Details of the Target
General Information of Target
| Target ID | LDTP04595 | |||||
|---|---|---|---|---|---|---|
| Target Name | Activated RNA polymerase II transcriptional coactivator p15 (SUB1) | |||||
| Gene Name | SUB1 | |||||
| Gene ID | 10923 | |||||
| Synonyms |
PC4; RPO2TC1; Activated RNA polymerase II transcriptional coactivator p15; Positive cofactor 4; PC4; SUB1 homolog; p14 |
|||||
| 3D Structure | ||||||
| Sequence |
MPKSKELVSSSSSGSDSDSEVDKKLKRKKQVAPEKPVKKQKTGETSRALSSSKQSSSSRD
DNMFQIGKMRYVSVRDFKGKVLIDIREYWMDPEGEMKPGRKGISLNPEQWSQLKEQISDI DDAVRKL |
|||||
| Target Bioclass |
Other
|
|||||
| Family |
Transcriptional coactivator PC4 family
|
|||||
| Subcellular location |
Nucleus
|
|||||
| Function |
General coactivator that functions cooperatively with TAFs and mediates functional interactions between upstream activators and the general transcriptional machinery. May be involved in stabilizing the multiprotein transcription complex. Binds single-stranded DNA. Also binds, in vitro, non-specifically to double-stranded DNA (ds DNA).
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
AZ-9 Probe Info |
 |
3.82 | LDD0393 | [1] | |
|
TH211 Probe Info |
 |
Y88(7.03) | LDD0260 | [2] | |
|
C-Sul Probe Info |
 |
8.79 | LDD0066 | [3] | |
|
ONAyne Probe Info |
 |
K53(2.62) | LDD0274 | [4] | |
|
STPyne Probe Info |
 |
K29(2.42); K35(2.80); K38(6.18); K53(6.95) | LDD0277 | [4] | |
|
OPA-S-S-alkyne Probe Info |
 |
K41(1.41); K39(1.42); K80(1.69); K97(6.03) | LDD3494 | [5] | |
|
AMP probe Probe Info |
 |
N.A. | LDD0200 | [6] | |
|
ATP probe Probe Info |
 |
K101(0.00); K68(0.00); K80(0.00); K53(0.00) | LDD0199 | [6] | |
|
ATP probe Probe Info |
 |
K68(0.00); K53(0.00) | LDD0035 | [7] | |
|
NHS Probe Info |
 |
K68(0.00); K29(0.00) | LDD0010 | [8] | |
|
SF Probe Info |
 |
K80(0.00); K53(0.00) | LDD0028 | [9] | |
|
Ox-W18 Probe Info |
 |
W89(0.00); W110(0.00) | LDD2175 | [10] | |
|
1c-yne Probe Info |
 |
N.A. | LDD0228 | [11] | |
|
1d-yne Probe Info |
 |
N.A. | LDD0357 | [11] | |
|
HHS-465 Probe Info |
 |
K38(0.00); K29(0.00); K53(0.00); K101(0.00) | LDD2240 | [12] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C207 Probe Info |
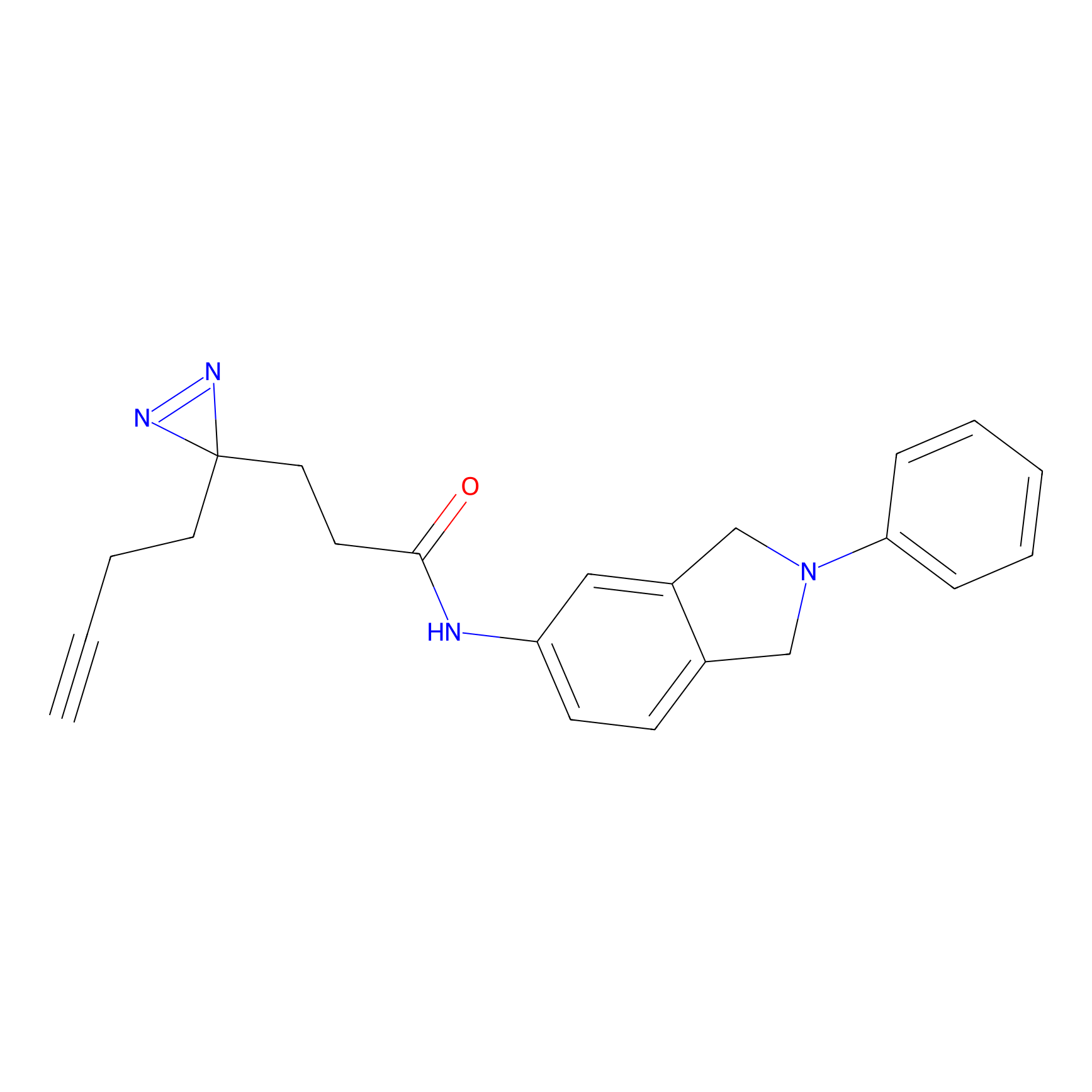 |
20.53 | LDD1882 | [13] | |
|
C354 Probe Info |
 |
7.57 | LDD2015 | [13] | |
|
C361 Probe Info |
 |
19.16 | LDD2022 | [13] | |
The Interaction Atlas With This Target
References
