Details of the Target
General Information of Target
| Target ID | LDTP10649 | |||||
|---|---|---|---|---|---|---|
| Target Name | 28S rRNA (cytosine-C(5))-methyltransferase (NSUN5) | |||||
| Gene Name | NSUN5 | |||||
| Gene ID | 55695 | |||||
| Synonyms |
NSUN5A; WBSCR20; WBSCR20A; 28S rRNA; cytosine-C(5))-methyltransferase; EC 2.1.1.-; NOL1-related protein; NOL1R; NOL1/NOP2/Sun domain family member 5; Williams-Beuren syndrome chromosomal region 20A protein
|
|||||
| 3D Structure | ||||||
| Sequence |
MTGYEARLITFGTWMYSVNKEQLARAGFYAIGQEDKVQCFHCGGGLANWKPKEDPWEQHA
KWYPGCKYLLEEKGHEYINNIHLTRSLEGALVQTTKKTPSLTKRISDTIFPNPMLQEAIR MGFDFKDVKKIMEERIQTSGSNYKTLEVLVADLVSAQKDTTENELNQTSLQREISPEEPL RRLQEEKLCKICMDRHIAVVFIPCGHLVTCKQCAEAVDRCPMCSAVIDFKQRVFMS |
|||||
| Target Bioclass |
Enzyme
|
|||||
| Family |
Class I-like SAM-binding methyltransferase superfamily, RsmB/NOP family
|
|||||
| Subcellular location |
Nucleus, nucleolus
|
|||||
| Function |
S-adenosyl-L-methionine-dependent methyltransferase that specifically methylates the C(5) position of cytosine 3782 (m5C3782) in 28S rRNA. m5C3782 promotes protein translation without affecting ribosome biogenesis and fidelity. Required for corpus callosum and cerebral cortex development.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
15.00 | LDD0402 | [1] | |
|
DBIA Probe Info |
 |
C41(5.35) | LDD3312 | [2] | |
|
AHL-Pu-1 Probe Info |
 |
C308(2.23) | LDD0173 | [3] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
C343(0.00); C404(0.00); C41(0.00); C146(0.00) | LDD0038 | [4] | |
|
IA-alkyne Probe Info |
 |
C343(0.00); C404(0.00); C146(0.00); C41(0.00) | LDD0036 | [4] | |
|
IPIAA_L Probe Info |
 |
N.A. | LDD0031 | [5] | |
|
Lodoacetamide azide Probe Info |
 |
C343(0.00); C404(0.00); C146(0.00); C308(0.00) | LDD0037 | [4] | |
|
NAIA_4 Probe Info |
 |
C146(0.00); C308(0.00); C404(0.00) | LDD2226 | [6] | |
|
TFBX Probe Info |
 |
N.A. | LDD0027 | [7] | |
|
Compound 10 Probe Info |
 |
N.A. | LDD2216 | [8] | |
|
Compound 11 Probe Info |
 |
N.A. | LDD2213 | [8] | |
|
IPM Probe Info |
 |
N.A. | LDD0005 | [9] | |
|
8d Probe Info |
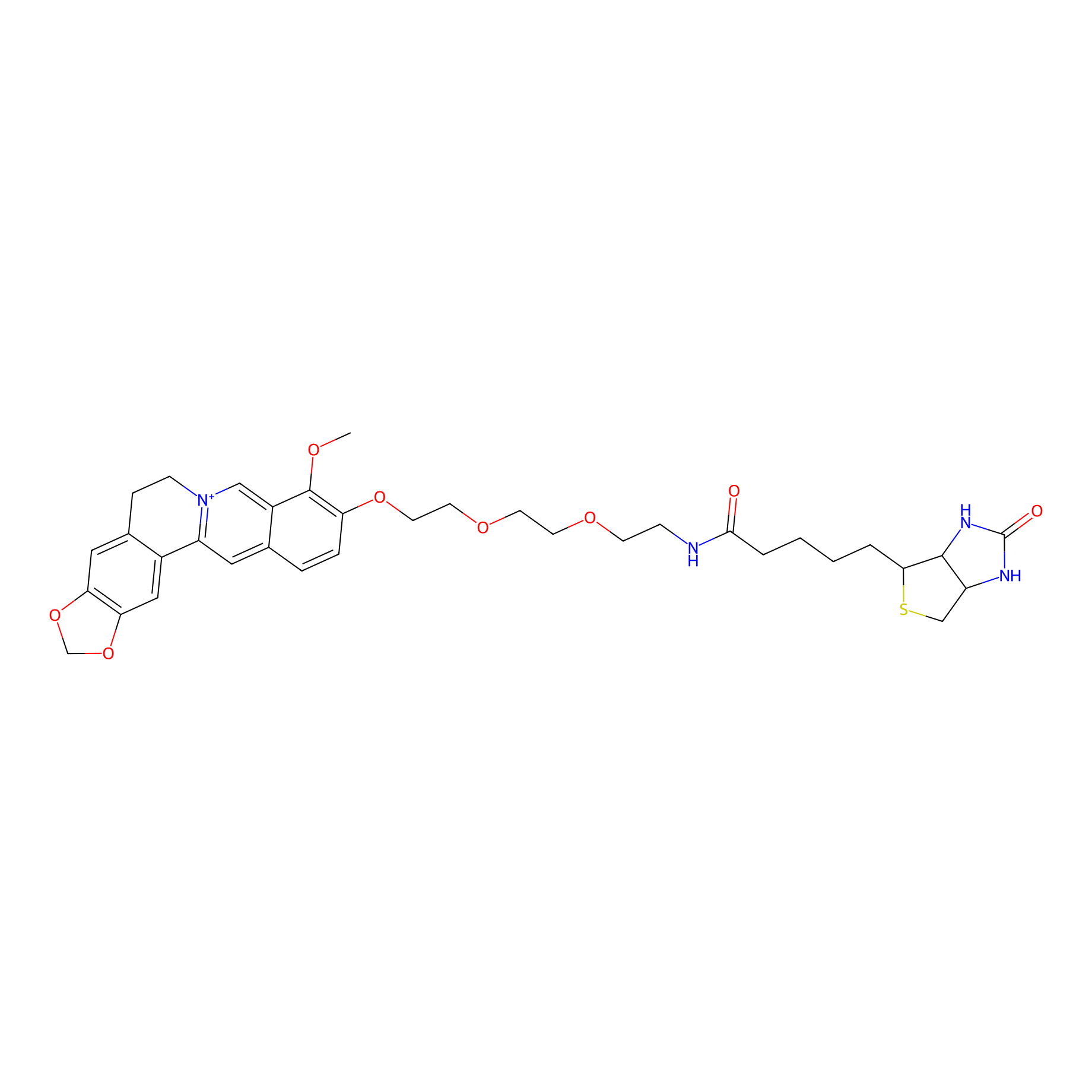 |
N.A. | LDD0305 | [10] | |
|
AOyne Probe Info |
 |
14.80 | LDD0443 | [11] | |
|
NAIA_5 Probe Info |
 |
C404(0.00); C41(0.00); C146(0.00); C234(0.00) | LDD2223 | [6] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C198 Probe Info |
 |
12.04 | LDD1874 | [12] | |
|
VE-P Probe Info |
 |
N.A. | LDD0396 | [13] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0132 | 15d | THP-1 | N.A. | LDD0305 | [10] |
| LDCM0026 | 4SU-RNA+native RNA | HEK-293T | C308(2.23) | LDD0173 | [3] |
| LDCM0156 | Aniline | NCI-H1299 | 12.48 | LDD0403 | [1] |
| LDCM0630 | CCW28-3 | 231MFP | C308(1.47) | LDD2214 | [14] |
| LDCM0632 | CL-Sc | Hep-G2 | C41(20.00); C343(1.76); C146(0.61) | LDD2227 | [6] |
| LDCM0625 | F8 | Ramos | C146(1.36); C343(0.90); C308(1.52); 1.03 | LDD2187 | [15] |
| LDCM0572 | Fragment10 | Ramos | C146(0.66); C343(0.80); 0.94 | LDD2189 | [15] |
| LDCM0573 | Fragment11 | Ramos | C146(0.17); C343(0.95); C308(0.82) | LDD2190 | [15] |
| LDCM0574 | Fragment12 | Ramos | C146(0.73); C343(0.78); 6.08 | LDD2191 | [15] |
| LDCM0575 | Fragment13 | Ramos | C146(0.93); C343(1.04) | LDD2192 | [15] |
| LDCM0576 | Fragment14 | Ramos | C146(0.73); C343(0.60); C308(0.51); 3.30 | LDD2193 | [15] |
| LDCM0579 | Fragment20 | Ramos | C146(0.53); C343(0.61) | LDD2194 | [15] |
| LDCM0580 | Fragment21 | Ramos | C146(0.78); C343(1.02) | LDD2195 | [15] |
| LDCM0582 | Fragment23 | Ramos | C146(0.89); C343(1.20) | LDD2196 | [15] |
| LDCM0578 | Fragment27 | Ramos | C146(0.93) | LDD2197 | [15] |
| LDCM0586 | Fragment28 | Ramos | C146(0.91); C343(0.55); C308(0.83) | LDD2198 | [15] |
| LDCM0588 | Fragment30 | Ramos | C146(1.06); C343(1.06); 2.05 | LDD2199 | [15] |
| LDCM0589 | Fragment31 | Ramos | C146(0.77); C343(0.99) | LDD2200 | [15] |
| LDCM0590 | Fragment32 | Ramos | C146(1.73); C343(0.57) | LDD2201 | [15] |
| LDCM0468 | Fragment33 | Ramos | C146(1.26); C343(0.80) | LDD2202 | [15] |
| LDCM0596 | Fragment38 | Ramos | C146(0.70) | LDD2203 | [15] |
| LDCM0566 | Fragment4 | Ramos | C146(0.94); C343(0.63); C308(0.95) | LDD2184 | [15] |
| LDCM0610 | Fragment52 | Ramos | C146(0.77) | LDD2204 | [15] |
| LDCM0614 | Fragment56 | Ramos | C146(1.07); C343(1.12); 1.78 | LDD2205 | [15] |
| LDCM0569 | Fragment7 | Ramos | C146(0.74); C343(0.59); C308(1.03); 0.85 | LDD2186 | [15] |
| LDCM0571 | Fragment9 | Ramos | C146(0.67); C343(0.67) | LDD2188 | [15] |
| LDCM0022 | KB02 | HEK-293T | C404(1.15); C308(0.98); C146(0.92); C362(1.00) | LDD1492 | [16] |
| LDCM0023 | KB03 | HEK-293T | C404(1.18); C308(1.05); C146(1.06); C362(1.06) | LDD1497 | [16] |
| LDCM0024 | KB05 | HMCB | C41(5.35) | LDD3312 | [2] |
| LDCM0131 | RA190 | MM1.R | C362(1.58) | LDD0304 | [17] |
References
