Details of the Target
General Information of Target
| Target ID | LDTP06885 | |||||
|---|---|---|---|---|---|---|
| Target Name | Glycerol-3-phosphate acyltransferase 3 (GPAT3) | |||||
| Gene Name | GPAT3 | |||||
| Gene ID | 84803 | |||||
| Synonyms |
AGPAT9; MAG1; Glycerol-3-phosphate acyltransferase 3; GPAT-3; EC 2.3.1.15; 1-acyl-sn-glycerol-3-phosphate O-acyltransferase 10; AGPAT 10; 1-acyl-sn-glycerol-3-phosphate O-acyltransferase 9; 1-AGP acyltransferase 9; 1-AGPAT 9; EC 2.3.1.51; Acyl-CoA:glycerol-3-phosphate acyltransferase 3; hGPAT3; Lung cancer metastasis-associated protein 1; Lysophosphatidic acid acyltransferase theta; LPAAT-theta; MAG-1
|
|||||
| 3D Structure | ||||||
| Sequence |
MEGAELAGKILSTWLTLVLGFILLPSVFGVSLGISEIYMKILVKTLEWATIRIEKGTPKE
SILKNSASVGIIQRDESPMEKGLSGLRGRDFELSDVFYFSKKGLEAIVEDEVTQRFSSEE LVSWNLLTRTNVNFQYISLRLTMVWVLGVIVRYCVLLPLRVTLAFIGISLLVIGTTLVGQ LPDSSLKNWLSELVHLTCCRICVRALSGTIHYHNKQYRPQKGGICVANHTSPIDVLILTT DGCYAMVGQVHGGLMGIIQRAMVKACPHVWFERSEMKDRHLVTKRLKEHIADKKKLPILI FPEGTCINNTSVMMFKKGSFEIGGTIHPVAIKYNPQFGDAFWNSSKYNMVSYLLRMMTSW AIVCDVWYMPPMTREEGEDAVQFANRVKSAIAIQGGLTELPWDGGLKRAKVKDIFKEEQQ KNYSKMIVGNGSLS |
|||||
| Target Bioclass |
Enzyme
|
|||||
| Family |
1-acyl-sn-glycerol-3-phosphate acyltransferase family
|
|||||
| Subcellular location |
Endoplasmic reticulum membrane
|
|||||
| Function |
Converts glycerol-3-phosphate to 1-acyl-sn-glycerol-3-phosphate (lysophosphatidic acid or LPA) by incorporating an acyl moiety at the sn-1 position of the glycerol backbone. Also converts LPA into 1,2-diacyl-sn-glycerol-3-phosphate (phosphatidic acid or PA) by incorporating an acyl moiety at the sn-2 position of the glycerol backbone. Protects cells against lipotoxicity.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
14.69 | LDD0402 | [1] | |
|
YN-1 Probe Info |
 |
100.00 | LDD0444 | [2] | |
|
STPyne Probe Info |
 |
K284(7.26); K287(5.43); K293(10.00); K388(1.23) | LDD0277 | [3] | |
|
BTD Probe Info |
 |
C266(1.07) | LDD2104 | [4] | |
|
AHL-Pu-1 Probe Info |
 |
C306(3.06) | LDD0169 | [5] | |
|
EA-probe Probe Info |
 |
C306(1.87) | LDD2210 | [6] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0221 | [7] | |
|
IA-alkyne Probe Info |
 |
C306(0.00); C266(0.00) | LDD0162 | [8] | |
|
IPM Probe Info |
 |
N.A. | LDD2156 | [9] | |
|
Methacrolein Probe Info |
 |
N.A. | LDD0218 | [7] | |
|
AOyne Probe Info |
 |
15.00 | LDD0443 | [10] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
A-DA Probe Info |
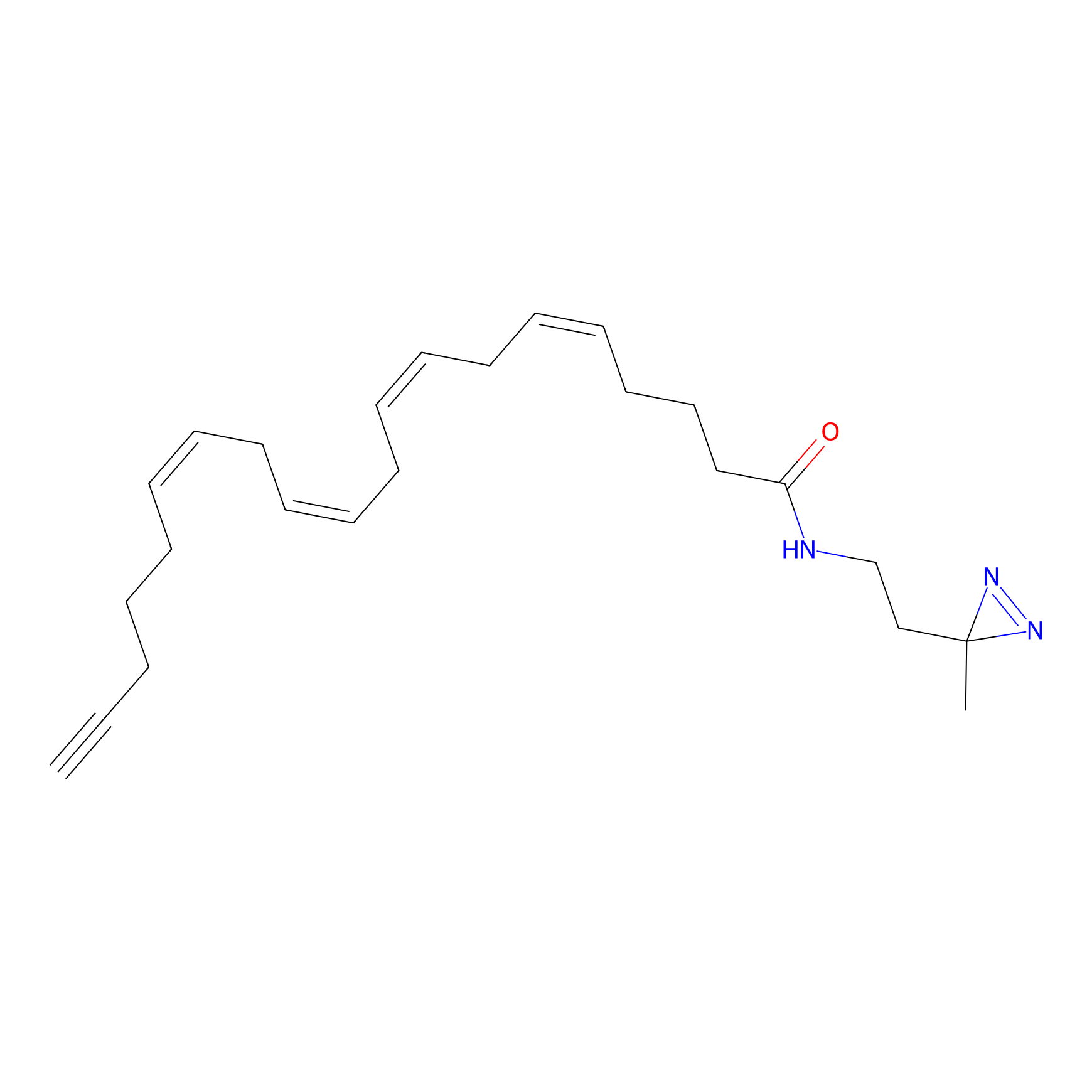 |
2.70 | LDD0143 | [11] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0548 | 1-(4-(Benzo[d][1,3]dioxol-5-ylmethyl)piperazin-1-yl)-2-nitroethan-1-one | MDA-MB-231 | C266(0.69) | LDD2142 | [4] |
| LDCM0026 | 4SU-RNA+native RNA | HEK-293T | C306(3.06) | LDD0169 | [5] |
| LDCM0156 | Aniline | NCI-H1299 | 12.35 | LDD0403 | [1] |
| LDCM0083 | Avasimibe | A-549 | 2.70 | LDD0143 | [11] |
| LDCM0630 | CCW28-3 | 231MFP | C306(2.01) | LDD2214 | [12] |
| LDCM0108 | Chloroacetamide | HeLa | N.A. | LDD0222 | [7] |
| LDCM0175 | Ethacrynic acid | HeLa | C306(1.87) | LDD2210 | [6] |
| LDCM0107 | IAA | HeLa | N.A. | LDD0221 | [7] |
| LDCM0511 | Nucleophilic fragment 18b | MDA-MB-231 | C266(1.07) | LDD2104 | [4] |
| LDCM0627 | NUDT7-COV-1 | HEK-293T | C266(1.42) | LDD2206 | [13] |
| LDCM0628 | OTUB2-COV-1 | HEK-293T | C266(0.91) | LDD2207 | [13] |
| LDCM0084 | Ro 48-8071 | A-549 | 2.93 | LDD0145 | [11] |
References
