Details of the Target
General Information of Target
| Target ID | LDTP07401 | |||||
|---|---|---|---|---|---|---|
| Target Name | Ragulator complex protein LAMTOR1 (LAMTOR1) | |||||
| Gene Name | LAMTOR1 | |||||
| Gene ID | 55004 | |||||
| Synonyms |
C11orf59; PDRO; Ragulator complex protein LAMTOR1; Late endosomal/lysosomal adaptor and MAPK and MTOR activator 1; Lipid raft adaptor protein p18; Protein associated with DRMs and endosomes; p27Kip1-releasing factor from RhoA; p27RF-Rho
|
|||||
| 3D Structure | ||||||
| Sequence |
MGCCYSSENEDSDQDREERKLLLDPSSPPTKALNGAEPNYHSLPSARTDEQALLSSILAK
TASNIIDVSAADSQGMEQHEYMDRARQYSTRLAVLSSSLTHWKKLPPLPSLTSQPHQVLA SEPIPFSDLQQVSRIAAYAYSALSQIRVDAKEELVVQFGIP |
|||||
| Target Bioclass |
Other
|
|||||
| Family |
LAMTOR1 family
|
|||||
| Subcellular location |
Late endosome membrane
|
|||||
| Function |
Key component of the Ragulator complex, a multiprotein complex involved in amino acid sensing and activation of mTORC1, a signaling complex promoting cell growth in response to growth factors, energy levels, and amino acids. Activated by amino acids through a mechanism involving the lysosomal V-ATPase, the Ragulator plays a dual role for the small GTPases Rag (RagA/RRAGA, RagB/RRAGB, RagC/RRAGC and/or RagD/RRAGD): it (1) acts as a guanine nucleotide exchange factor (GEF), activating the small GTPases Rag and (2) mediates recruitment of Rag GTPases to the lysosome membrane. Activated Ragulator and Rag GTPases function as a scaffold recruiting mTORC1 to lysosomes where it is in turn activated. LAMTOR1 is directly responsible for anchoring the Ragulator complex to the lysosomal membrane. LAMTOR1 wraps around the other subunits of the Ragulator complex to hold them in place and interacts with the Rag GTPases, thereby playing a key role in the recruitment of the mTORC1 complex to lysosomes. Also involved in the control of embryonic stem cells differentiation via non-canonical RagC/RRAGC and RagD/RRAGD activation: together with FLCN, it is necessary to recruit and activate RagC/RRAGC and RagD/RRAGD at the lysosomes, and to induce exit of embryonic stem cells from pluripotency via non-canonical, mTOR-independent TFE3 inactivation. Also required for late endosomes/lysosomes biogenesis it may regulate both the recycling of receptors through endosomes and the MAPK signaling pathway through recruitment of some of its components to late endosomes. May be involved in cholesterol homeostasis regulating LDL uptake and cholesterol release from late endosomes/lysosomes. May also play a role in RHOA activation.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
7.72 | LDD0402 | [1] | |
|
STPyne Probe Info |
 |
K103(10.00); K20(1.49); K60(6.67) | LDD0277 | [2] | |
|
Probe 1 Probe Info |
 |
Y40(10.90) | LDD3495 | [3] | |
|
Alkyne-RA190 Probe Info |
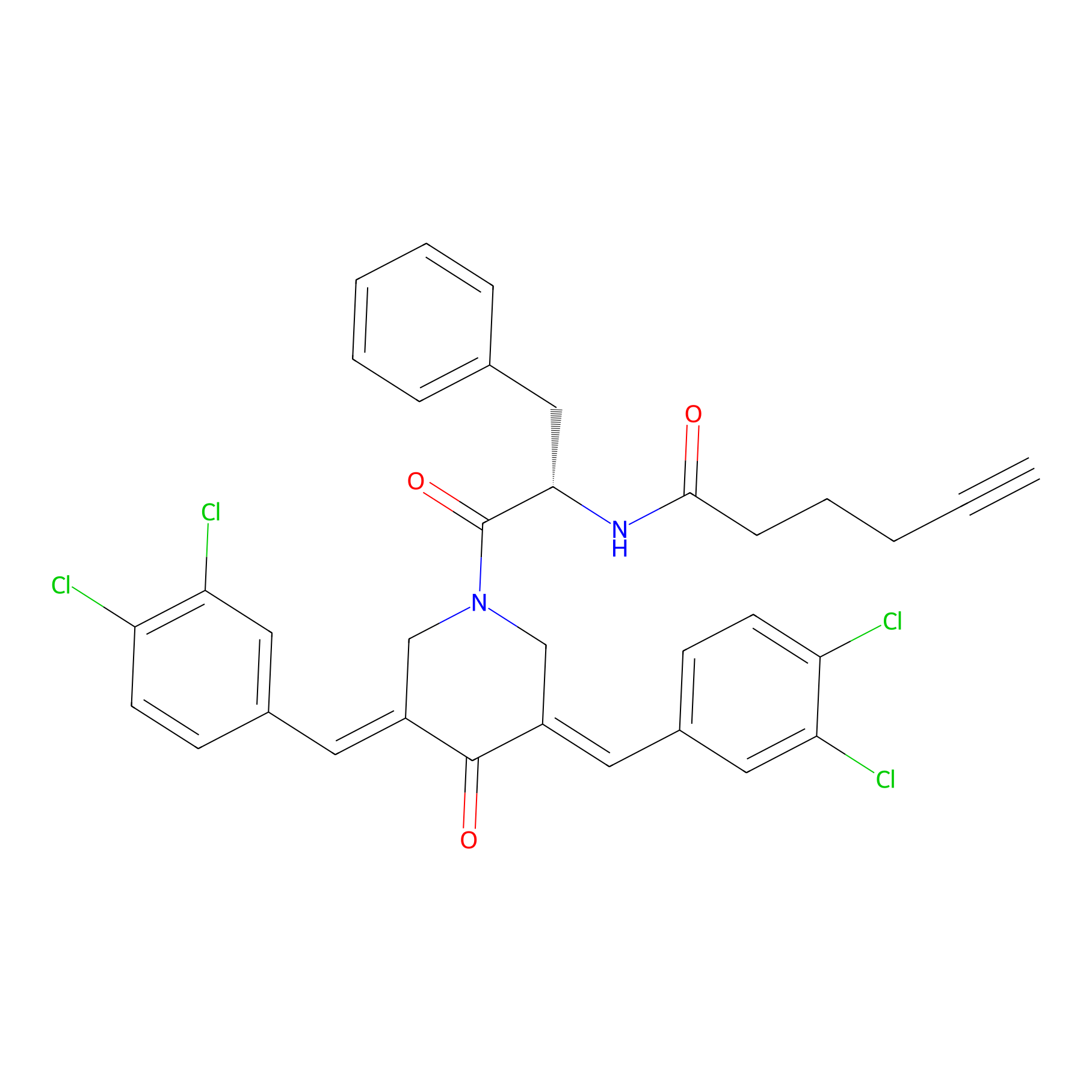 |
2.25 | LDD0299 | [4] | |
|
5E-2FA Probe Info |
 |
H41(0.00); H116(0.00) | LDD2235 | [5] | |
|
SF Probe Info |
 |
Y40(0.00); Y138(0.00) | LDD0028 | [6] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0217 | [7] | |
|
AOyne Probe Info |
 |
15.00 | LDD0443 | [8] | |
|
HHS-465 Probe Info |
 |
N.A. | LDD2240 | [9] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C310 Probe Info |
 |
7.84 | LDD1977 | [10] | |
|
FFF probe11 Probe Info |
 |
8.11 | LDD0471 | [11] | |
|
FFF probe12 Probe Info |
 |
9.71 | LDD0473 | [11] | |
|
FFF probe13 Probe Info |
 |
13.90 | LDD0475 | [11] | |
|
FFF probe14 Probe Info |
 |
20.00 | LDD0477 | [11] | |
|
FFF probe2 Probe Info |
 |
14.11 | LDD0463 | [11] | |
|
FFF probe3 Probe Info |
 |
11.17 | LDD0464 | [11] | |
Competitor(s) Related to This Target
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transporter and channel
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Neutral amino acid transporter 9 (SLC38A9) | Amino acid/polyamine transporter 2 family | Q8NBW4 | |||
Immunoglobulin
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Platelet endothelial cell adhesion molecule (PECAM1) | . | P16284 | |||
Other
References
