Details of the Target
General Information of Target
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| CCK81 | SNV: p.E496G | DBIA Probe Info | |||
| D283MED | SNV: p.P724S | . | |||
| DU145 | SNV: p.M885L | DBIA Probe Info | |||
| HCT116 | Deletion: p.V637CfsTer22 | . | |||
| HEC1 | SNV: p.R643W | DBIA Probe Info | |||
| HEC1B | SNV: p.M97T; p.R643W | . | |||
| HUPT3 | SNV: p.P645A | DBIA Probe Info | |||
| MDAMB436 | SNV: p.C395Y | . | |||
| MFE319 | SNV: p.L384P | . | |||
| U937 | Substitution: p.S9I | DBIA Probe Info | |||
| WM793 | SNV: p.K801M | . | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
A-EBA Probe Info |
 |
50.00 | LDD0214 | [1] | |
|
DBIA Probe Info |
 |
C89(2.35) | LDD3310 | [2] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0221 | [3] | |
|
CY-1 Probe Info |
 |
D369(0.00); E371(0.00); K372(0.00); Q378(0.00) | LDD0246 | [4] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
C677(0.00); C463(0.00); C89(0.00); C352(0.00) | LDD0038 | [5] | |
|
IA-alkyne Probe Info |
 |
C661(0.00); C463(0.00); C677(0.00); C89(0.00) | LDD0036 | [5] | |
|
Lodoacetamide azide Probe Info |
 |
C61(0.00); C661(0.00); C677(0.00); C89(0.00) | LDD0037 | [5] | |
|
KY-26 Probe Info |
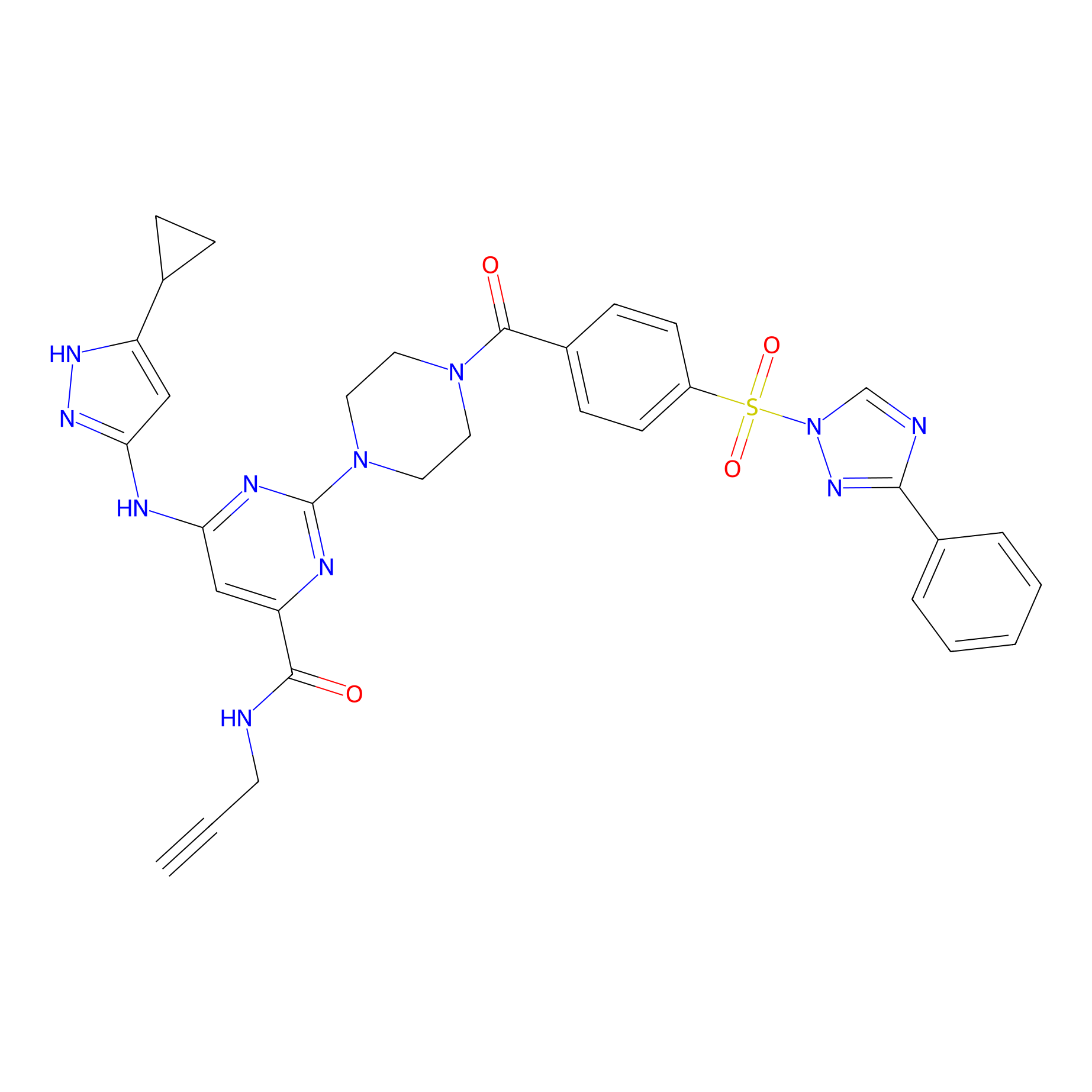 |
N.A. | LDD0301 | [6] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
DFG-out-3 Probe Info |
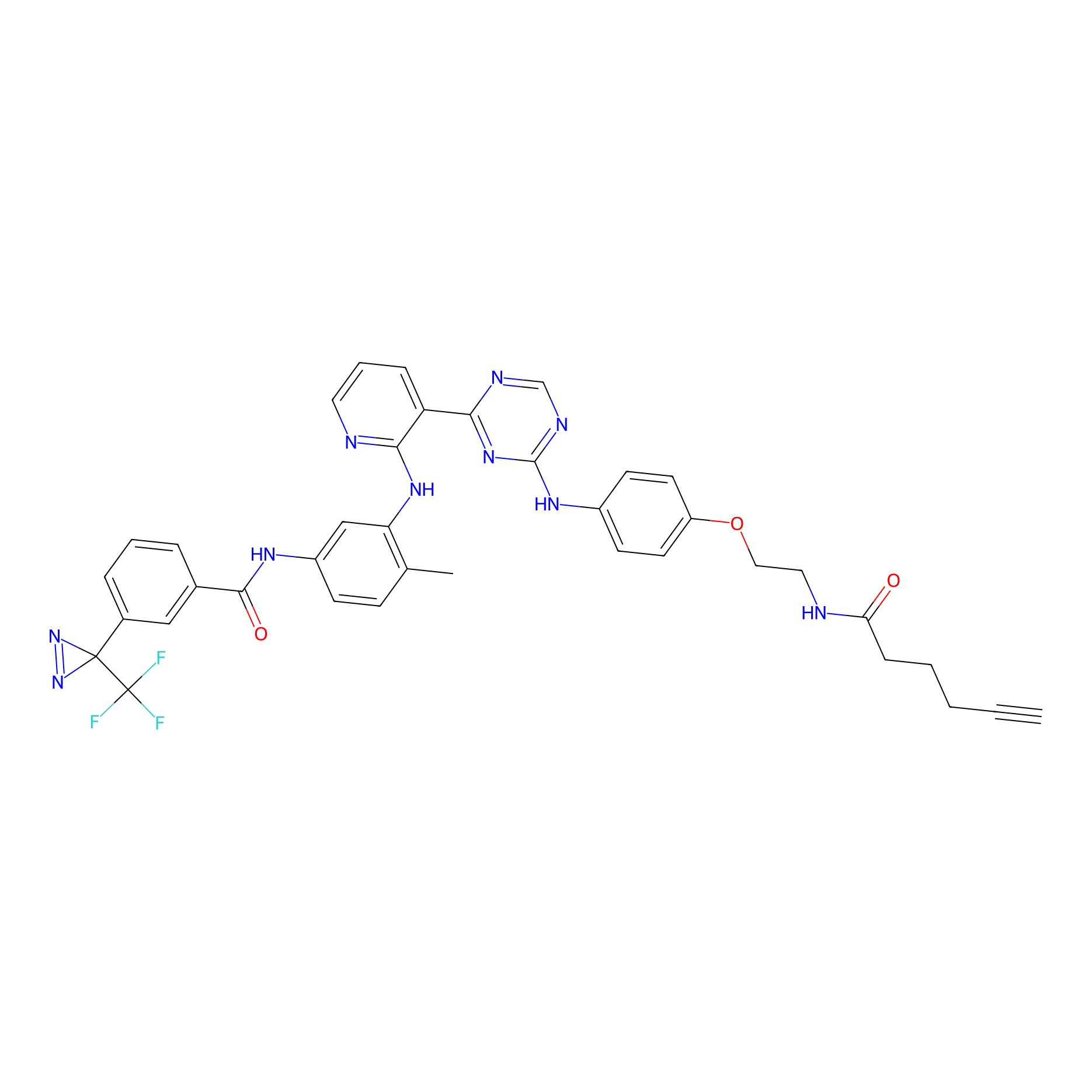 |
2.70 | LDD0074 | [7] | |
|
DFG-out-4 Probe Info |
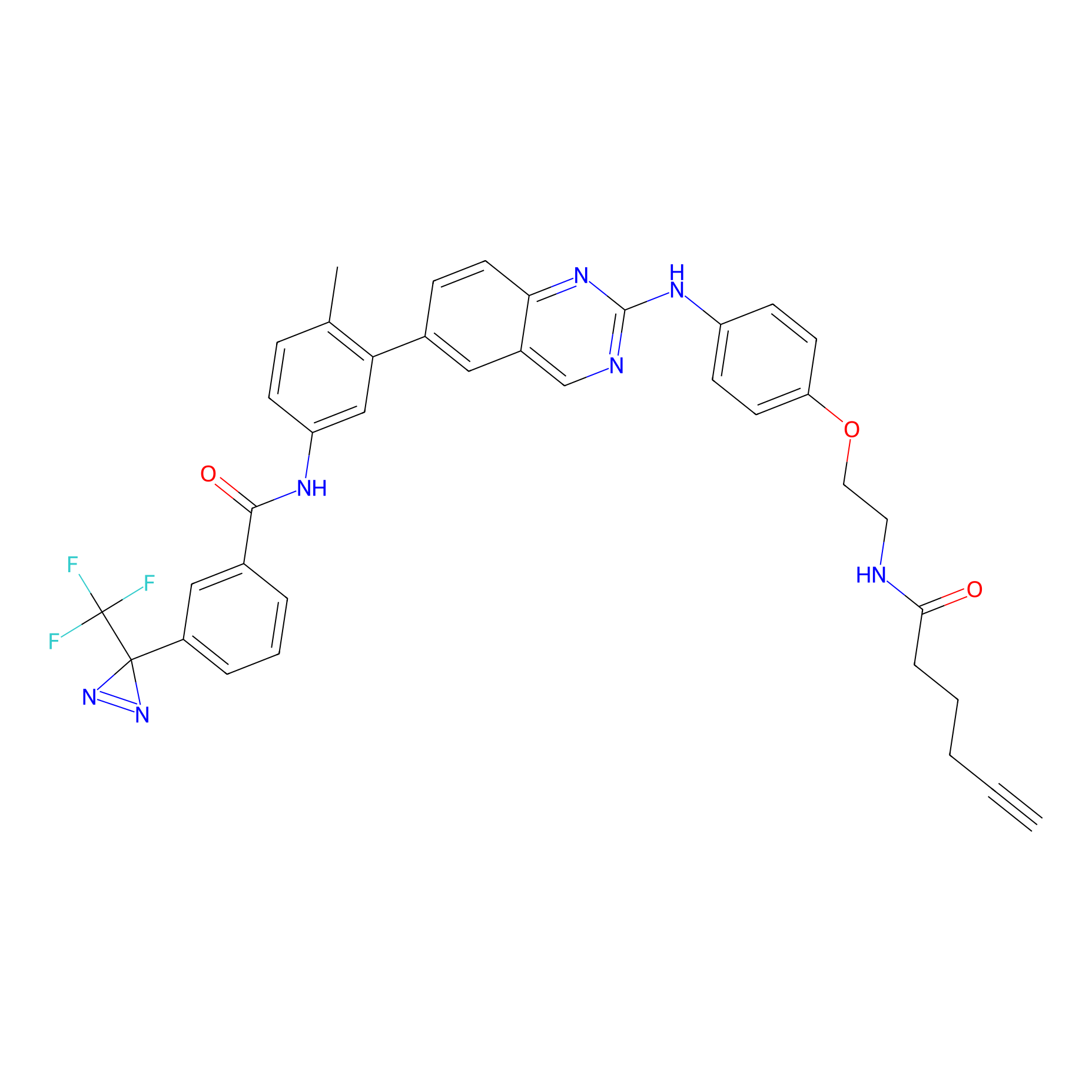 |
3.90 | LDD0075 | [7] | |
|
Probe 12 Probe Info |
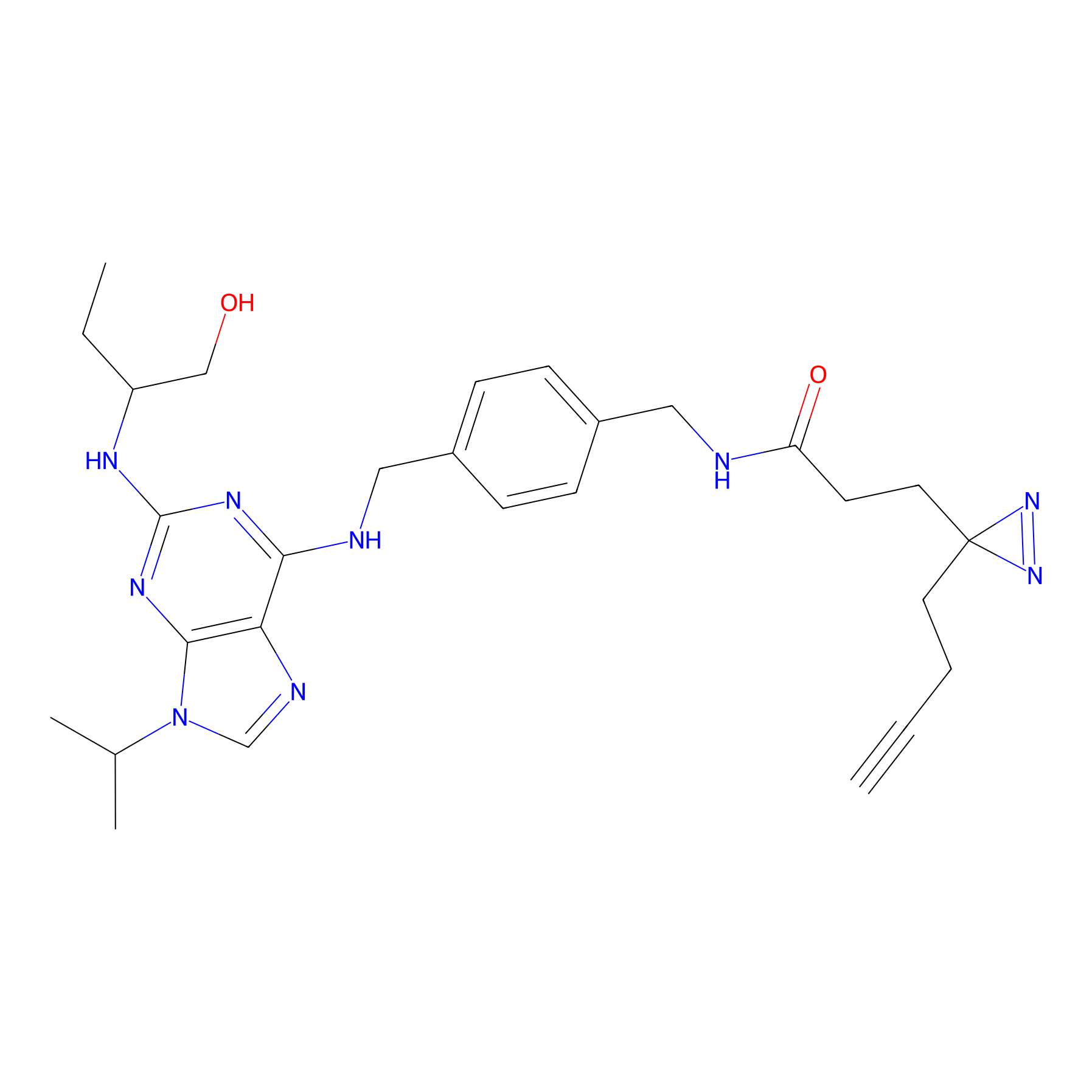 |
N.A. | LDD0420 | [8] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0165 | Compound 9 | HL-60 | N.A. | LDD0420 | [8] |
| LDCM0017 | DFG-out-2 | A431 | 3.90 | LDD0075 | [7] |
| LDCM0625 | F8 | Ramos | C89(1.09); C61(0.54); C44(1.22); C463(1.45) | LDD2187 | [9] |
| LDCM0572 | Fragment10 | Ramos | C89(1.09); C61(1.41); C44(0.71); C463(1.42) | LDD2189 | [9] |
| LDCM0573 | Fragment11 | Ramos | C89(18.41); C61(4.44); C44(0.71); C463(1.46) | LDD2190 | [9] |
| LDCM0574 | Fragment12 | Ramos | C89(1.07); C61(1.99); C44(0.80); C463(1.15) | LDD2191 | [9] |
| LDCM0575 | Fragment13 | Ramos | C89(1.06); C61(1.09); C44(0.85); C463(0.44) | LDD2192 | [9] |
| LDCM0576 | Fragment14 | Ramos | C89(0.46); C61(0.35); C44(1.16); C463(1.00) | LDD2193 | [9] |
| LDCM0579 | Fragment20 | Ramos | C89(1.23); C61(1.82); C44(0.86); C562(2.23) | LDD2194 | [9] |
| LDCM0580 | Fragment21 | Ramos | C89(0.96); C61(1.00); C44(0.92); C463(0.90) | LDD2195 | [9] |
| LDCM0582 | Fragment23 | Ramos | C89(1.04); C61(1.45); C44(0.97); C463(0.94) | LDD2196 | [9] |
| LDCM0578 | Fragment27 | Ramos | C89(1.00); C61(1.04); C44(1.12); C463(1.39) | LDD2197 | [9] |
| LDCM0586 | Fragment28 | Ramos | C89(1.00); C61(0.73); C44(0.53); C463(0.62) | LDD2198 | [9] |
| LDCM0588 | Fragment30 | Ramos | C89(1.13); C61(1.60); C44(1.04); C463(0.81) | LDD2199 | [9] |
| LDCM0589 | Fragment31 | Ramos | C89(0.97); C61(1.00); C44(0.88); C463(0.67) | LDD2200 | [9] |
| LDCM0590 | Fragment32 | Ramos | C89(0.74); C61(1.08); C44(0.59); C463(2.01) | LDD2201 | [9] |
| LDCM0468 | Fragment33 | Ramos | C89(0.96); C61(1.19); C44(0.74); C463(0.99) | LDD2202 | [9] |
| LDCM0596 | Fragment38 | Ramos | C89(0.96); C61(1.18); C44(0.75); C463(1.45) | LDD2203 | [9] |
| LDCM0566 | Fragment4 | Ramos | C89(1.03); C61(0.64); C44(0.85); C463(0.84) | LDD2184 | [9] |
| LDCM0610 | Fragment52 | Ramos | C89(2.08); C61(1.75); C44(1.11); C463(1.02) | LDD2204 | [9] |
| LDCM0614 | Fragment56 | Ramos | C89(1.26); C61(1.51); C44(0.99); C463(1.09) | LDD2205 | [9] |
| LDCM0569 | Fragment7 | Ramos | C89(1.13); C61(0.92); C44(0.90); C463(1.22) | LDD2186 | [9] |
| LDCM0571 | Fragment9 | Ramos | C89(1.70); C61(2.13); C44(1.15); C463(1.48) | LDD2188 | [9] |
| LDCM0107 | IAA | HeLa | N.A. | LDD0221 | [3] |
| LDCM0022 | KB02 | Ramos | C89(1.64); C61(0.56); C44(0.80); C463(1.36) | LDD2182 | [9] |
| LDCM0023 | KB03 | Jurkat | C972(4.51) | LDD0315 | [10] |
| LDCM0024 | KB05 | COLO792 | C89(2.35) | LDD3310 | [2] |
| LDCM0131 | RA190 | MM1.R | C899(1.92); C972(1.77); C677(1.28); C44(1.10) | LDD0304 | [11] |
| LDCM0016 | Ranjitkar_cp1 | A431 | 2.70 | LDD0074 | [7] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transporter and channel
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Amyloid-beta precursor protein (APP) | APP family | P05067 | |||
Other
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Nephrocystin-1 (NPHP1) | Nephrocystin-1 family | O15259 | |||
| Nephrocystin-4 (NPHP4) | NPHP4 family | O75161 | |||
The Drug(s) Related To This Target
Approved
| Drug Name | Drug Type | External ID | |||
|---|---|---|---|---|---|
| Fostamatinib | Small molecular drug | DB12010 | |||
| Leflunomide | Small molecular drug | DB01097 | |||
Investigative
References
