Details of the Target
General Information of Target
| Target ID | LDTP01474 | |||||
|---|---|---|---|---|---|---|
| Target Name | Ubiquitin carboxyl-terminal hydrolase 19 (USP19) | |||||
| Gene Name | USP19 | |||||
| Gene ID | 10869 | |||||
| Synonyms |
KIAA0891; ZMYND9; Ubiquitin carboxyl-terminal hydrolase 19; EC 3.4.19.12; Deubiquitinating enzyme 19; Ubiquitin thioesterase 19; Ubiquitin-specific-processing protease 19; Zinc finger MYND domain-containing protein 9
|
|||||
| 3D Structure | ||||||
| Sequence |
MSGGASATGPRRGPPGLEDTTSKKKQKDRANQESKDGDPRKETGSRYVAQAGLEPLASGD
PSASASHAAGITGSRHRTRLFFPSSSGSASTPQEEQTKEGACEDPHDLLATPTPELLLDW RQSAEEVIVKLRVGVGPLQLEDVDAAFTDTDCVVRFAGGQQWGGVFYAEIKSSCAKVQTR KGSLLHLTLPKKVPMLTWPSLLVEADEQLCIPPLNSQTCLLGSEENLAPLAGEKAVPPGN DPVSPAMVRSRNPGKDDCAKEEMAVAADAATLVDEPESMVNLAFVKNDSYEKGPDSVVVH VYVKEICRDTSRVLFREQDFTLIFQTRDGNFLRLHPGCGPHTTFRWQVKLRNLIEPEQCT FCFTASRIDICLRKRQSQRWGGLEAPAARVGGAKVAVPTGPTPLDSTPPGGAPHPLTGQE EARAVEKDKSKARSEDTGLDSVATRTPMEHVTPKPETHLASPKPTCMVPPMPHSPVSGDS VEEEEEEEKKVCLPGFTGLVNLGNTCFMNSVIQSLSNTRELRDFFHDRSFEAEINYNNPL GTGGRLAIGFAVLLRALWKGTHHAFQPSKLKAIVASKASQFTGYAQHDAQEFMAFLLDGL HEDLNRIQNKPYTETVDSDGRPDEVVAEEAWQRHKMRNDSFIVDLFQGQYKSKLVCPVCA KVSITFDPFLYLPVPLPQKQKVLPVFYFAREPHSKPIKFLVSVSKENSTASEVLDSLSQS VHVKPENLRLAEVIKNRFHRVFLPSHSLDTVSPSDTLLCFELLSSELAKERVVVLEVQQR PQVPSVPISKCAACQRKQQSEDEKLKRCTRCYRVGYCNQLCQKTHWPDHKGLCRPENIGY PFLVSVPASRLTYARLAQLLEGYARYSVSVFQPPFQPGRMALESQSPGCTTLLSTGSLEA GDSERDPIQPPELQLVTPMAEGDTGLPRVWAAPDRGPVPSTSGISSEMLASGPIEVGSLP AGERVSRPEAAVPGYQHPSEAMNAHTPQFFIYKIDSSNREQRLEDKGDTPLELGDDCSLA LVWRNNERLQEFVLVASKELECAEDPGSAGEAARAGHFTLDQCLNLFTRPEVLAPEEAWY CPQCKQHREASKQLLLWRLPNVLIVQLKRFSFRSFIWRDKINDLVEFPVRNLDLSKFCIG QKEEQLPSYDLYAVINHYGGMIGGHYTACARLPNDRSSQRSDVGWRLFDDSTVTTVDESQ VVTRYAYVLFYRRRNSPVERPPRAGHSEHHPDLGPAAEAAASQASRIWQELEAEEEPVPE GSGPLGPWGPQDWVGPLPRGPTTPDEGCLRYFVLGTVAALVALVLNVFYPLVSQSRWR |
|||||
| Target Bioclass |
Enzyme
|
|||||
| Family |
Peptidase C19 family
|
|||||
| Subcellular location |
Endoplasmic reticulum membrane
|
|||||
| Function |
Deubiquitinating enzyme that regulates the degradation of various proteins. Deubiquitinates and prevents proteasomal degradation of RNF123 which in turn stimulates CDKN1B ubiquitin-dependent degradation thereby playing a role in cell proliferation. Involved in decreased protein synthesis in atrophying skeletal muscle. Modulates transcription of major myofibrillar proteins. Also involved in turnover of endoplasmic-reticulum-associated degradation (ERAD) substrates. Regulates the stability of BIRC2/c-IAP1 and BIRC3/c-IAP2 by preventing their ubiquitination. Required for cells to mount an appropriate response to hypoxia and rescues HIF1A from degradation in a non-catalytic manner. Plays an important role in 17 beta-estradiol (E2)-inhibited myogenesis. Decreases the levels of ubiquitinated proteins during skeletal muscle formation and acts to repress myogenesis. Exhibits a preference towards 'Lys-63'-linked ubiquitin chains.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
15.00 | LDD0402 | [1] | |
|
DA-P2 Probe Info |
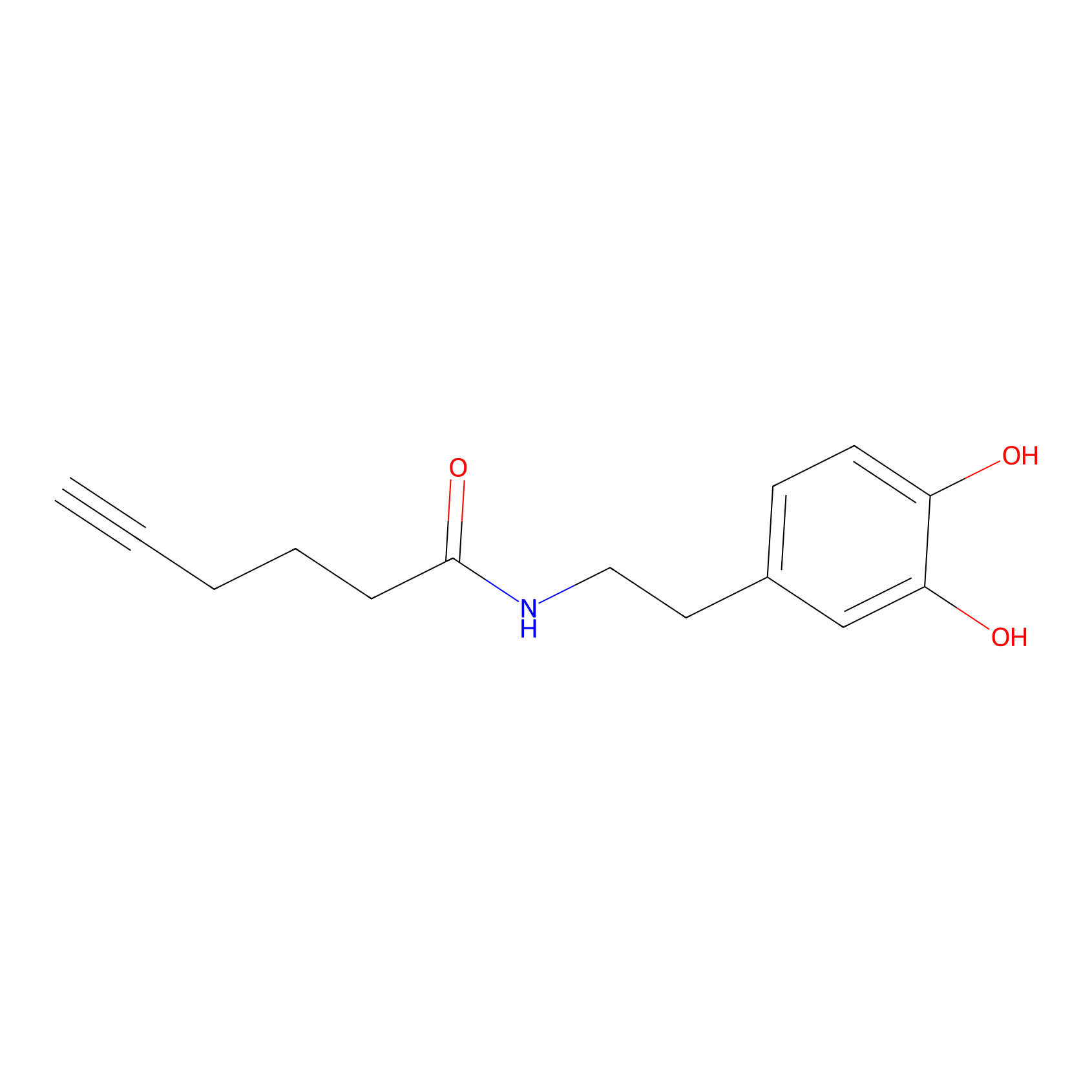 |
4.98 | LDD0348 | [2] | |
|
C-Sul Probe Info |
 |
7.02 | LDD0066 | [3] | |
|
DBIA Probe Info |
 |
C936(1.73) | LDD3310 | [4] | |
|
BTD Probe Info |
 |
C338(1.18) | LDD2107 | [5] | |
|
DA-P3 Probe Info |
 |
4.25 | LDD0180 | [2] | |
|
HHS-475 Probe Info |
 |
Y584(0.15); Y992(0.89) | LDD0264 | [6] | |
|
BDBM50514113 Probe Info |
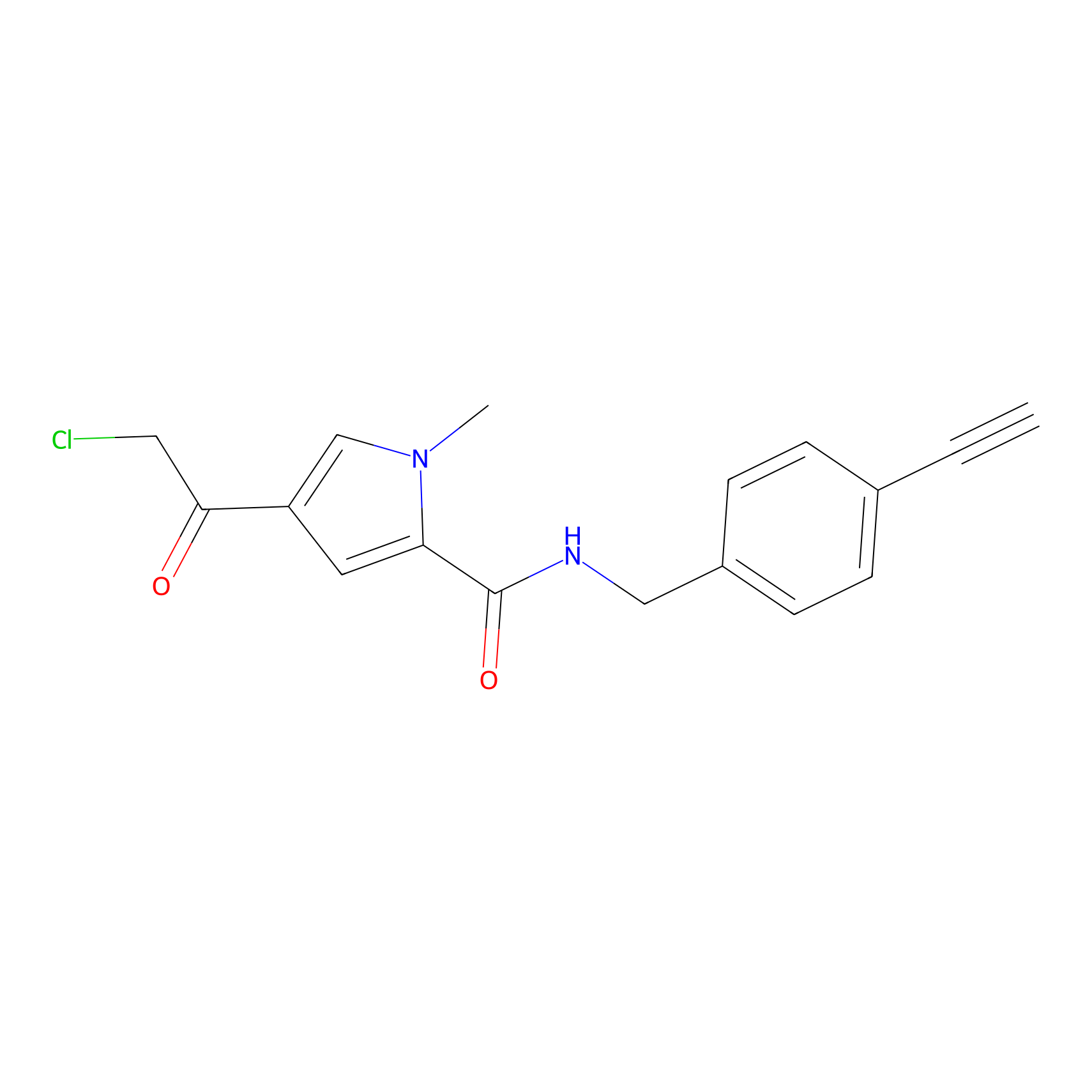 |
1.49 | LDD0041 | [7] | |
|
5E-2FA Probe Info |
 |
N.A. | LDD2235 | [8] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
C102(0.00); C833(0.00); C759(0.00) | LDD0038 | [9] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0032 | [10] | |
|
IPIAA_L Probe Info |
 |
N.A. | LDD0031 | [11] | |
|
Lodoacetamide azide Probe Info |
 |
C833(0.00); C759(0.00) | LDD0037 | [9] | |
|
JW-RF-010 Probe Info |
 |
N.A. | LDD0026 | [12] | |
|
TFBX Probe Info |
 |
N.A. | LDD0027 | [12] | |
|
IPM Probe Info |
 |
N.A. | LDD0005 | [13] | |
|
SF Probe Info |
 |
Y992(0.00); K993(0.00) | LDD0028 | [14] | |
|
VSF Probe Info |
 |
N.A. | LDD0007 | [13] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0217 | [15] | |
|
Methacrolein Probe Info |
 |
C338(0.00); C1042(0.00) | LDD0218 | [15] | |
|
W1 Probe Info |
 |
N.A. | LDD0236 | [16] | |
|
AOyne Probe Info |
 |
11.70 | LDD0443 | [17] | |
|
NAIA_5 Probe Info |
 |
N.A. | LDD2223 | [18] | |
|
HHS-465 Probe Info |
 |
K993(0.00); Y992(0.00) | LDD2240 | [19] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0524 | 2-Cyano-N-(2-morpholin-4-yl-ethyl)-acetamide | MDA-MB-231 | C1138(0.95) | LDD2117 | [5] |
| LDCM0156 | Aniline | NCI-H1299 | 11.47 | LDD0403 | [1] |
| LDCM0108 | Chloroacetamide | HeLa | H335(0.00); C1042(0.00) | LDD0222 | [15] |
| LDCM0028 | Dobutamine | HEK-293T | 4.25 | LDD0180 | [2] |
| LDCM0213 | Electrophilic fragment 2 | MDA-MB-231 | C817(1.00); C821(0.99) | LDD1702 | [5] |
| LDCM0031 | Epigallocatechin gallate | HEK-293T | 18.67 | LDD0183 | [2] |
| LDCM0116 | HHS-0101 | DM93 | Y584(0.15); Y992(0.89) | LDD0264 | [6] |
| LDCM0117 | HHS-0201 | DM93 | Y992(1.71); Y584(1.96) | LDD0265 | [6] |
| LDCM0118 | HHS-0301 | DM93 | Y992(0.58); Y584(0.87) | LDD0266 | [6] |
| LDCM0119 | HHS-0401 | DM93 | Y992(0.20) | LDD0267 | [6] |
| LDCM0120 | HHS-0701 | DM93 | Y992(1.49); Y584(2.05) | LDD0268 | [6] |
| LDCM0022 | KB02 | HEK-293T | C1042(0.90); C466(1.03); C833(0.91); C1017(0.97) | LDD1492 | [20] |
| LDCM0023 | KB03 | HEK-293T | C1042(1.11); C466(1.28); C833(1.28); C1017(1.16) | LDD1497 | [20] |
| LDCM0024 | KB05 | COLO792 | C936(1.73) | LDD3310 | [4] |
| LDCM0030 | Luteolin | HEK-293T | 7.20 | LDD0182 | [2] |
| LDCM0514 | Nucleophilic fragment 20a | MDA-MB-231 | C338(1.18) | LDD2107 | [5] |
| LDCM0530 | Nucleophilic fragment 28a | MDA-MB-231 | C1138(1.01); C338(0.66) | LDD2123 | [5] |
| LDCM0550 | Nucleophilic fragment 5a | MDA-MB-231 | C338(1.27) | LDD2144 | [5] |
| LDCM0029 | Quercetin | HEK-293T | 7.87 | LDD0181 | [2] |
| LDCM0002 | Ward_cp1 | U2OS | 1.49 | LDD0041 | [7] |
References
