Details of the Target
General Information of Target
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C-Sul Probe Info |
 |
2.70 | LDD0066 | [1] | |
|
ONAyne Probe Info |
 |
K118(2.56) | LDD0274 | [2] | |
|
AZ-9 Probe Info |
 |
D73(0.95); E43(10.00); E75(10.00) | LDD2208 | [3] | |
|
ATP probe Probe Info |
 |
N.A. | LDD0199 | [4] | |
|
aONE Probe Info |
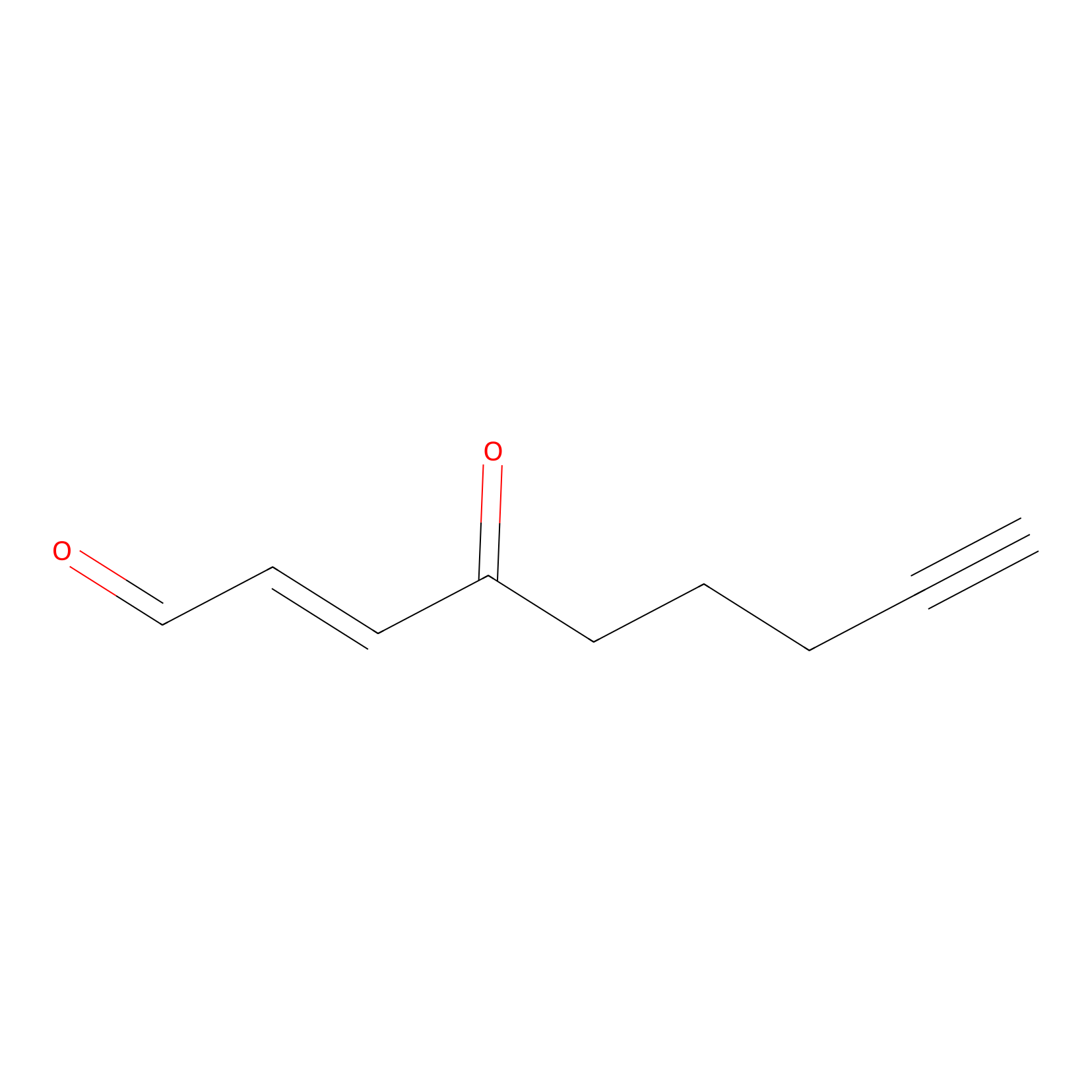 |
N.A. | LDD0002 | [5] | |
|
NHS Probe Info |
 |
K47(0.00); K107(0.00); K17(0.00); K35(0.00) | LDD0010 | [5] | |
|
OSF Probe Info |
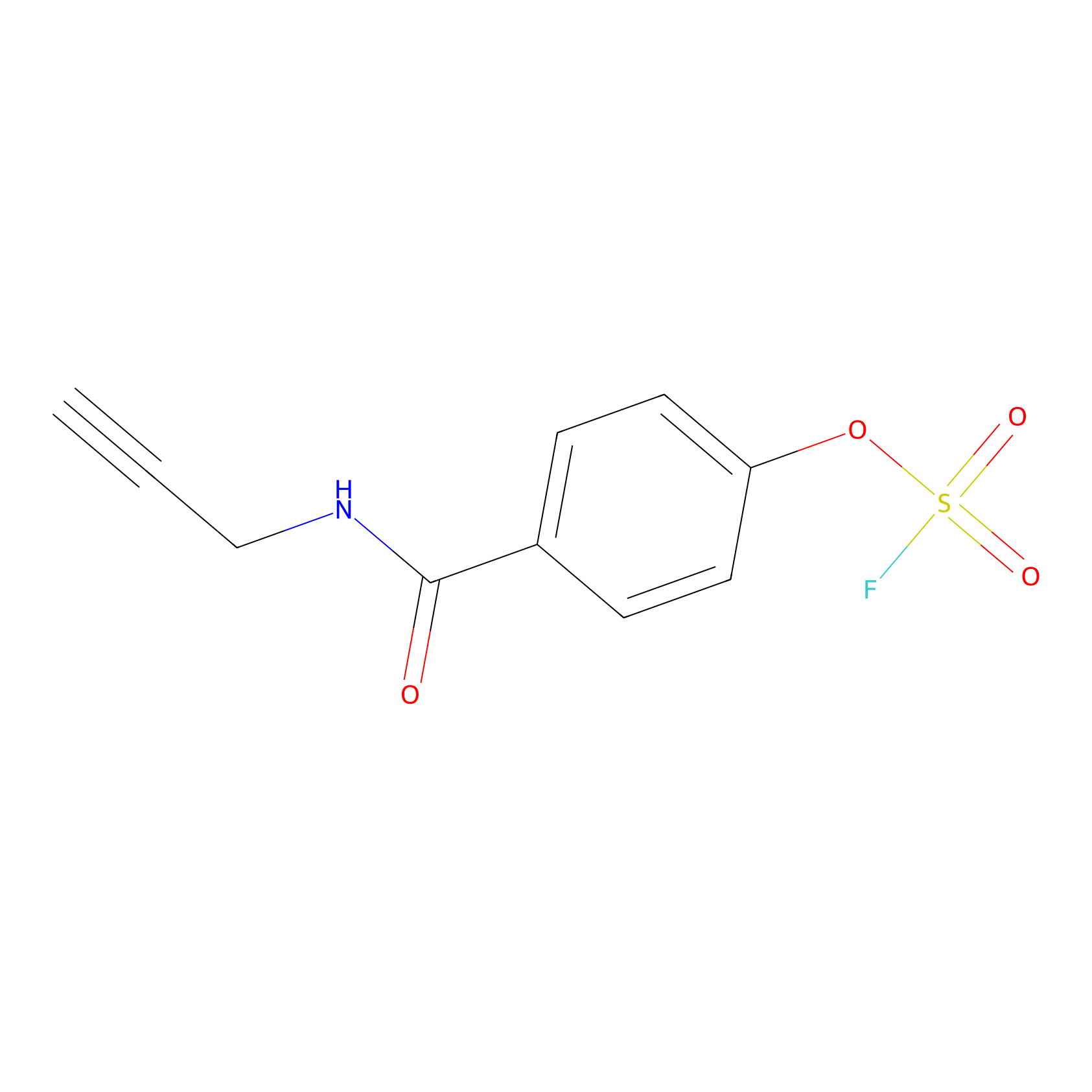 |
N.A. | LDD0029 | [6] | |
|
SF Probe Info |
 |
K65(0.00); K131(0.00); K91(0.00); K76(0.00) | LDD0028 | [6] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
FFF probe12 Probe Info |
 |
6.26 | LDD0473 | [7] | |
|
Photocelecoxib Probe Info |
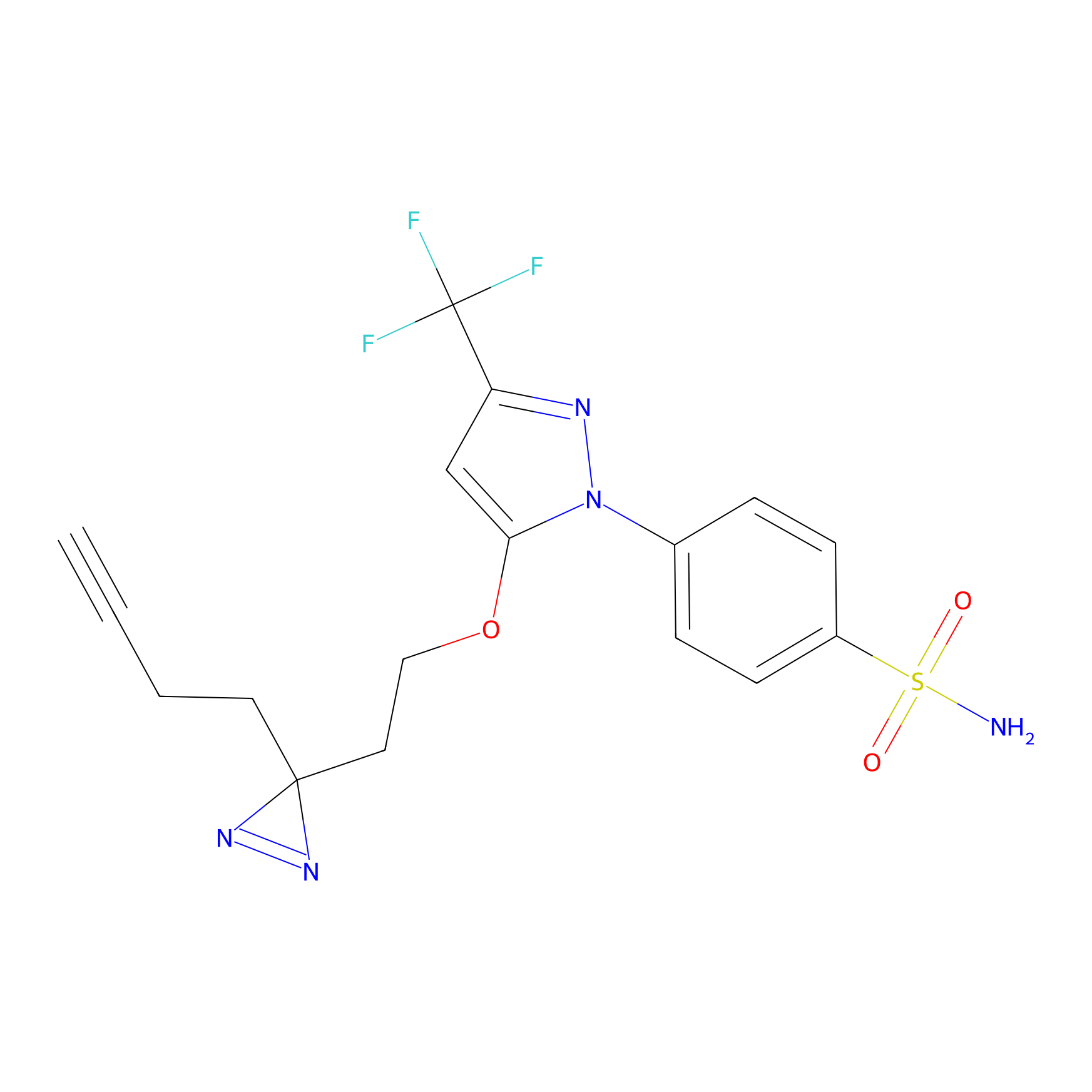 |
V74(0.00); E75(0.00) | LDD0153 | [8] | |
|
Photonaproxen Probe Info |
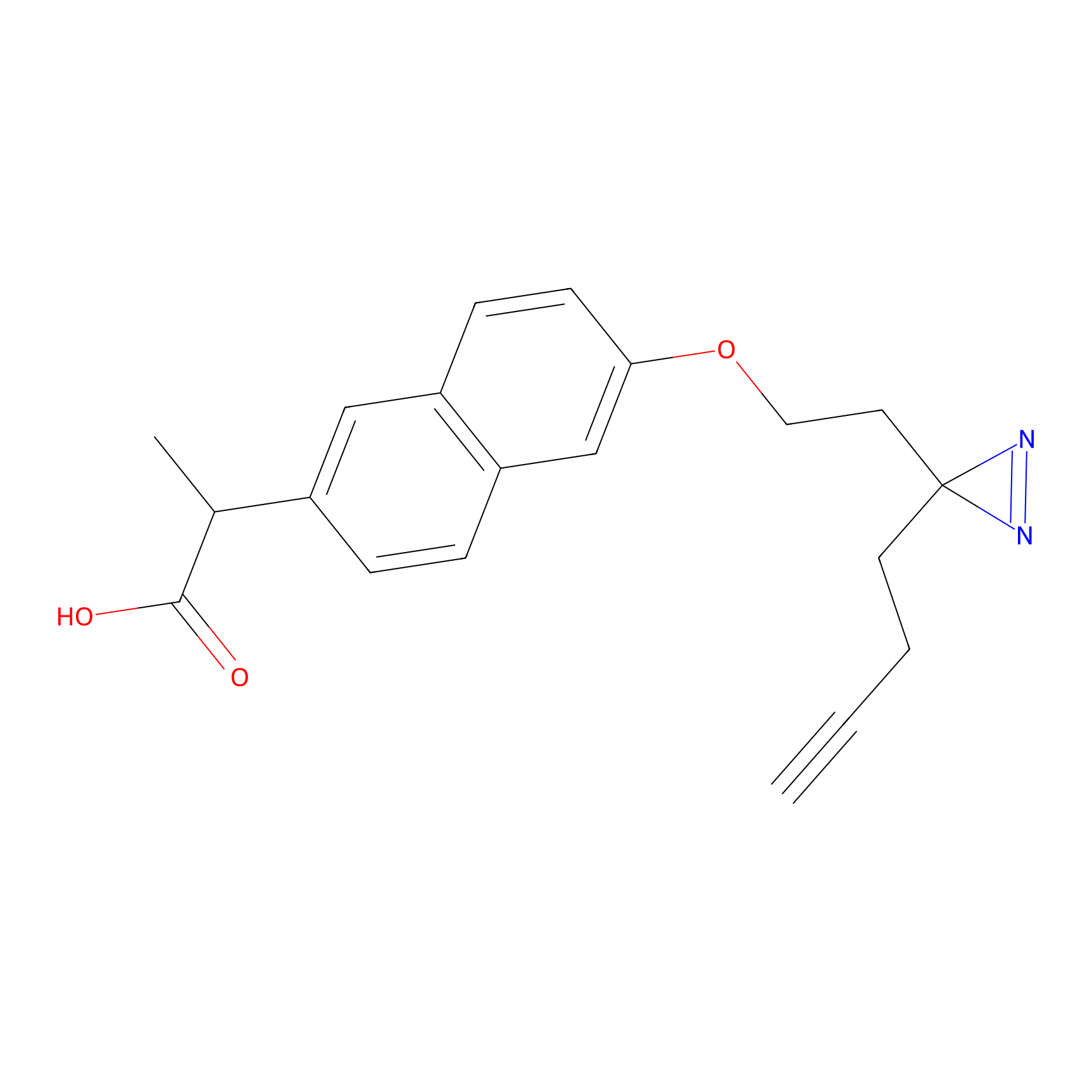 |
N.A. | LDD0157 | [8] | |
References
