Details of the Target
General Information of Target
| Target ID | LDTP09509 | |||||
|---|---|---|---|---|---|---|
| Target Name | Rap guanine nucleotide exchange factor 6 (RAPGEF6) | |||||
| Gene Name | RAPGEF6 | |||||
| Gene ID | 51735 | |||||
| Synonyms |
PDZGEF2; Rap guanine nucleotide exchange factor 6; PDZ domain-containing guanine nucleotide exchange factor 2; PDZ-GEF2; RA-GEF-2 |
|||||
| 3D Structure | ||||||
| Sequence |
MNSPVDPGARQALRKKPPERTPEDLNTIYSYLHGMEILSNLREHQLRLMSARARYERYSG
NQVLFCSETIARCWYILLSGSVLVKGSMVLPPCSFGKQFGGKRGCDCLVLEPSEMIVVEN AKDNEDSILQREIPARQSRRRFRKINYKGERQTITDDVEVNSYLSLPADLTKMHLTENPH PQVTHVSSSQSGCSIASDSGSSSLSDIYQATESEVGDVDLTRLPEGPVDSEDDEEEDEEI DRTDPLQGRDLVRECLEKEPADKTDDDIEQLLEFMHQLPAFANMTMSVRRELCSVMIFEV VEQAGAIILEDGQELDSWYVILNGTVEISHPDGKVENLFMGNSFGITPTLDKQYMHGIVR TKVDDCQFVCIAQQDYWRILNHVEKNTHKVEEEGEIVMVHEHRELDRSGTRKGHIVIKAT PERLIMHLIEEHSIVDPTYIEDFLLTYRTFLESPLDVGIKLLEWFKIDSLRDKVTRIVLL WVNNHFNDFEGDPAMTRFLEEFEKNLEDTKMNGHLRLLNIACAAKAKWRQVVLQKASRES PLQFSLNGGSEKGFGIFVEGVEPGSKAADSGLKRGDQIMEVNGQNFENITFMKAVEILRN NTHLALTVKTNIFVFKELLFRTEQEKSGVPHIPKIAEKKSNRHSIQHVPGDIEQTSQEKG SKKVKANTVSGGRNKIRKILDKTRFSILPPKLFSDGGLSQSQDDSIVGTRHCRHSLAIMP IPGTLSSSSPDLLQPTTSMLDFSNPSDIPDQVIRVFKVDQQSCYIIISKDTTAKEVVFHA VHEFGLTGASDTYSLCEVSVTPEGVIKQRRLPDQFSKLADRIQLNGRYYLKNNMETETLC SDEDAQELVKESQLSMLQLSTIEVATQLSMRDFDLFRNIEPTEYIDDLFKLNSKTGNTHL KRFEDIVNQETFWVASEILTEANQLKRMKIIKHFIKIALHCRECKNFNSMFAIISGLNLA SVARLRGTWEKLPSKYEKHLQDLQDIFDPSRNMAKYRNILSSQSMQPPIIPLFPVVKKDM TFLHEGNDSKVDGLVNFEKLRMISKEIRQVVRMTSANMDPAMMFRQRSLSQGSTNSNMLD VQGGAHKKRARRSSLLNAKKLYEDAQMARKVKQYLSSLDVETDEEKFQMMSLQWEPAYGT LTKNLSEKRSAKSSEMSPVPMRSAGQTTKAHLHQPHRVSQVLQVPAVNLHPIRKKGQTKD PALNTSLPQKVLGTTEEISGKKHTEDTISVASSLHSSPPASPQGSPHKGYTLIPSAKSDN LSDSSHSEISSRSSIVSNCSVDSMSAALQDERCSSQALAVPESTGALEKTEHASGIGDHS QHGPGWTLLKPSLIKCLAVSSSVSNEEISQEHIIIEAADSGRGSWTSCSSSSHDNFQSLP NPKSWDFLNSYRHTHLDDPIAEVEPTDSEPYSCSKSCSRTCGQCKGSLERKSWTSSSSLS DTYEPNYGTVKQRVLESTPAESSEGLDPKDATDPVYKTVTSSTEKGLIVYCVTSPKKDDR YREPPPTPPGYLGISLADLKEGPHTHLKPPDYSVAVQRSKMMHNSLSRLPPASLSSNLVA CVPSKIVTQPQRHNLQPFHPKLGDVTDADSEADENEQVSAV |
|||||
| Target Bioclass |
Other
|
|||||
| Subcellular location |
Cytoplasm
|
|||||
| Function | Guanine nucleotide exchange factor (GEF) for Rap1A, Rap2A and M-Ras GTPases. Does not interact with cAMP. | |||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| CHL1 | SNV: p.Y439F | . | |||
| EFO27 | SNV: p.R538H | . | |||
| HT115 | SNV: p.V805A | . | |||
| IGR37 | SNV: p.V707L | DBIA Probe Info | |||
| IGR39 | SNV: p.V707L | DBIA Probe Info | |||
| IM95 | SNV: p.H714Y | DBIA Probe Info | |||
| MCC13 | SNV: p.A717T | . | |||
| NALM6 | SNV: p.A282V | . | |||
| NCIH1993 | SNV: p.G1486C | . | |||
| NCIH2286 | SNV: p.S1267I | . | |||
| NCIH2291 | SNV: p.Y1138C | DBIA Probe Info | |||
| OVCAR8 | SNV: p.T1205A | . | |||
| RL | SNV: p.D820A | . | |||
| YSCCC | SNV: p.Q1071P | . | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
15.00 | LDD0402 | [1] | |
|
STPyne Probe Info |
 |
K682(0.94) | LDD0277 | [2] | |
|
BTD Probe Info |
 |
C941(2.60) | LDD2093 | [3] | |
|
AHL-Pu-1 Probe Info |
 |
C1293(17.21) | LDD0169 | [4] | |
|
EA-probe Probe Info |
 |
N.A. | LDD0440 | [5] | |
|
DBIA Probe Info |
 |
C93(4.05) | LDD0209 | [6] | |
|
IPM Probe Info |
 |
C522(2.72); C941(0.74) | LDD1702 | [3] | |
|
IA-alkyne Probe Info |
 |
C522(0.00); C255(0.00) | LDD0036 | [7] | |
|
Lodoacetamide azide Probe Info |
 |
C1293(0.00); C255(0.00); C522(0.00); C93(0.00) | LDD0037 | [7] | |
|
NAIA_4 Probe Info |
 |
C1293(0.00); C1368(0.00); C1413(0.00) | LDD2226 | [8] | |
|
NAIA_5 Probe Info |
 |
N.A. | LDD2224 | [8] | |
|
TFBX Probe Info |
 |
N.A. | LDD0027 | [9] | |
|
Compound 10 Probe Info |
 |
N.A. | LDD2216 | [10] | |
|
Compound 11 Probe Info |
 |
N.A. | LDD2213 | [10] | |
|
VSF Probe Info |
 |
N.A. | LDD0007 | [11] | |
|
Phosphinate-6 Probe Info |
 |
N.A. | LDD0018 | [12] | |
|
Methacrolein Probe Info |
 |
N.A. | LDD0218 | [13] | |
|
AOyne Probe Info |
 |
15.00 | LDD0443 | [14] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C046 Probe Info |
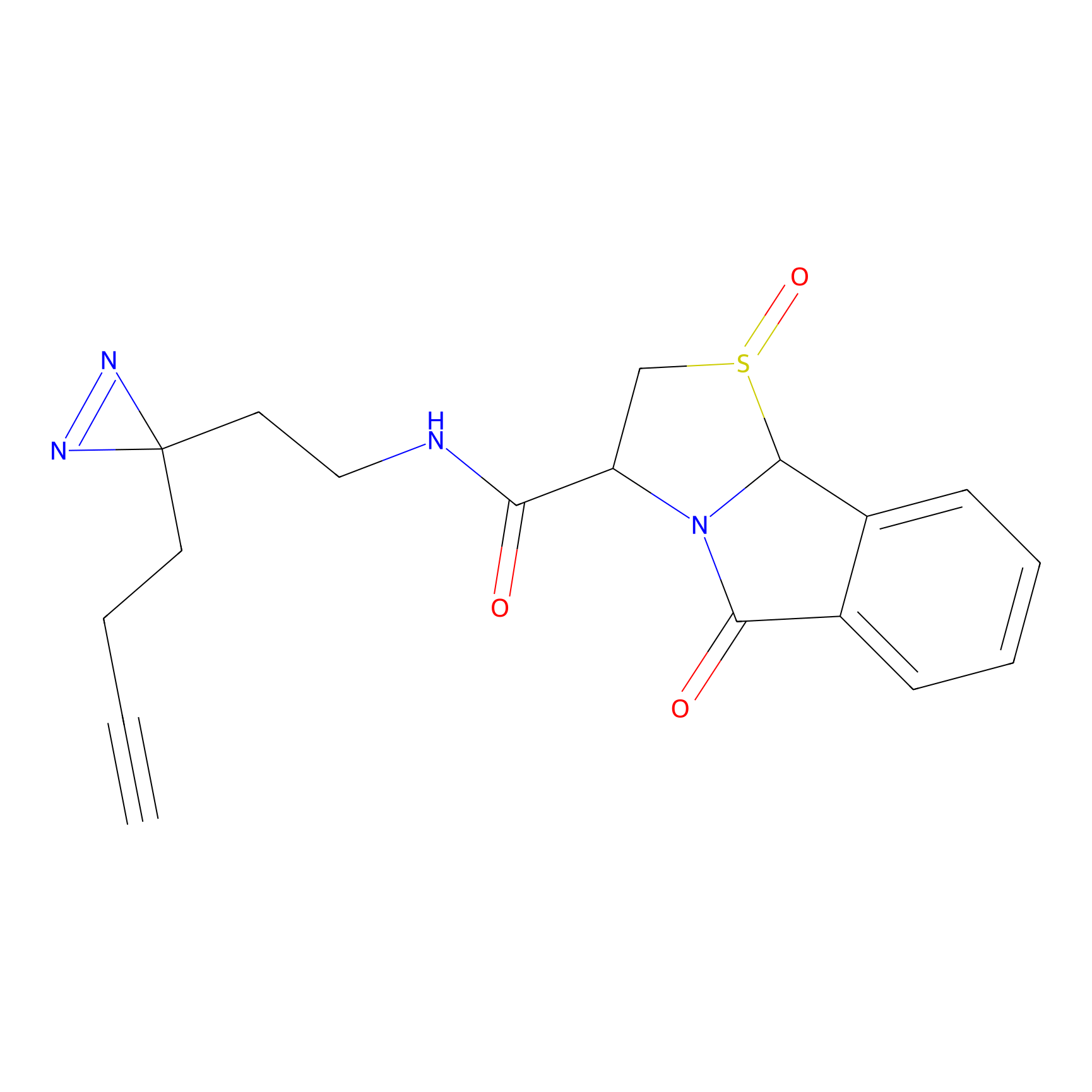 |
12.13 | LDD1745 | [15] | |
|
C064 Probe Info |
 |
8.57 | LDD1761 | [15] | |
|
C347 Probe Info |
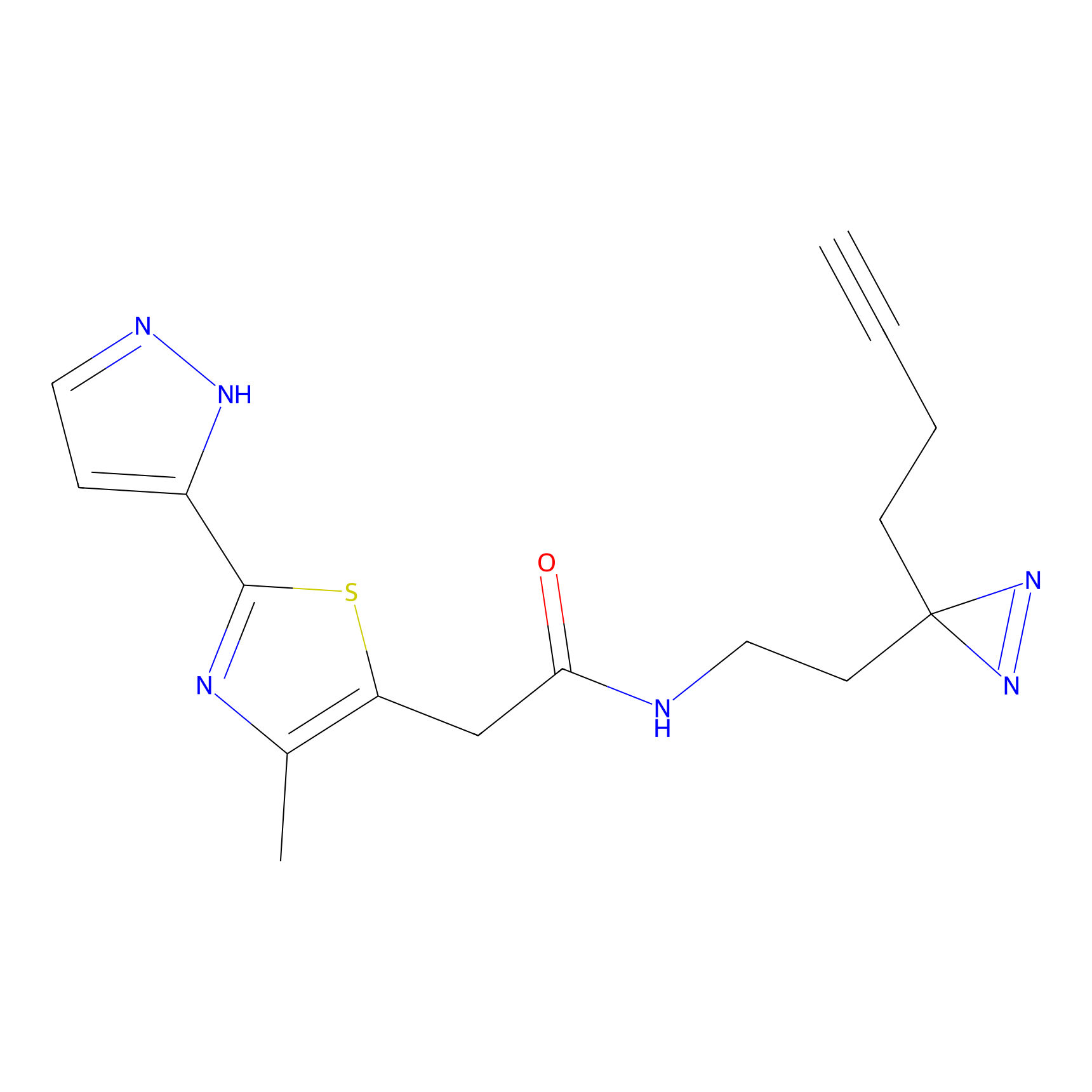 |
8.75 | LDD2008 | [15] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0026 | 4SU-RNA+native RNA | HEK-293T | C1293(17.21) | LDD0169 | [4] |
| LDCM0156 | Aniline | NCI-H1299 | 11.48 | LDD0403 | [1] |
| LDCM0632 | CL-Sc | Hep-G2 | C522(20.00); C1491(0.46) | LDD2227 | [8] |
| LDCM0213 | Electrophilic fragment 2 | MDA-MB-231 | C522(2.72); C941(0.74) | LDD1702 | [3] |
| LDCM0175 | Ethacrynic acid | HeLa | N.A. | LDD0440 | [5] |
| LDCM0625 | F8 | Ramos | C1561(1.01); C93(1.35); C1293(1.50) | LDD2187 | [16] |
| LDCM0572 | Fragment10 | Ramos | C1561(0.92); C1293(1.17); C1368(1.54) | LDD2189 | [16] |
| LDCM0573 | Fragment11 | Ramos | C1561(4.47); C93(0.39); C1293(0.99); C1368(20.00) | LDD2190 | [16] |
| LDCM0574 | Fragment12 | Ramos | C1561(1.07); C93(0.79); C1293(0.94); C1368(3.14) | LDD2191 | [16] |
| LDCM0575 | Fragment13 | Ramos | C1561(0.88); C93(1.09); C1293(1.25); C1368(0.89) | LDD2192 | [16] |
| LDCM0576 | Fragment14 | Ramos | C1561(0.85); C1491(1.04); C93(0.52); C1293(0.94) | LDD2193 | [16] |
| LDCM0579 | Fragment20 | Ramos | C1561(0.90); C93(1.61); C1293(0.64); C1368(3.68) | LDD2194 | [16] |
| LDCM0580 | Fragment21 | Ramos | C1561(0.81); C1491(0.83); C93(0.60); C1293(1.11) | LDD2195 | [16] |
| LDCM0582 | Fragment23 | Ramos | C1561(1.31); C93(0.66); C1293(1.61); C1368(3.28) | LDD2196 | [16] |
| LDCM0578 | Fragment27 | Ramos | C1561(0.91); C1491(0.94); C93(0.78); C1293(1.28) | LDD2197 | [16] |
| LDCM0586 | Fragment28 | Ramos | C1561(0.97); C93(0.44); C1293(0.65); C1368(1.02) | LDD2198 | [16] |
| LDCM0588 | Fragment30 | Ramos | C1561(0.88); C1491(0.61); C93(1.22); C1293(0.82) | LDD2199 | [16] |
| LDCM0589 | Fragment31 | Ramos | C1561(1.00); C1491(0.90); C93(0.82); C1293(0.75) | LDD2200 | [16] |
| LDCM0590 | Fragment32 | Ramos | C1561(0.60); C93(3.00); C1293(0.98) | LDD2201 | [16] |
| LDCM0468 | Fragment33 | Ramos | C1561(0.80); C1491(0.52); C93(0.76); C1293(0.85) | LDD2202 | [16] |
| LDCM0596 | Fragment38 | Ramos | C1561(0.86); C93(0.54); C1293(0.71); C1368(1.02) | LDD2203 | [16] |
| LDCM0566 | Fragment4 | Ramos | C1561(1.73); C93(1.11); C1293(1.11) | LDD2184 | [16] |
| LDCM0610 | Fragment52 | Ramos | C1561(1.05); C93(1.46); C1293(1.35); C1368(1.06) | LDD2204 | [16] |
| LDCM0614 | Fragment56 | Ramos | C1561(1.24); C93(1.77); C1293(1.92); C1368(1.19) | LDD2205 | [16] |
| LDCM0569 | Fragment7 | Ramos | C1561(1.30); C93(1.40); C1293(0.80); C1368(1.03) | LDD2186 | [16] |
| LDCM0571 | Fragment9 | Ramos | C1561(1.14); C1293(1.04); C1368(1.29) | LDD2188 | [16] |
| LDCM0022 | KB02 | HEK-293T | C1413(0.94); C93(0.83); C522(0.97); C1561(1.01) | LDD1492 | [17] |
| LDCM0023 | KB03 | Jurkat | C93(4.05) | LDD0209 | [6] |
| LDCM0024 | KB05 | HMCB | C522(5.99) | LDD3312 | [18] |
| LDCM0500 | Nucleophilic fragment 13a | MDA-MB-231 | C941(2.60) | LDD2093 | [3] |
The Interaction Atlas With This Target
References
