Details of the Target
General Information of Target
| Target ID | LDTP09505 | |||||
|---|---|---|---|---|---|---|
| Target Name | GPI ethanolamine phosphate transferase 3 (PIGO) | |||||
| Gene Name | PIGO | |||||
| Gene ID | 84720 | |||||
| Synonyms |
GPI ethanolamine phosphate transferase 3; EC 2.-.-.-; Phosphatidylinositol-glycan biosynthesis class O protein; PIG-O |
|||||
| 3D Structure | ||||||
| Sequence |
MQKASVLLFLAWVCFLFYAGIALFTSGFLLTRLELTNHSSCQEPPGPGSLPWGSQGKPGA
CWMASRFSRVVLVLIDALRFDFAQPQHSHVPREPPVSLPFLGKLSSLQRILEIQPHHARL YRSQVDPPTTTMQRLKALTTGSLPTFIDAGSNFASHAIVEDNLIKQLTSAGRRVVFMGDD TWKDLFPGAFSKAFFFPSFNVRDLDTVDNGILEHLYPTMDSGEWDVLIAHFLGVDHCGHK HGPHHPEMAKKLSQMDQVIQGLVERLENDTLLVVAGDHGMTTNGDHGGDSELEVSAALFL YSPTAVFPSTPPEEPEVIPQVSLVPTLALLLGLPIPFGNIGEVMAELFSGGEDSQPHSSA LAQASALHLNAQQVSRFLHTYSAATQDLQAKELHQLQNLFSKASADYQWLLQSPKGAEAT LPTVIAELQQFLRGARAMCIESWARFSLVRMAGGTALLAASCFICLLASQWAISPGFPFC PLLLTPVAWGLVGAIAYAGLLGTIELKLDLVLLGAVAAVSSFLPFLWKAWAGWGSKRPLA TLFPIPGPVLLLLLFRLAVFFSDSFVVAEARATPFLLGSFILLLVVQLHWEGQLLPPKLL TMPRLGTSATTNPPRHNGAYALRLGIGLLLCTRLAGLFHRCPEETPVCHSSPWLSPLASM VGGRAKNLWYGACVAALVALLAAVRLWLRRYGNLKSPEPPMLFVRWGLPLMALGTAAYWA LASGADEAPPRLRVLVSGASMVLPRAVAGLAASGLALLLWKPVTVLVKAGAGAPRTRTVL TPFSGPPTSQADLDYVVPQIYRHMQEEFRGRLERTKSQGPLTVAAYQLGSVYSAAMVTAL TLLAFPLLLLHAERISLVFLLLFLQSFLLLHLLAAGIPVTTPGPFTVPWQAVSAWALMAT QTFYSTGHQPVFPAIHWHAAFVGFPEGHGSCTWLPALLVGANTFASHLLFAVGCPLLLLW PFLCESQGLRKRQQPPGNEADARVRPEEEEEPLMEMRLRDAPQHFYAALLQLGLKYLFIL GIQILACALAASILRRHLMVWKVFAPKFIFEAVGFIVSSVGLLLGIALVMRVDGAVSSWF RQLFLAQQR |
|||||
| Target Bioclass |
Enzyme
|
|||||
| Family |
PIGG/PIGN/PIGO family, PIGO subfamily
|
|||||
| Subcellular location |
Endoplasmic reticulum membrane
|
|||||
| Function |
Ethanolamine phosphate transferase involved in glycosylphosphatidylinositol-anchor biosynthesis. Transfers ethanolamine phosphate to the GPI third mannose which links the GPI-anchor to the C-terminus of the proteins by an amide bond.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Target Site Mutations in Different Cell Lines
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
DBIA Probe Info |
 |
C439(2.33) | LDD3383 | [1] | |
|
Johansson_61 Probe Info |
 |
_(20.00) | LDD1485 | [2] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
A-DA Probe Info |
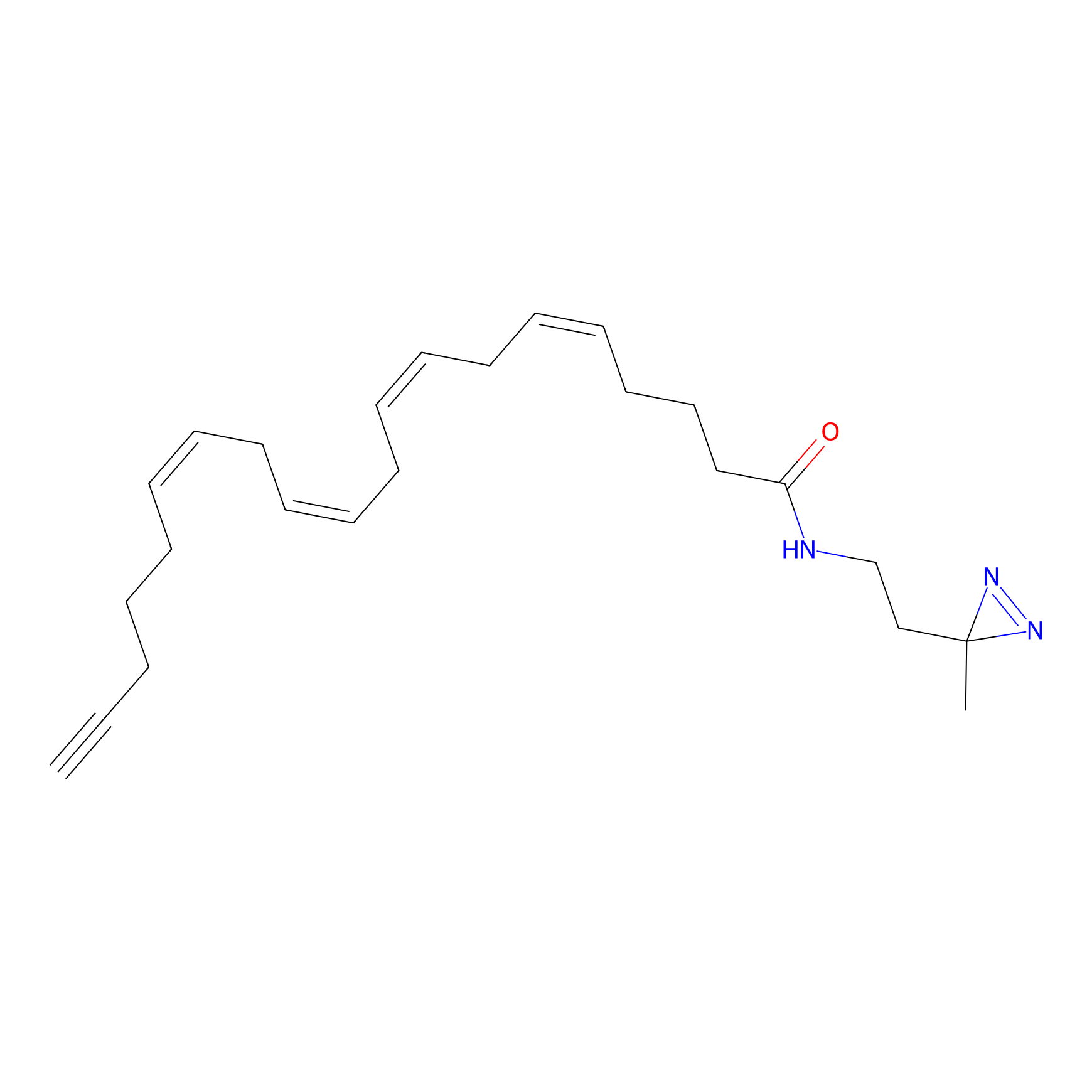 |
2.28 | LDD0145 | [3] | |
|
OEA-DA Probe Info |
 |
9.64 | LDD0046 | [4] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0615 | Fragment63-R | Jurkat | _(20.00) | LDD1487 | [2] |
| LDCM0617 | Fragment63-S | Jurkat | _(20.00) | LDD1490 | [2] |
| LDCM0569 | Fragment7 | Jurkat | _(20.00) | LDD1485 | [2] |
| LDCM0022 | KB02 | A2780 | C439(1.48) | LDD2254 | [1] |
| LDCM0023 | KB03 | A2780 | C439(2.12) | LDD2671 | [1] |
| LDCM0024 | KB05 | OVCAR-5 | C439(2.33) | LDD3383 | [1] |
| LDCM0084 | Ro 48-8071 | A-549 | 2.28 | LDD0145 | [3] |
References
