Details of the Target
General Information of Target
| Target ID | LDTP07908 | |||||
|---|---|---|---|---|---|---|
| Target Name | Histone H2A.V (H2AZ2) | |||||
| Gene Name | H2AZ2 | |||||
| Gene ID | 94239 | |||||
| Synonyms |
H2AFV; H2AV; Histone H2A.V; H2A.F/Z; H2A.Z variant histone 2 |
|||||
| 3D Structure | ||||||
| Sequence |
MAGGKAGKDSGKAKAKAVSRSQRAGLQFPVGRIHRHLKTRTTSHGRVGATAAVYSAAILE
YLTAEVLELAGNASKDLKVKRITPRHLQLAIRGDEELDSLIKATIAGGGVIPHIHKSLIG KKGQQKTA |
|||||
| Target Bioclass |
Other
|
|||||
| Family |
Histone H2A family
|
|||||
| Subcellular location |
Nucleus
|
|||||
| Function |
Variant histone H2A which replaces conventional H2A in a subset of nucleosomes. Nucleosomes wrap and compact DNA into chromatin, limiting DNA accessibility to the cellular machineries which require DNA as a template. Histones thereby play a central role in transcription regulation, DNA repair, DNA replication and chromosomal stability. DNA accessibility is regulated via a complex set of post-translational modifications of histones, also called histone code, and nucleosome remodeling. May be involved in the formation of constitutive heterochromatin. May be required for chromosome segregation during cell division.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
ONAyne Probe Info |
 |
K8(1.64) | LDD0274 | [2] | |
|
STPyne Probe Info |
 |
K16(6.67) | LDD0277 | [2] | |
|
W1 Probe Info |
 |
H86(1.01) | LDD0239 | [3] | |
|
NHS Probe Info |
 |
N.A. | LDD0010 | [4] | |
|
OSF Probe Info |
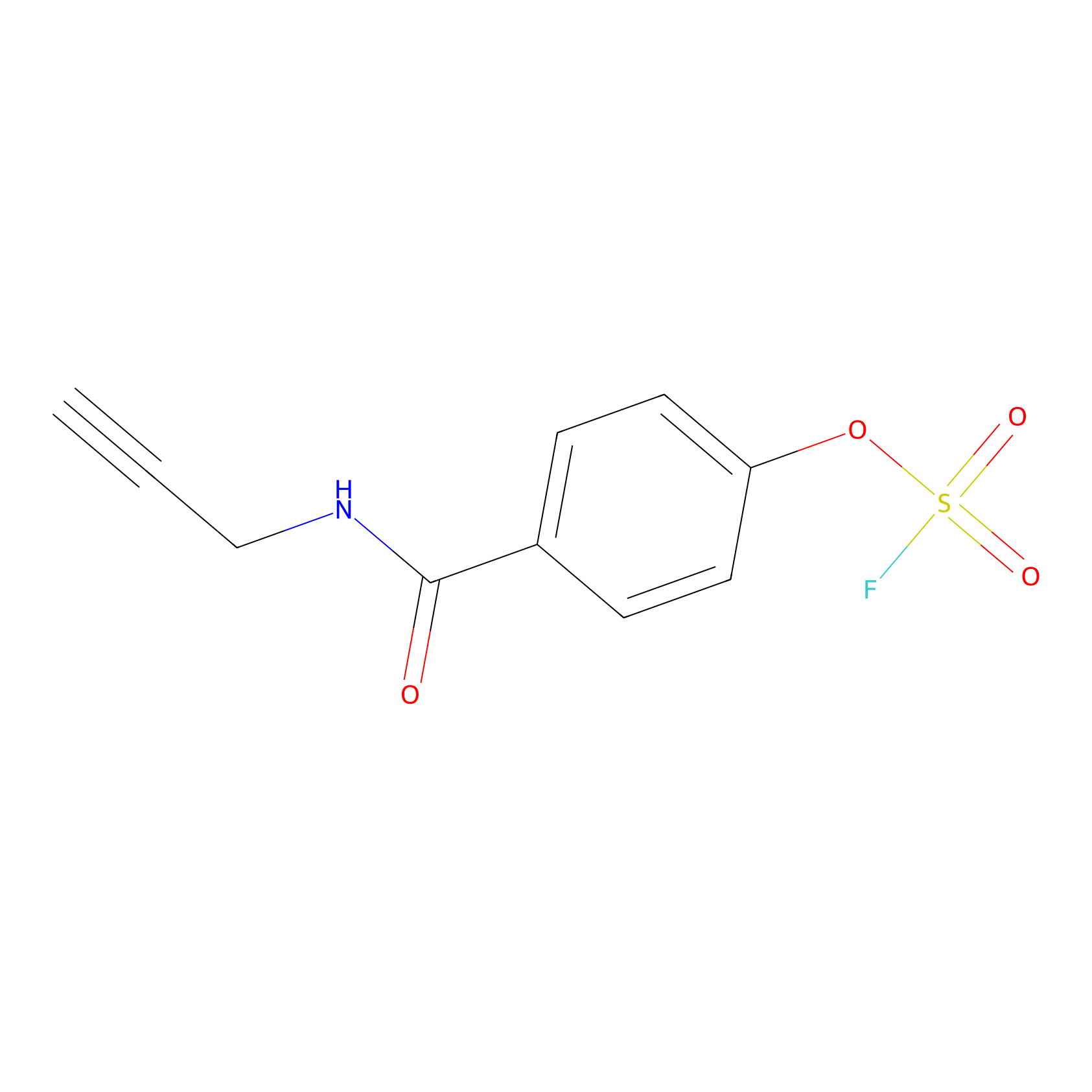 |
N.A. | LDD0029 | [5] | |
|
SF Probe Info |
 |
K116(0.00); K8(0.00) | LDD0028 | [5] | |
|
Acrolein Probe Info |
 |
H113(0.00); H115(0.00) | LDD0217 | [6] | |
|
Crotonaldehyde Probe Info |
 |
H113(0.00); H115(0.00) | LDD0219 | [6] | |
|
Methacrolein Probe Info |
 |
H113(0.00); H115(0.00) | LDD0218 | [6] | |
|
HHS-465 Probe Info |
 |
N.A. | LDD2240 | [7] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
BD-F Probe Info |
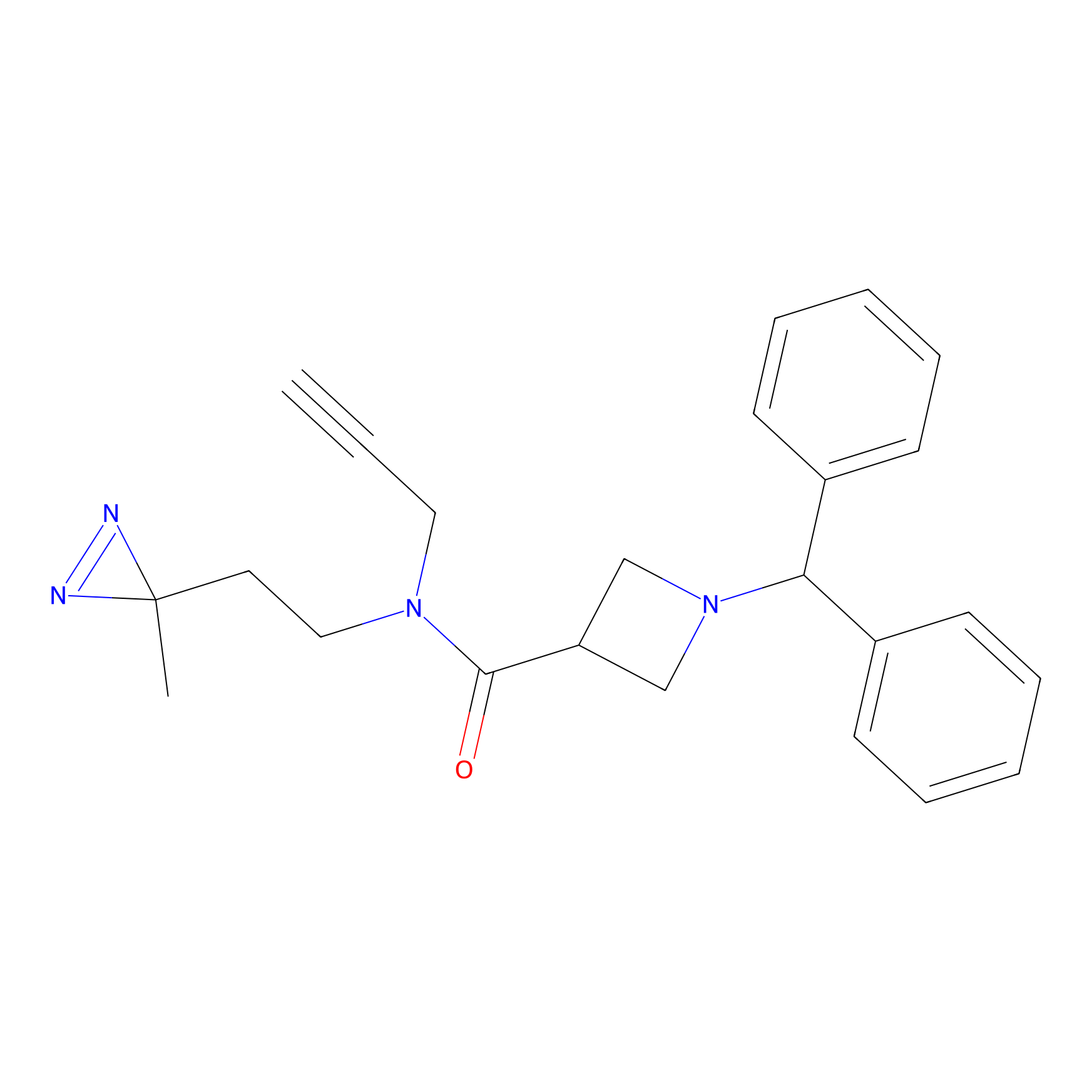 |
E68(0.00); Y61(0.00); A64(0.00); I58(0.00) | LDD0024 | [8] | |
|
LD-F Probe Info |
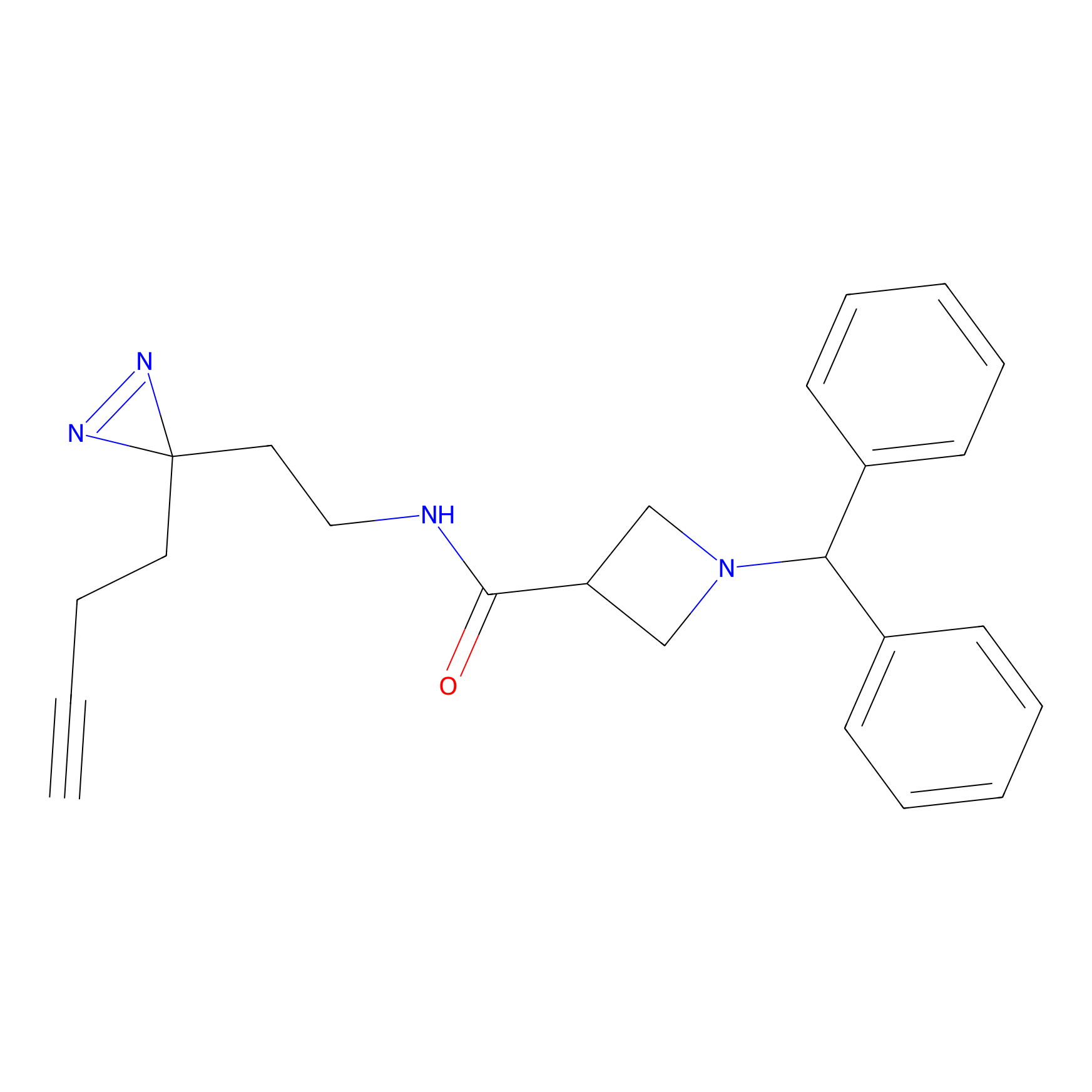 |
E60(0.00); E65(0.00); T63(0.00); L59(0.00) | LDD0015 | [8] | |
|
TM-F Probe Info |
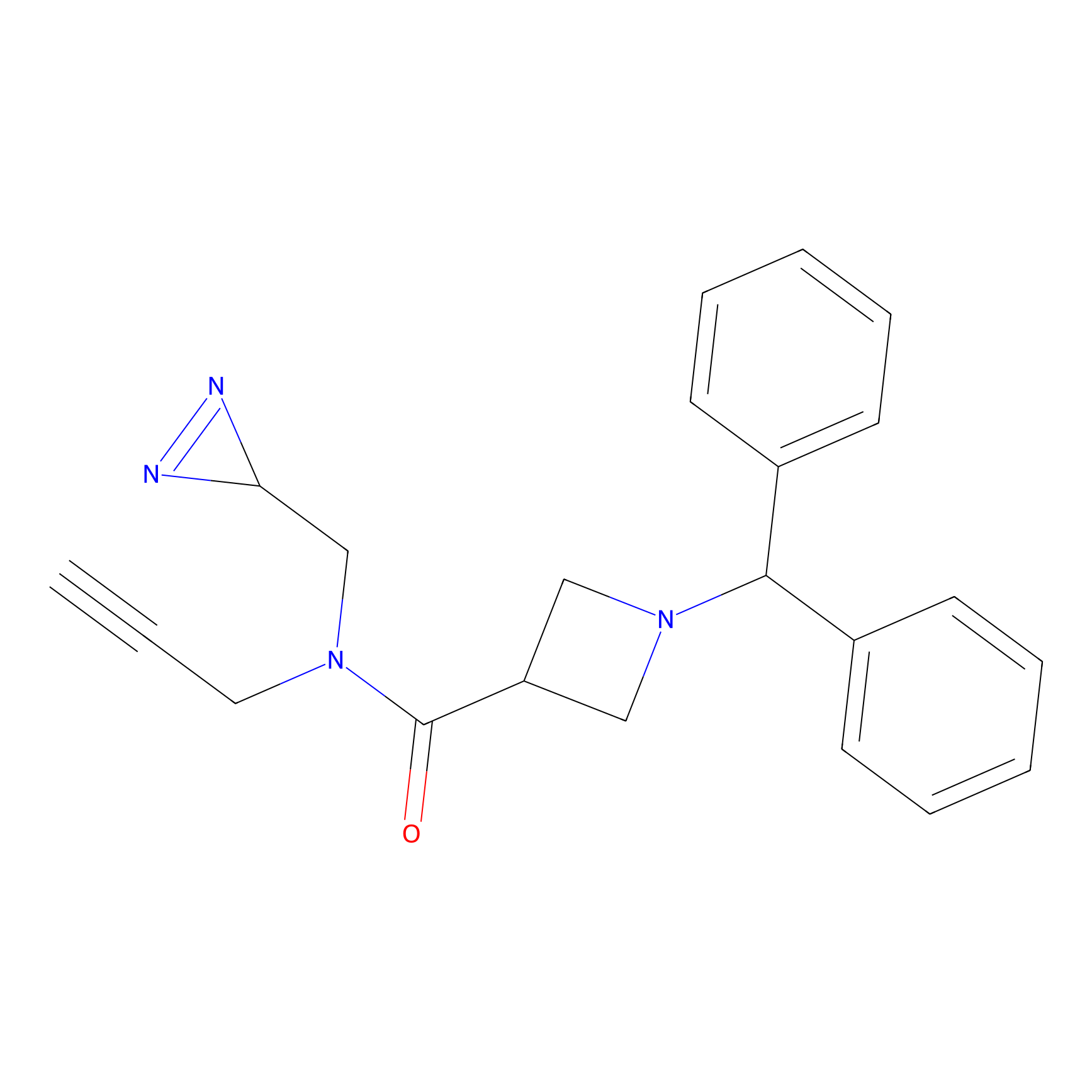 |
L59(0.00); L67(0.00); A56(0.00); V66(0.00) | LDD0020 | [8] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0108 | Chloroacetamide | HeLa | H115(0.00); H113(0.00) | LDD0222 | [6] |
| LDCM0107 | IAA | HeLa | H113(0.00); H115(0.00) | LDD0221 | [6] |
| LDCM0109 | NEM | HeLa | H113(0.00); H115(0.00); H86(0.00) | LDD0223 | [6] |
| LDCM0112 | W16 | Hep-G2 | H86(1.01) | LDD0239 | [3] |
| LDCM0113 | W17 | Hep-G2 | H86(0.93) | LDD0240 | [3] |
References
