Details of the Target
General Information of Target
| Target ID | LDTP18400 | |||||
|---|---|---|---|---|---|---|
| Target Name | DENN domain-containing protein 2D (DENND2D) | |||||
| Gene Name | DENND2D | |||||
| Gene ID | 79961 | |||||
| Synonyms |
DENN domain-containing protein 2D |
|||||
| 3D Structure | ||||||
| Sequence |
MSNFLHLKYNEKSVSVTKALTVRFLTKRFIGEYASNFESIYKKHLCLERKQLNLEIYDPC
SQTQKAKFSLTSELHWADGFVIVYDISDRSSFAFAKALIYRIREPQTSHCKRAVESAVFL VGNKRDLCHVREVGWEEGQKLALENRCQFCELSAAEQSLEVEMMFIRIIKDILINFKLKE KRRPSGSKSMAKLINNVFGKRRKSV |
|||||
| Target Bioclass |
Other
|
|||||
| Subcellular location |
Cytoplasm
|
|||||
| Function | Guanine nucleotide exchange factor (GEF) which may activate RAB9A and RAB9B. Promotes the exchange of GDP to GTP, converting inactive GDP-bound Rab proteins into their active GTP-bound form. | |||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Target Site Mutations in Different Cell Lines
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
Johansson_61 Probe Info |
 |
_(20.00) | LDD1487 | [1] | |
|
YY4-yne Probe Info |
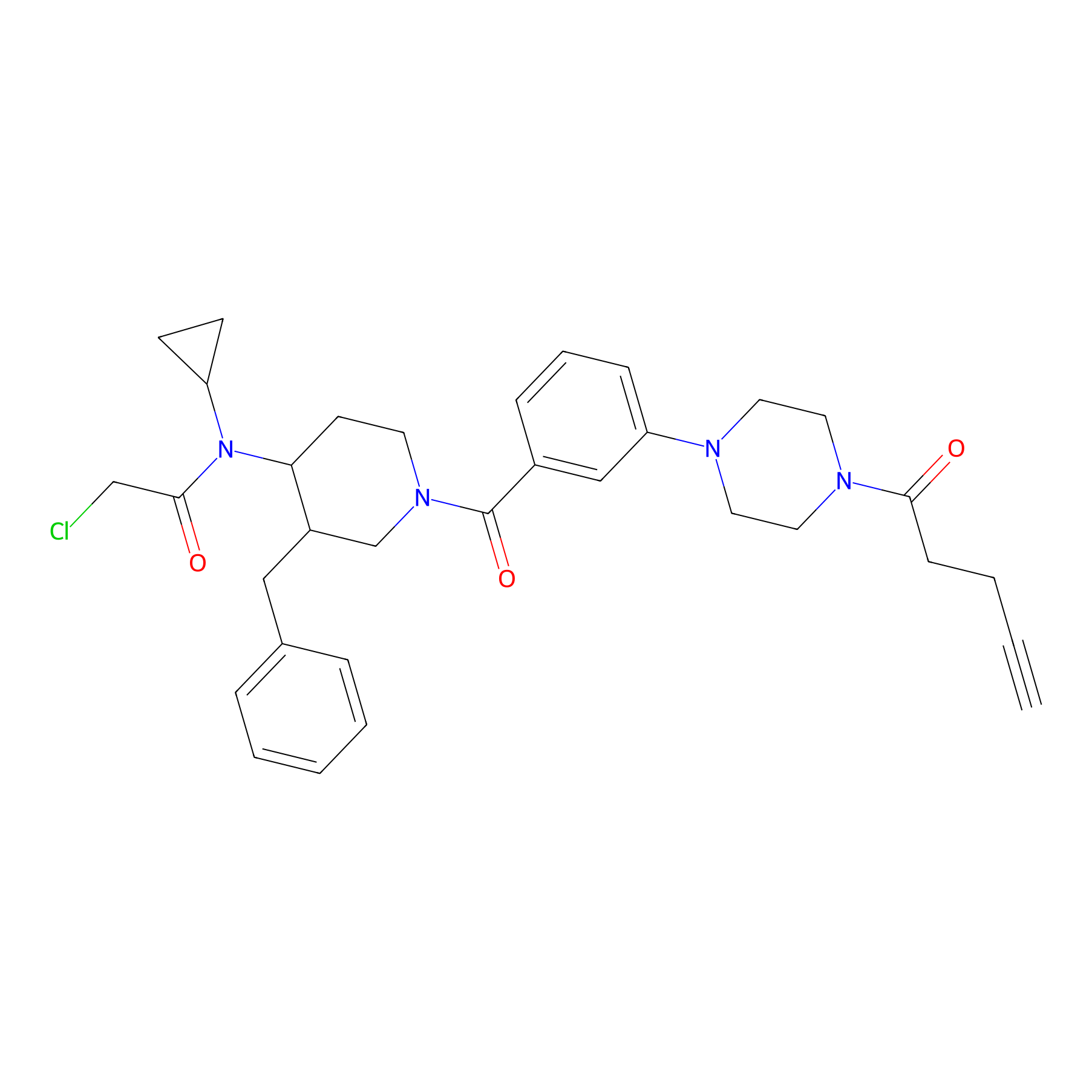 |
4.68 | LDD0400 | [2] | |
|
HPAP Probe Info |
 |
4.43 | LDD0064 | [3] | |
|
DBIA Probe Info |
 |
C104(38.94) | LDD0209 | [4] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0162 | [5] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0616 | Fragment61 | Jurkat | _(10.77) | LDD1489 | [1] |
| LDCM0615 | Fragment63-R | Jurkat | _(20.00) | LDD1487 | [1] |
| LDCM0022 | KB02 | AGS | C104(1.18) | LDD2263 | [6] |
| LDCM0023 | KB03 | Jurkat | C104(38.94) | LDD0209 | [4] |
| LDCM0024 | KB05 | HMCB | C104(1.68) | LDD3312 | [6] |
| LDCM0014 | Panhematin | hPBMC | 4.43 | LDD0064 | [3] |
| LDCM0154 | YY4 | T cell | 4.68 | LDD0400 | [2] |
The Interaction Atlas With This Target
References
