Details of the Target
General Information of Target
| Target ID | LDTP01067 | |||||
|---|---|---|---|---|---|---|
| Target Name | Endothelial differentiation-related factor 1 (EDF1) | |||||
| Gene Name | EDF1 | |||||
| Gene ID | 8721 | |||||
| Synonyms |
Endothelial differentiation-related factor 1; EDF-1; Multiprotein-bridging factor 1; MBF1 |
|||||
| 3D Structure | ||||||
| Sequence |
MAESDWDTVTVLRKKGPTAAQAKSKQAILAAQRRGEDVETSKKWAAGQNKQHSITKNTAK
LDRETEELHHDRVTLEVGKVIQQGRQSKGLTQKDLATKINEKPQVIADYESGRAIPNNQV LGKIERAIGLKLRGKDIGKPIEKGPRAK |
|||||
| Target Bioclass |
Other
|
|||||
| Subcellular location |
Cytoplasm
|
|||||
| Function |
Transcriptional coactivator stimulating NR5A1 and ligand-dependent NR1H3/LXRA and PPARG transcriptional activities. Enhances the DNA-binding activity of ATF1, ATF2, CREB1 and NR5A1. Regulates nitric oxid synthase activity probably by sequestering calmodulin in the cytoplasm. May function in endothelial cells differentiation, hormone-induced cardiomyocytes hypertrophy and lipid metabolism.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Target Site Mutations in Different Cell Lines
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C-Sul Probe Info |
 |
9.91 | LDD0066 | [1] | |
|
ONAyne Probe Info |
 |
K131(10.00) | LDD0274 | [2] | |
|
STPyne Probe Info |
 |
K102(1.69); K131(7.69); K135(5.40); K139(10.00) | LDD0277 | [2] | |
|
AMP probe Probe Info |
 |
N.A. | LDD0200 | [3] | |
|
ATP probe Probe Info |
 |
K15(0.00); K50(0.00); K56(0.00); K79(0.00) | LDD0199 | [3] | |
|
m-APA Probe Info |
 |
N.A. | LDD2232 | [4] | |
|
ATP probe Probe Info |
 |
N.A. | LDD0035 | [5] | |
|
1d-yne Probe Info |
 |
N.A. | LDD0356 | [6] | |
|
SF Probe Info |
 |
K25(0.00); K143(0.00) | LDD0028 | [7] | |
|
Ox-W18 Probe Info |
 |
W6(0.00); W44(0.00) | LDD2175 | [8] | |
|
1c-yne Probe Info |
 |
K98(0.00); K102(0.00) | LDD0228 | [6] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0217 | [9] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C003 Probe Info |
 |
33.36 | LDD1713 | [10] | |
|
C052 Probe Info |
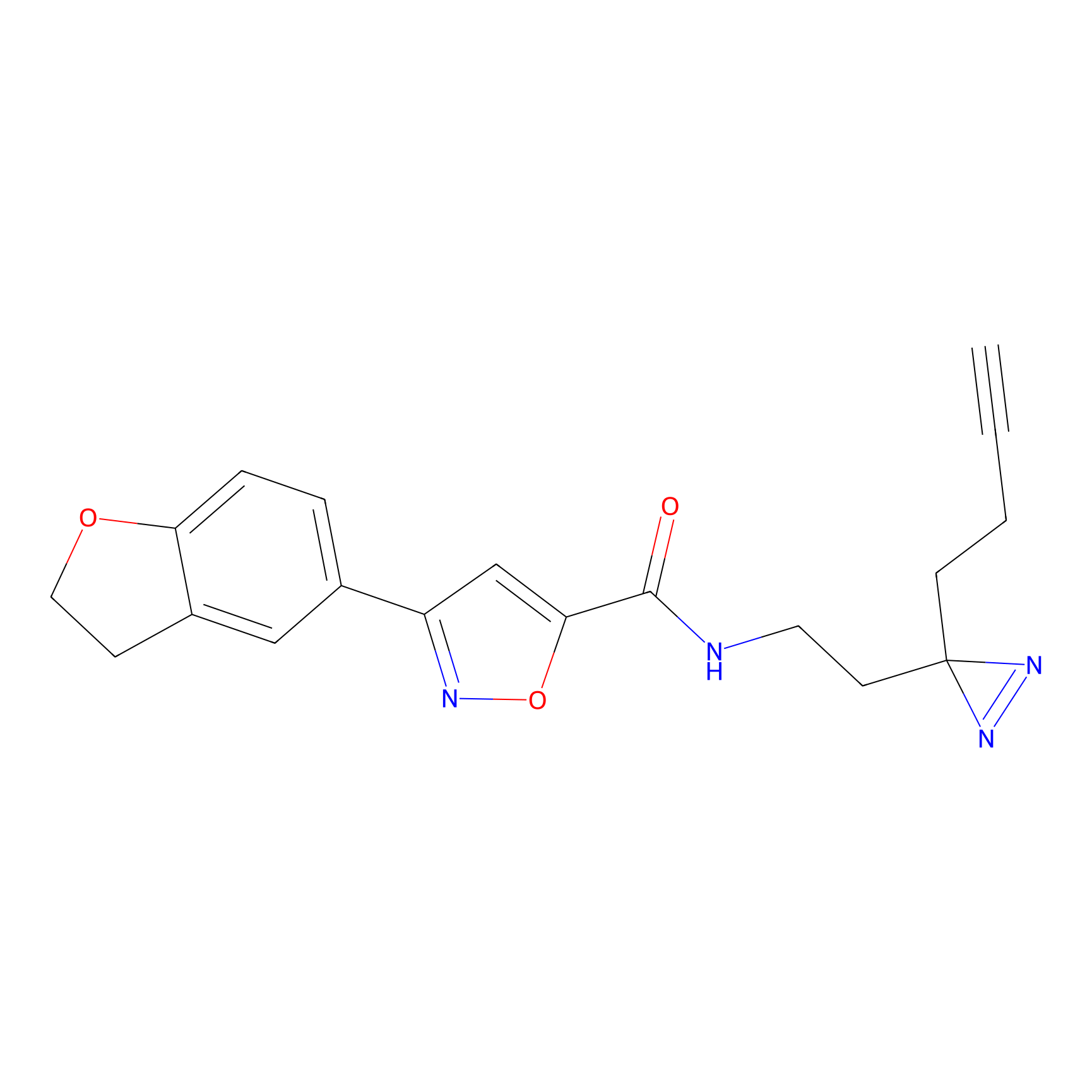 |
5.35 | LDD1750 | [10] | |
|
C087 Probe Info |
 |
8.75 | LDD1779 | [10] | |
|
C189 Probe Info |
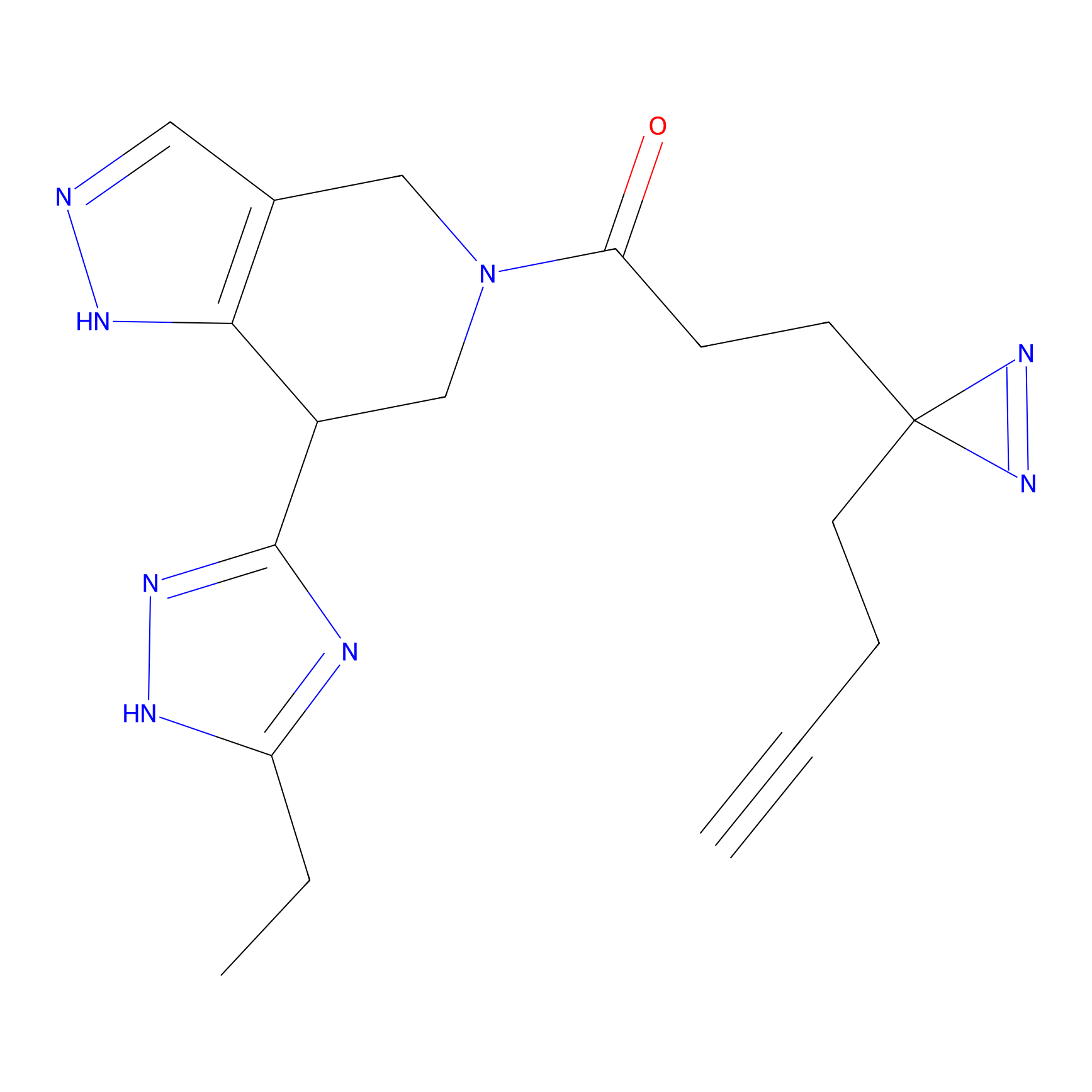 |
5.62 | LDD1866 | [10] | |
|
C362 Probe Info |
 |
27.47 | LDD2023 | [10] | |
|
C364 Probe Info |
 |
15.78 | LDD2025 | [10] | |
Competitor(s) Related to This Target
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Transporter and channel
Transcription factor
References
