] | |
|
m-APA Probe Info |
 |
15.00 | LDD0403 | [5] | |
|
Jackson_14 Probe Info |
 |
2.04 | LDD0123 | [6] | |
|
AMP probe Probe Info |
 |
K152(0.00); K153(0.00) | LDD0200 | [7] | |
|
ATP probe Probe Info |
 |
K152(0.00); K153(0.00); K165(0.00); K11(0.00) | LDD0199 | [7] | |
|
ATP probe Probe Info |
 |
K152(0.00); K153(0.00) | LDD0035 | [8] | |
|
IPM Probe Info |
 |
N.A. | LDD0025 | [9] | |
|
NHS Probe Info |
 |
K152(0.00); K11(0.00); K165(0.00); K153(0.00) | LDD0010 | [10] | |
|
SF Probe Info |
 |
N.A. | LDD0028 | [11] | |
|
STPyne Probe Info |
 |
K30(0.00); K11(0.00) | LDD0009 | [10] | |
|
VSF Probe Info |
 |
C321(0.00); C279(0.00) | LDD0007 | [10] | |
|
Phosphinate-6 Probe Info |
 |
N.A. | LDD0018 | [12] | |
|
1c-yne Probe Info |
 |
K175(0.00); K40(0.00); K11(0.00); K69(0.00) | LDD0228 | [13] | |
|
AOyne Probe Info |
 |
15.00 | LDD0443 | [14] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
STS-1 Probe Info |
 |
N.A. | LDD0068 | [15] | |
Competitor(s) Related to This Target
The Interaction Atlas With This Target
References
