Details of the Target
General Information of Target
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
TH211 Probe Info |
 |
Y84(16.42); Y38(11.06); Y43(10.74); Y41(6.07) | LDD0257 | [1] | |
|
TH216 Probe Info |
 |
Y38(20.00); Y43(20.00); Y84(20.00); Y41(10.90) | LDD0259 | [1] | |
|
OPA-S-S-alkyne Probe Info |
 |
K86(8.96); K109(11.43) | LDD3494 | [2] | |
|
Probe 1 Probe Info |
 |
Y41(140.90); Y84(61.09) | LDD3495 | [3] | |
|
HHS-475 Probe Info |
 |
Y84(1.13) | LDD0264 | [4] | |
|
HHS-465 Probe Info |
 |
Y84(2.95) | LDD2237 | [5] | |
|
5E-2FA Probe Info |
 |
N.A. | LDD2235 | [6] | |
|
m-APA Probe Info |
 |
N.A. | LDD2231 | [6] | |
|
aONE Probe Info |
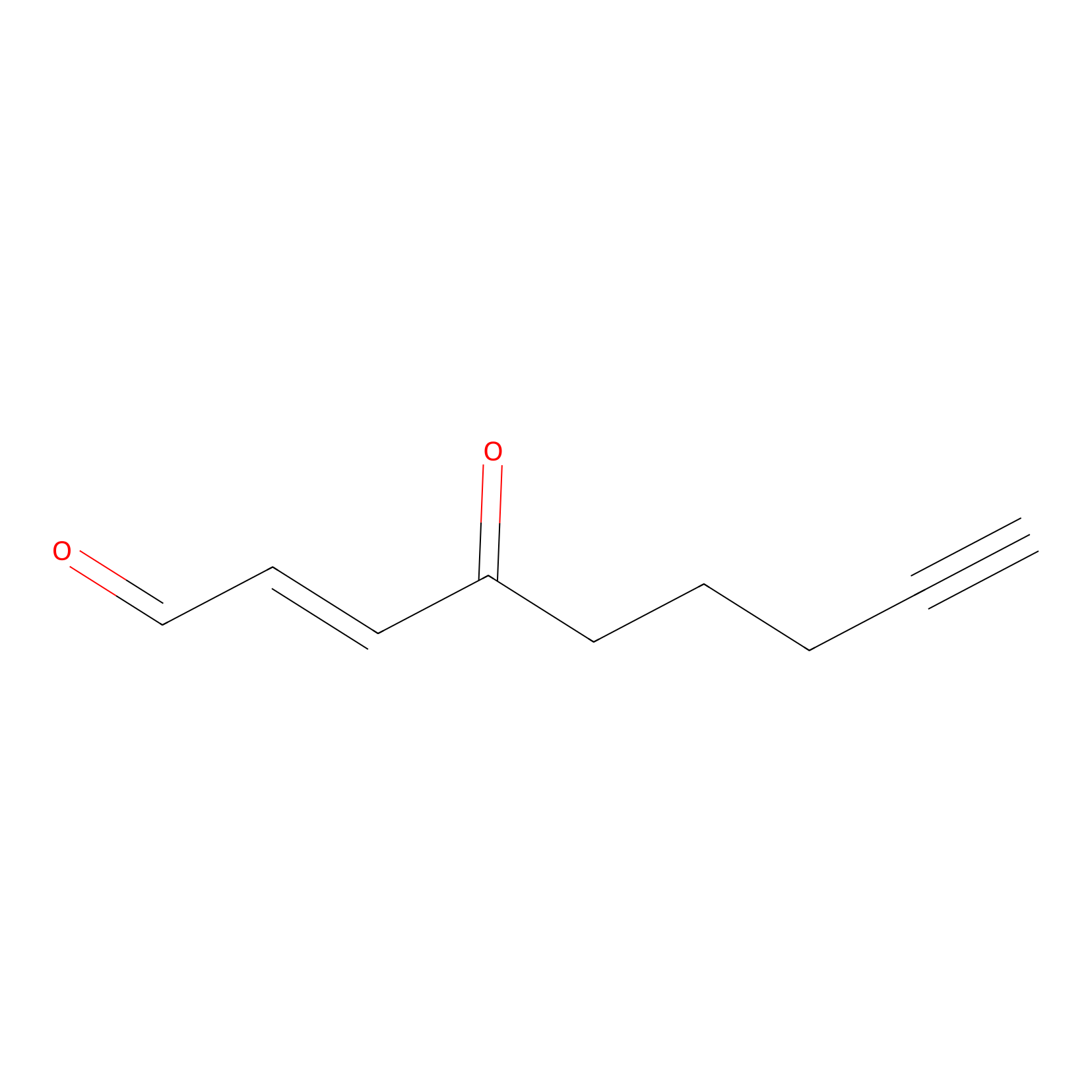 |
K109(0.00); K117(0.00) | LDD0002 | [7] | |
|
OSF Probe Info |
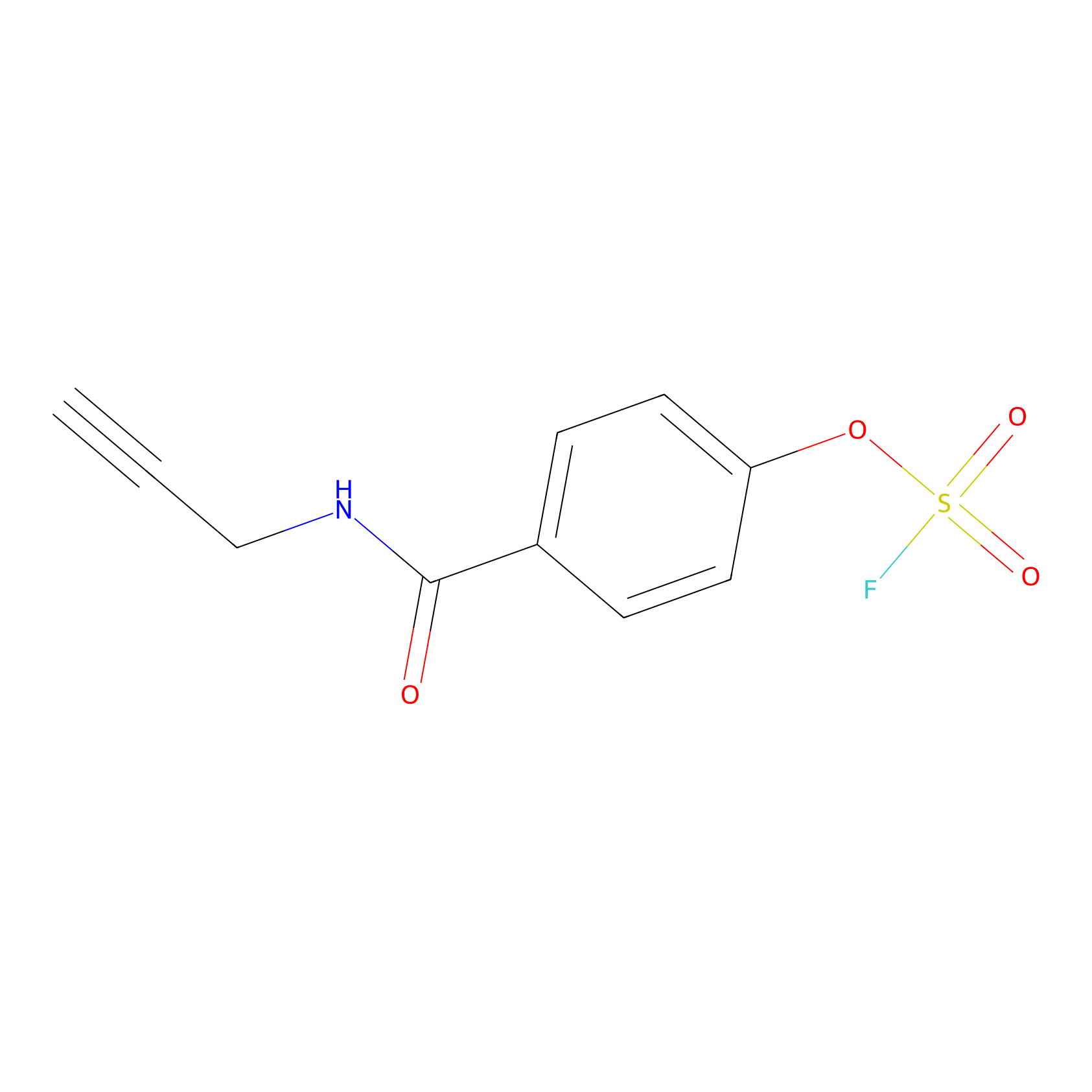 |
Y84(0.00); H83(0.00); Y41(0.00); H110(0.00) | LDD0029 | [8] | |
|
SF Probe Info |
 |
K117(0.00); K12(0.00); Y38(0.00); K121(0.00) | LDD0028 | [8] | |
|
1c-yne Probe Info |
 |
K44(0.00); K58(0.00); K117(0.00) | LDD0228 | [9] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0217 | [10] | |
|
Crotonaldehyde Probe Info |
 |
N.A. | LDD0219 | [10] | |
|
Methacrolein Probe Info |
 |
N.A. | LDD0218 | [10] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
VE-P Probe Info |
 |
N.A. | LDD0396 | [11] | |
|
Photocelecoxib Probe Info |
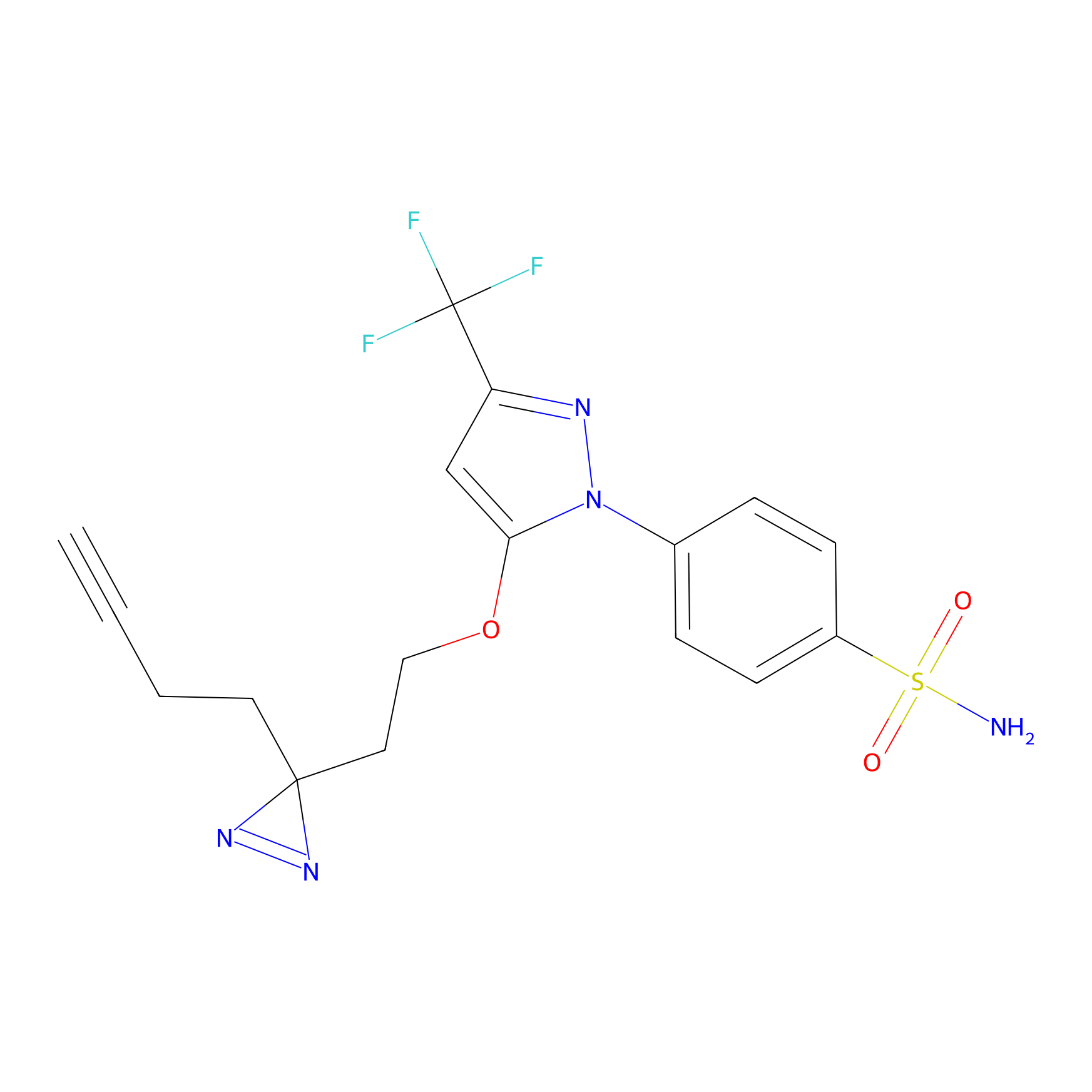 |
H50(0.00); S65(0.00); V67(0.00); D69(0.00) | LDD0019 | [12] | |
|
Photoindomethacin Probe Info |
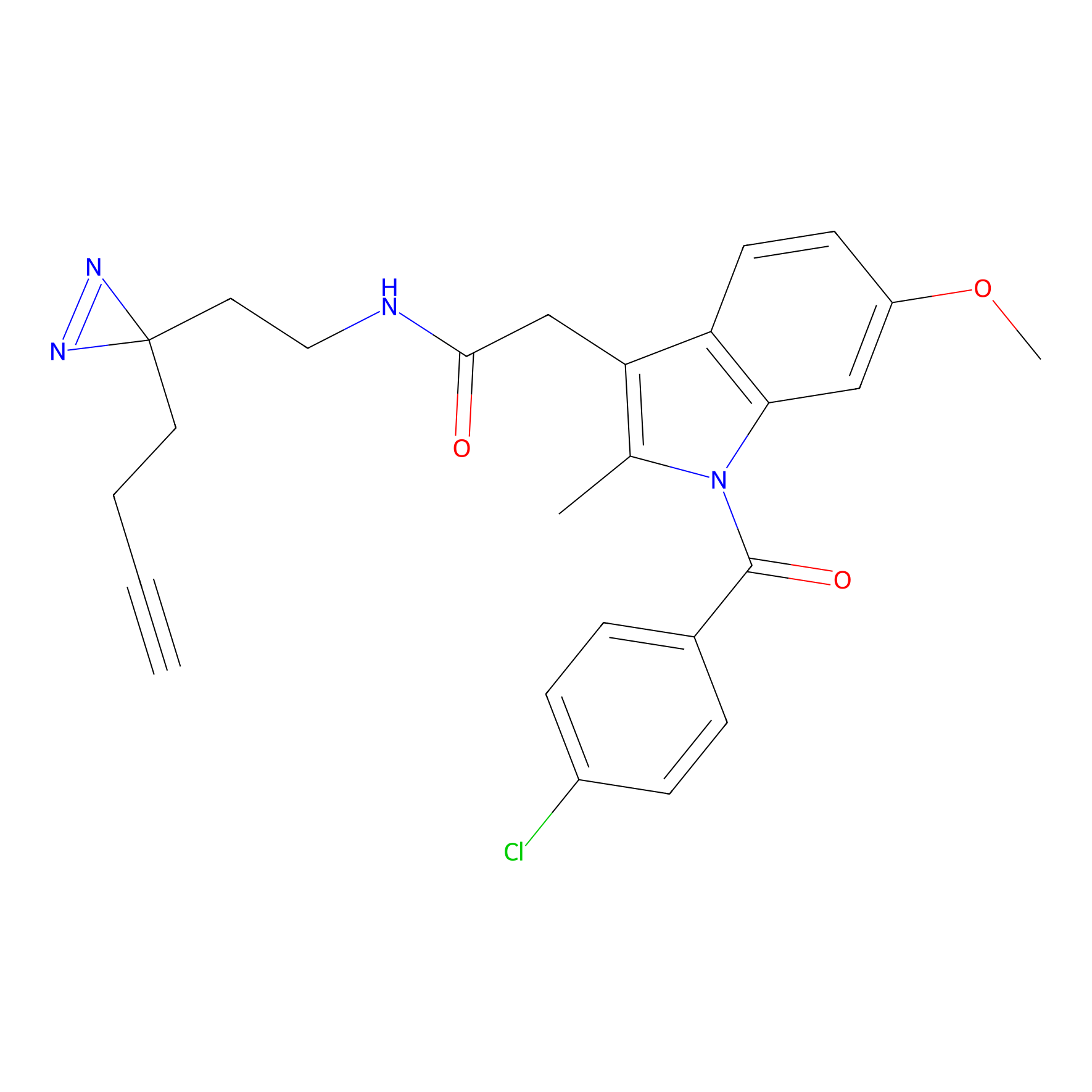 |
N.A. | LDD0155 | [12] | |
|
Photonaproxen Probe Info |
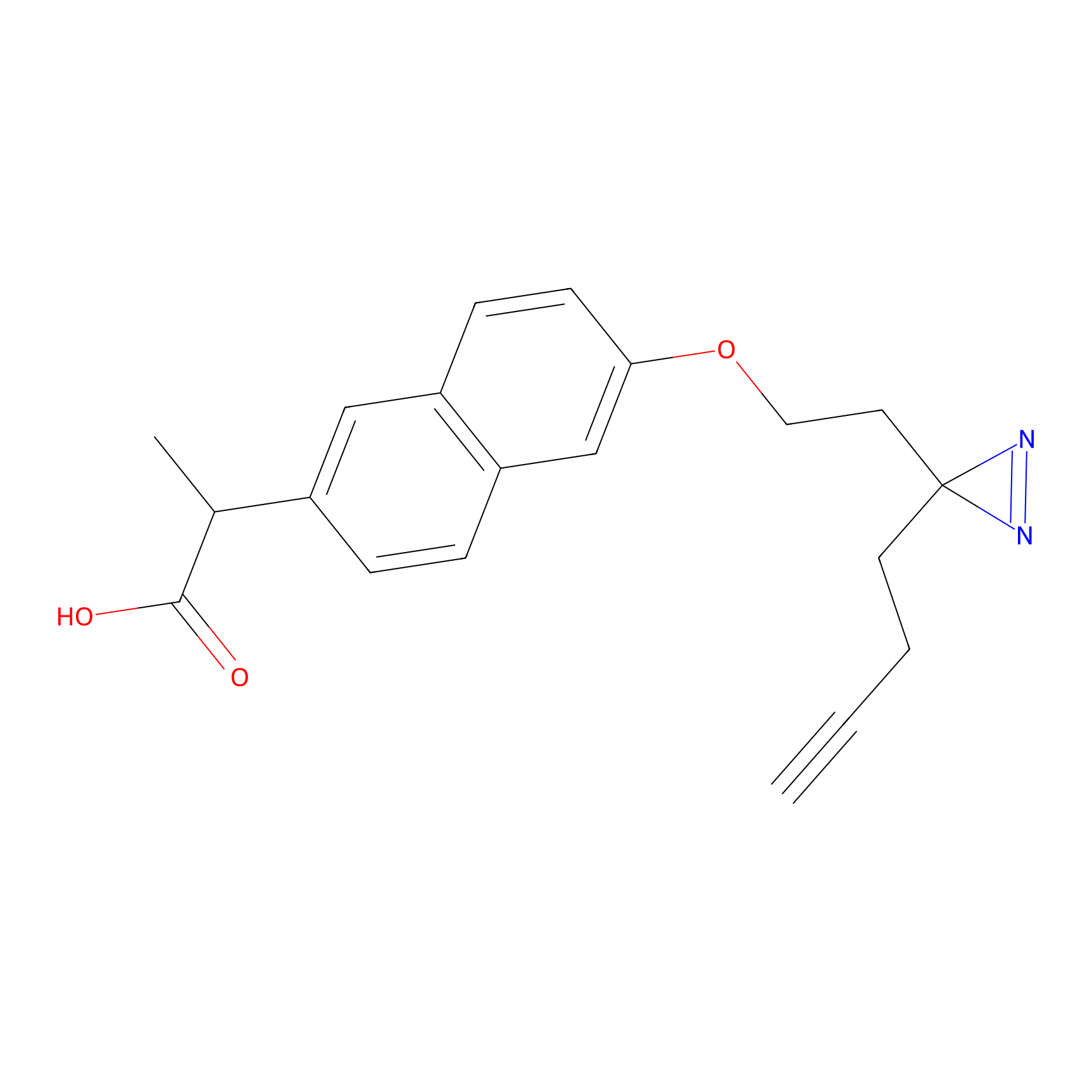 |
D69(0.00); F71(0.00); E72(0.00); R73(0.00) | LDD0156 | [12] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0108 | Chloroacetamide | HeLa | N.A. | LDD0222 | [10] |
| LDCM0116 | HHS-0101 | DM93 | Y84(1.13) | LDD0264 | [4] |
| LDCM0117 | HHS-0201 | DM93 | Y84(1.00) | LDD0265 | [4] |
| LDCM0118 | HHS-0301 | DM93 | Y84(1.27) | LDD0266 | [4] |
| LDCM0119 | HHS-0401 | DM93 | Y84(1.26) | LDD0267 | [4] |
| LDCM0120 | HHS-0701 | DM93 | Y84(1.74) | LDD0268 | [4] |
| LDCM0107 | IAA | HeLa | N.A. | LDD0221 | [10] |
| LDCM0109 | NEM | HeLa | N.A. | LDD0223 | [10] |
The Interaction Atlas With This Target
References
