Details of the Target
General Information of Target
| Target ID | LDTP06870 | |||||
|---|---|---|---|---|---|---|
| Target Name | Protein GREB1 (GREB1) | |||||
| Gene Name | GREB1 | |||||
| Gene ID | 9687 | |||||
| Synonyms |
KIAA0575; Protein GREB1; Gene regulated in breast cancer 1 protein |
|||||
| 3D Structure | ||||||
| Sequence |
MGNSYAGQLKTTRFEEVLHNSIEASLRSNNLVPRPIFSQLYLEAEQQLAALEGGSRVDNE
EEEEEGEGGLETNGPPNPFQLHPLPEGCCTTDGFCQAGKDLRLVSISNEPMDVPAGFLLV GVKSPSLPDHLLVCAVDKRFLPDDNGHNALLGFSGNCVGCGKKGFCYFTEFSNHINLKLT TQPKKQKHLKYYLVRNAQGTLTKGPLICWKGSEFRSRQIPASTCSSSLFPALESTAAFPS EPVPGTNPSILMGAQQAGPASDHPSLNAAMGPAVFNGKDSPKCQQLAKNNLLALPRPSAL GILSNSGPPKKRHKGWSPESPSAPDGGCPQGGGNRAKYESAGMSCVPQVGLVGPASVTFP VVASGEPVSVPDNLLKICKAKPVIFKGHGNFPYLCGNLNDVVVSPLLYTCYQNSQSVSRA YEQYGASAIQPISEEMQLLLTVYYLVQLAADQVPLMEDLEQIFLRSWRESHLTEIRQYQQ APPQPFPPAPSAAAPVTSAQLPWLASLAASSCNDSVHVIECAYSLAEGLSEMFRLLVEGK LAKTNYVVIICACRSAAIDSCIAVTGKYQARILSESLLTPAEYQKEVNYELVTGKVDSLG AFFSTLCPEGDIDILLDKFHQENQGHISSSLAASSVTKAASLDVSGTPVCTSYNLEPHSI RPFQLAVAQKLLSHVCSIADSSTQNLDLGSFEKVDFLICIPPSEVTYQQTLLHVWHSGVL LELGLKKEHMTKQRVEQYVLKLDTEAQTKFKAFLQNSFQNPHTLFVLIHDHAHWDLVSST VHNLYSQSDPSVGLVDRLLNCREVKEAPNIVTLHVTSFPYALQTQHTLISPYNEIHWPAS CSNGVDLYHENKKYFGLSEFIESTLSGHSLPLLRYDSSFEAMVTALGKRFPRLHSAVIRT FVLVQHYAAALMAVSGLPQMKNYTSVETLEITQNLLNSPKQCPCGHGLMVLLRVPCSPLA VVAYERLAHVRARLALEEHFEIILGSPSSGVTVGKHFVKQLRMWQKIEDVEWRPQTYLEL EGLPCILIFSGMDPHGESLPRSLRYCDLRLINSSCLVRTALEQELGLAAYFVSNEVPLEK GARNEALESDAEKLSSTDNEDEELGTEGSTSEKRSPMKRERSRSHDSASSSLSSKASGSA LGGESSAQPTALPQGEHARSPQPRGPAEEGRAPGEKQRPRASQGPPSAISRHSPGPTPQP DCSLRTGQRSVQVSVTSSCSQLSSSSGSSSSSVAPAAGTWVLQASQCSLTKACRQPPIVF LPKLVYDMVVSTDSSGLPKAASLLPSPSVMWASSFRPLLSKTMTSTEQSLYYRQWTVPRP SHMDYGNRAEGRVDGFHPRRLLLSGPPQIGKTGAYLQFLSVLSRMLVRLTEVDVYDEEEI NINLREESDWHYLQLSDPWPDLELFKKLPFDYIIHDPKYEDASLICSHYQGIKSEDRGMS RKPEDLYVRRQTARMRLSKYAAYNTYHHCEQCHQYMGFHPRYQLYESTLHAFAFSYSMLG EEIQLHFIIPKSKEHHFVFSQPGGQLESMRLPLVTDKSHEYIKSPTFTPTTGRHEHGLFN LYHAMDGASHLHVLVVKEYEMAIYKKYWPNHIMLVLPSIFNSAGVGAAHFLIKELSYHNL ELERNRQEELGIKPQDIWPFIVISDDSCVMWNVVDVNSAGERSREFSWSERNVSLKHIMQ HIEAAPDIMHYALLGLRKWSSKTRASEVQEPFSRCHVHNFIILNVDLTQNVQYNQNRFLC DDVDFNLRVHSAGLLLCRFNRFSVMKKQIVVGGHRSFHITSKVSDNSAAVVPAQYICAPD SKHTFLAAPAQLLLEKFLQHHSHLFFPLSLKNHDHPVLSVDCYLNLGSQISVCYVSSRPH SLNISCSDLLFSGLLLYLCDSFVGASFLKKFHFLKGATLCVICQDRSSLRQTVVRLELED EWQFRLRDEFQTANAREDRPLFFLTGRHI |
|||||
| Target Bioclass |
Other
|
|||||
| Family |
GREB1 family
|
|||||
| Subcellular location |
Membrane
|
|||||
| Function | May play a role in estrogen-stimulated cell proliferation. Acts as a regulator of hormone-dependent cancer growth in breast and prostate cancers. | |||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| 22RV1 | SNV: p.S298P | . | |||
| A172 | SNV: p.L18F | . | |||
| A2058 | SNV: p.S1271F | . | |||
| AN3CA | SNV: p.P1235L | . | |||
| CALU6 | SNV: p.R1947Ter | . | |||
| CCK81 | Insertion: p.S1288LfsTer89 SNV: p.H471R |
. | |||
| CHL1 | SNV: p.F1924V | . | |||
| DOTC24510 | SNV: p.F663L | . | |||
| DU145 | SNV: p.G98R;p.K1513Ter Substitution: p.E722Ter |
. | |||
| HCC1419 | SNV: p.L127P | . | |||
| HT115 | SNV: p.P392T | . | |||
| HT1376 | SNV: p.D876H | . | |||
| IGROV1 | SNV: p.Y738C | DBIA Probe Info | |||
| JHH4 | SNV: p.W1291Ter | . | |||
| JURKAT | SNV: p.D1033G | . | |||
| MFE319 | SNV: p.L459M | . | |||
| NCIH1993 | SNV: p.R1449L | . | |||
| OAW28 | SNV: p.E1614Q | . | |||
| OSRC2 | SNV: p.S1096C | . | |||
| RKO | SNV: p.K1301E | . | |||
| SBC5 | SNV: p.T12I | . | |||
| SKMEL5 | SNV: p.Q1212Ter | . | |||
| SW1116 | SNV: p.G1567S | . | |||
| SW1990 | SNV: p.R1939L | . | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
STPyne Probe Info |
 |
K1176(9.03) | LDD0277 | [1] | |
|
IPM Probe Info |
 |
C1202(0.00); C1055(0.00) | LDD0241 | [2] | |
|
Probe 1 Probe Info |
 |
Y738(20.21); Y1447(36.27) | LDD3495 | [3] | |
|
DBIA Probe Info |
 |
C200(1.84) | LDD3340 | [4] | |
|
Curcusone 37 Probe Info |
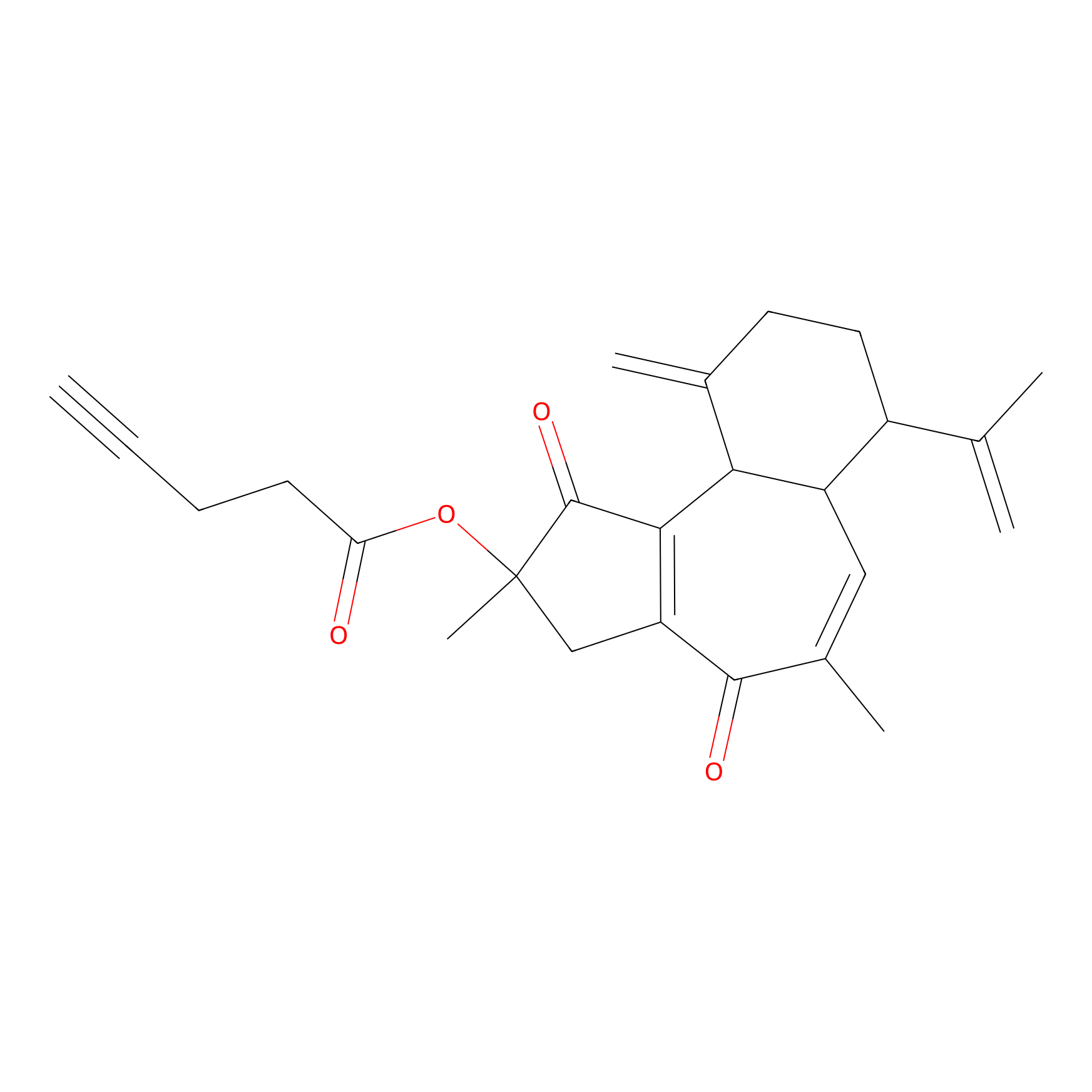 |
4.36 | LDD0188 | [5] | |
|
AHL-Pu-1 Probe Info |
 |
C1202(6.09) | LDD0171 | [6] | |
|
NAIA_5 Probe Info |
 |
C1253(0.00); C166(0.00); C328(0.00); C208(0.00) | LDD2223 | [7] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0026 | 4SU-RNA+native RNA | DM93 | C1202(6.09) | LDD0171 | [6] |
| LDCM0284 | AC24 | HEK-293T | C208(0.94) | LDD1523 | [8] |
| LDCM0293 | AC32 | HEK-293T | C208(0.89) | LDD1532 | [8] |
| LDCM0302 | AC40 | HEK-293T | C208(0.93) | LDD1541 | [8] |
| LDCM0310 | AC48 | HEK-293T | C208(1.04) | LDD1549 | [8] |
| LDCM0319 | AC56 | HEK-293T | C208(1.05) | LDD1558 | [8] |
| LDCM0328 | AC64 | HEK-293T | C208(1.01) | LDD1567 | [8] |
| LDCM0345 | AC8 | HEK-293T | C208(1.02) | LDD1569 | [8] |
| LDCM0275 | AKOS034007705 | HEK-293T | C208(1.04) | LDD1514 | [8] |
| LDCM0632 | CL-Sc | Hep-G2 | C345(20.00); C1055(1.58); C561(1.27); C956(1.27) | LDD2227 | [7] |
| LDCM0367 | CL1 | HEK-293T | C208(1.19) | LDD1571 | [8] |
| LDCM0369 | CL100 | HEK-293T | C208(0.99) | LDD1573 | [8] |
| LDCM0370 | CL101 | HEK-293T | C208(1.08) | LDD1574 | [8] |
| LDCM0373 | CL104 | HEK-293T | C208(0.88) | LDD1577 | [8] |
| LDCM0374 | CL105 | HEK-293T | C208(0.99) | LDD1578 | [8] |
| LDCM0377 | CL108 | HEK-293T | C208(1.02) | LDD1581 | [8] |
| LDCM0378 | CL109 | HEK-293T | C208(0.95) | LDD1582 | [8] |
| LDCM0382 | CL112 | HEK-293T | C208(0.96) | LDD1586 | [8] |
| LDCM0383 | CL113 | HEK-293T | C208(1.20) | LDD1587 | [8] |
| LDCM0386 | CL116 | HEK-293T | C208(0.79) | LDD1590 | [8] |
| LDCM0387 | CL117 | HEK-293T | C208(1.07) | LDD1591 | [8] |
| LDCM0390 | CL12 | HEK-293T | C208(1.02) | LDD1594 | [8] |
| LDCM0391 | CL120 | HEK-293T | C208(0.89) | LDD1595 | [8] |
| LDCM0392 | CL121 | HEK-293T | C208(0.95) | LDD1596 | [8] |
| LDCM0395 | CL124 | HEK-293T | C208(0.85) | LDD1599 | [8] |
| LDCM0396 | CL125 | HEK-293T | C208(0.91) | LDD1600 | [8] |
| LDCM0399 | CL128 | HEK-293T | C208(0.92) | LDD1603 | [8] |
| LDCM0400 | CL13 | HEK-293T | C208(1.13) | LDD1604 | [8] |
| LDCM0403 | CL16 | HEK-293T | C208(0.81) | LDD1607 | [8] |
| LDCM0412 | CL24 | HEK-293T | C208(1.03) | LDD1616 | [8] |
| LDCM0413 | CL25 | HEK-293T | C208(0.88) | LDD1617 | [8] |
| LDCM0416 | CL28 | HEK-293T | C208(0.93) | LDD1620 | [8] |
| LDCM0425 | CL36 | HEK-293T | C208(1.00) | LDD1629 | [8] |
| LDCM0426 | CL37 | HEK-293T | C208(0.84) | LDD1630 | [8] |
| LDCM0429 | CL4 | HEK-293T | C208(0.89) | LDD1633 | [8] |
| LDCM0430 | CL40 | HEK-293T | C208(0.80) | LDD1634 | [8] |
| LDCM0438 | CL48 | HEK-293T | C208(0.99) | LDD1642 | [8] |
| LDCM0439 | CL49 | HEK-293T | C208(0.94) | LDD1643 | [8] |
| LDCM0443 | CL52 | HEK-293T | C208(1.03) | LDD1646 | [8] |
| LDCM0452 | CL60 | HEK-293T | C208(1.04) | LDD1655 | [8] |
| LDCM0453 | CL61 | HEK-293T | C208(1.04) | LDD1656 | [8] |
| LDCM0456 | CL64 | HEK-293T | C208(0.85) | LDD1659 | [8] |
| LDCM0465 | CL72 | HEK-293T | C208(1.00) | LDD1668 | [8] |
| LDCM0466 | CL73 | HEK-293T | C208(0.96) | LDD1669 | [8] |
| LDCM0469 | CL76 | HEK-293T | C208(0.87) | LDD1672 | [8] |
| LDCM0478 | CL84 | HEK-293T | C208(0.96) | LDD1681 | [8] |
| LDCM0479 | CL85 | HEK-293T | C208(0.92) | LDD1682 | [8] |
| LDCM0482 | CL88 | HEK-293T | C208(0.94) | LDD1685 | [8] |
| LDCM0491 | CL96 | HEK-293T | C208(0.90) | LDD1694 | [8] |
| LDCM0492 | CL97 | HEK-293T | C208(1.07) | LDD1695 | [8] |
| LDCM0033 | Curcusone1d | MCF-7 | 4.36 | LDD0188 | [5] |
| LDCM0634 | CY-0357 | Hep-G2 | C1202(1.16); C956(0.53) | LDD2228 | [7] |
| LDCM0022 | KB02 | A2058 | C200(1.49) | LDD2253 | [4] |
| LDCM0023 | KB03 | A2058 | C200(1.93) | LDD2670 | [4] |
| LDCM0024 | KB05 | NB1 | C200(1.84) | LDD3340 | [4] |
References
