Details of the Target
General Information of Target
| Target ID | LDTP13305 | |||||
|---|---|---|---|---|---|---|
| Target Name | SAP30-binding protein (SAP30BP) | |||||
| Gene Name | SAP30BP | |||||
| Gene ID | 29115 | |||||
| Synonyms |
HCNGP; HTRG; HTRP; SAP30-binding protein; Transcriptional regulator protein HCNGP |
|||||
| 3D Structure | ||||||
| Sequence |
MSRGPEEVNRLTESTYRNVMEQFNPGLRNLINLGKNYEKAVNAMILAGKAYYDGVAKIGE
IATGSPVSTELGHVLIEISSTHKKLNESLDENFKKFHKEIIHELEKKIELDVKYMNATLK RYQTEHKNKLESLEKSQAELKKIRRKSQGSRNALKYEHKEIEYVETVTSRQSEIQKFIAD GCKEALLEEKRRFCFLVDKHCGFANHIHYYHLQSAELLNSKLPRWQETCVDAIKVPEKIM NMIEEIKTPASTPVSGTPQASPMIERSNVVRKDYDTLSKCSPKMPPAPSGRAYTSPLIDM FNNPATAAPNSQRVNNSTGTSEDPSLQRSVSVATGLNMMKKQKVKTIFPHTAGSNKTLLS FAQGDVITLLIPEEKDGWLYGEHDVSKARGWFPSSYTKLLEENETEAVTVPTPSPTPVRS ISTVNLSENSSVVIPPPDYLECLSMGAAADRRADSARTTSTFKAPASKPETAAPNDANGT AKPPFLSGENPFATVKLRPTVTNDRSAPIIR |
|||||
| Target Bioclass |
Transcription factor
|
|||||
| Family |
HCNGP family
|
|||||
| Subcellular location |
Nucleus
|
|||||
| Function |
Plays a role in transcriptional repression by promoting histone deacetylase activity, leading to deacetylation of histone H3. May be involved in the regulation of beta-2-microglobulin genes.; (Microbial infection) Involved in transcriptional repression of HHV-1 genes TK and gC.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| HCT15 | SNV: p.G3R | . | |||
| JURKAT | SNV: p.D89E | Compound 10 Probe Info | |||
| MCC13 | SNV: p.S104F | DBIA Probe Info | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
12.55 | LDD0402 | [1] | |
|
STPyne Probe Info |
 |
K228(5.88) | LDD0277 | [2] | |
|
Probe 1 Probe Info |
 |
Y46(14.19); Y61(35.51); Y151(16.72) | LDD3495 | [3] | |
|
DBIA Probe Info |
 |
C127(2.26) | LDD3319 | [4] | |
|
BTD Probe Info |
 |
C127(0.59) | LDD2095 | [5] | |
|
AHL-Pu-1 Probe Info |
 |
C172(6.92) | LDD0169 | [6] | |
|
HHS-475 Probe Info |
 |
Y151(1.05) | LDD0264 | [7] | |
|
HHS-465 Probe Info |
 |
Y151(10.00) | LDD2237 | [8] | |
|
ATP probe Probe Info |
 |
N.A. | LDD0199 | [9] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0036 | [10] | |
|
Lodoacetamide azide Probe Info |
 |
N.A. | LDD0037 | [10] | |
|
NAIA_4 Probe Info |
 |
C127(0.00); C172(0.00) | LDD2226 | [11] | |
|
TFBX Probe Info |
 |
N.A. | LDD0027 | [12] | |
|
WYneN Probe Info |
 |
N.A. | LDD0021 | [13] | |
|
Compound 10 Probe Info |
 |
N.A. | LDD2216 | [14] | |
|
Compound 11 Probe Info |
 |
N.A. | LDD2213 | [14] | |
|
ENE Probe Info |
 |
N.A. | LDD0006 | [13] | |
|
IPM Probe Info |
 |
N.A. | LDD0005 | [13] | |
|
NPM Probe Info |
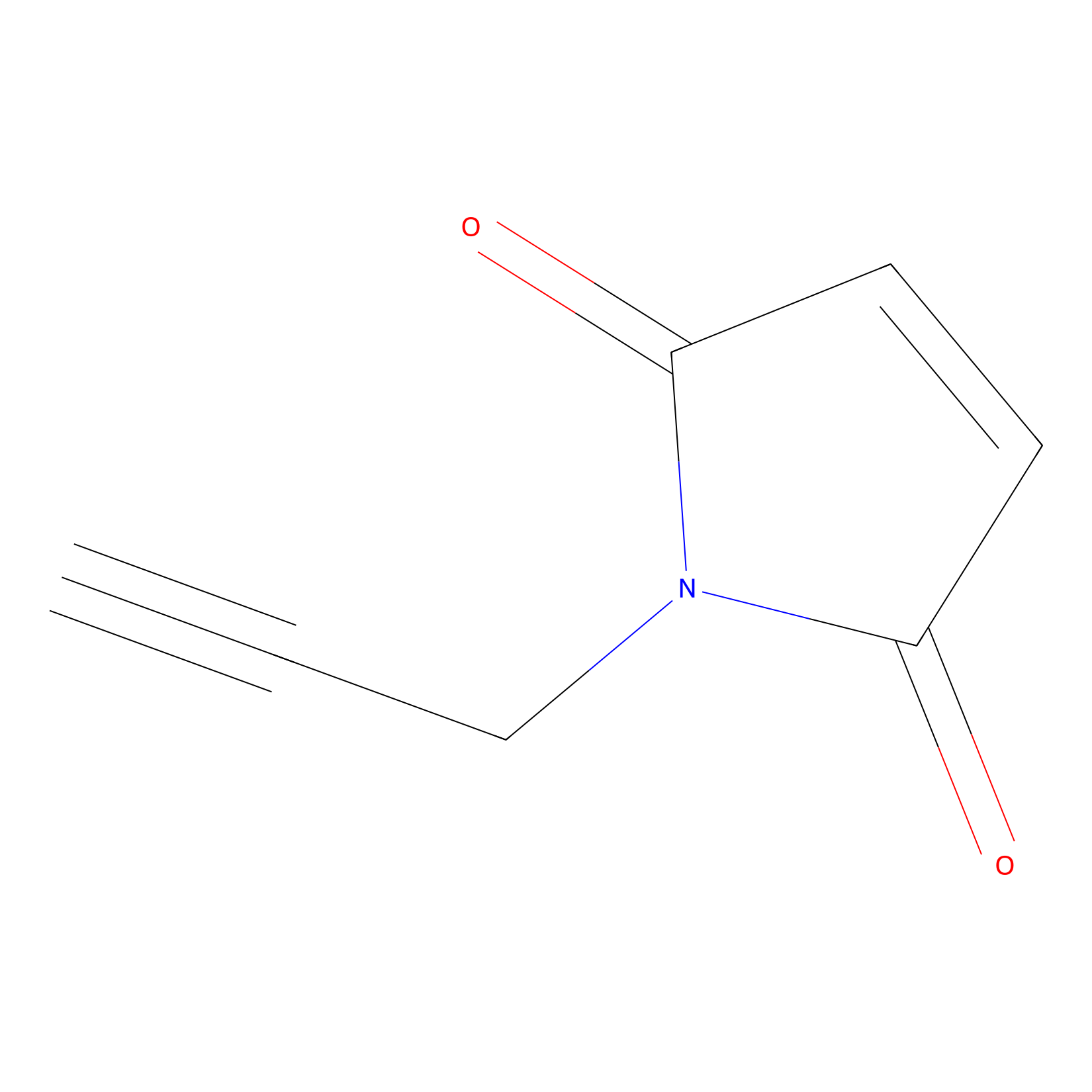 |
N.A. | LDD0016 | [13] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0217 | [15] | |
|
AOyne Probe Info |
 |
15.00 | LDD0443 | [16] | |
|
NAIA_5 Probe Info |
 |
C127(0.00); C172(0.00) | LDD2223 | [11] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C040 Probe Info |
 |
6.87 | LDD1740 | [17] | |
|
C041 Probe Info |
 |
5.66 | LDD1741 | [17] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0502 | 1-(Cyanoacetyl)piperidine | MDA-MB-231 | C127(0.59) | LDD2095 | [5] |
| LDCM0524 | 2-Cyano-N-(2-morpholin-4-yl-ethyl)-acetamide | MDA-MB-231 | C127(1.27) | LDD2117 | [5] |
| LDCM0539 | 3-(4-Isopropylpiperazin-1-yl)-3-oxopropanenitrile | MDA-MB-231 | C127(0.47) | LDD2132 | [5] |
| LDCM0026 | 4SU-RNA+native RNA | HEK-293T | C172(6.92) | LDD0169 | [6] |
| LDCM0156 | Aniline | NCI-H1299 | 9.73 | LDD0403 | [1] |
| LDCM0020 | ARS-1620 | HCC44 | C172(1.07) | LDD2171 | [18] |
| LDCM0108 | Chloroacetamide | HeLa | N.A. | LDD0222 | [15] |
| LDCM0632 | CL-Sc | Hep-G2 | C127(1.34); C127(1.28); C172(0.98) | LDD2227 | [11] |
| LDCM0213 | Electrophilic fragment 2 | MDA-MB-231 | C127(1.81) | LDD1702 | [5] |
| LDCM0625 | F8 | Ramos | C127(1.18) | LDD2187 | [19] |
| LDCM0572 | Fragment10 | Ramos | C127(1.78) | LDD2189 | [19] |
| LDCM0573 | Fragment11 | Ramos | C127(20.00) | LDD2190 | [19] |
| LDCM0574 | Fragment12 | Ramos | C127(1.52) | LDD2191 | [19] |
| LDCM0575 | Fragment13 | Ramos | C127(2.07) | LDD2192 | [19] |
| LDCM0576 | Fragment14 | Ramos | C127(1.04) | LDD2193 | [19] |
| LDCM0579 | Fragment20 | Ramos | C127(1.85) | LDD2194 | [19] |
| LDCM0580 | Fragment21 | Ramos | C127(1.89) | LDD2195 | [19] |
| LDCM0582 | Fragment23 | Ramos | C127(0.90) | LDD2196 | [19] |
| LDCM0578 | Fragment27 | Ramos | C127(5.39) | LDD2197 | [19] |
| LDCM0586 | Fragment28 | Ramos | C127(1.06) | LDD2198 | [19] |
| LDCM0588 | Fragment30 | Ramos | C127(1.32) | LDD2199 | [19] |
| LDCM0589 | Fragment31 | Ramos | C127(1.50) | LDD2200 | [19] |
| LDCM0590 | Fragment32 | Ramos | C127(2.13) | LDD2201 | [19] |
| LDCM0468 | Fragment33 | Ramos | C127(1.41) | LDD2202 | [19] |
| LDCM0596 | Fragment38 | Ramos | C127(1.79) | LDD2203 | [19] |
| LDCM0566 | Fragment4 | Ramos | C127(0.86) | LDD2184 | [19] |
| LDCM0610 | Fragment52 | Ramos | C127(2.03) | LDD2204 | [19] |
| LDCM0614 | Fragment56 | Ramos | C127(1.40) | LDD2205 | [19] |
| LDCM0569 | Fragment7 | Ramos | C127(0.89) | LDD2186 | [19] |
| LDCM0571 | Fragment9 | Ramos | C127(1.53) | LDD2188 | [19] |
| LDCM0116 | HHS-0101 | DM93 | Y151(1.05) | LDD0264 | [7] |
| LDCM0118 | HHS-0301 | DM93 | Y151(0.90) | LDD0266 | [7] |
| LDCM0119 | HHS-0401 | DM93 | Y151(1.67) | LDD0267 | [7] |
| LDCM0120 | HHS-0701 | DM93 | Y151(2.11) | LDD0268 | [7] |
| LDCM0022 | KB02 | HEK-293T | C127(0.98); C172(0.95) | LDD1492 | [20] |
| LDCM0023 | KB03 | HEK-293T | C127(0.96); C172(1.07) | LDD1497 | [20] |
| LDCM0024 | KB05 | MEWO | C127(2.26) | LDD3319 | [4] |
| LDCM0507 | Nucleophilic fragment 16b | MDA-MB-231 | C127(0.45) | LDD2100 | [5] |
| LDCM0530 | Nucleophilic fragment 28a | MDA-MB-231 | C127(1.20) | LDD2123 | [5] |
| LDCM0547 | Nucleophilic fragment 41 | MDA-MB-231 | C127(0.71) | LDD2141 | [5] |
| LDCM0021 | THZ1 | HCT 116 | C172(1.07) | LDD2173 | [18] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Transcription factor
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| THAP domain-containing protein 1 (THAP1) | THAP1 family | Q9NVV9 | |||
Other
References
